The goal of jab is to conveniently calculate Jeffrey’s approximate Bayes factor (JAB; Wagenmakers, 2022) for a wide variety of statistical analyses.
You can install the development version of jab like so:
remotes::install_github("crsh/jab")jab automatically supports calculation of JAB for any analysis that outputs a Wald test and for which broom returns an estimate and a standard error. The user additionally needs to specify a prior distribution for estimate in the scale used to calculate the Wald statistic.
Take the example of standard linear regression. JAB can be easily
calculated for all regression coefficients. We simply submit the results
from the orthodox frequentist analysis to jab() and specify a prior
distribution—let’s use a scaled central Cauchy distribution. Note that
JAB gives evidence for the null hypothesis relative to the alternative.
library("jab")
library("ggplot2")
# Fit regression model
data(attitude)
attitude_z <- data.frame(scale(attitude))
attitude_lm <- lm(rating ~ 0 + ., data = attitude_z)
attitude_tidy_lm <- broom::tidy(attitude_lm)
attitude_tidy_lm
#> # A tibble: 6 × 5
#> term estimate std.error statistic p.value
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 complaints 0.671 0.172 3.89 0.000694
#> 2 privileges -0.0734 0.134 -0.550 0.588
#> 3 learning 0.309 0.159 1.94 0.0640
#> 4 raises 0.0698 0.185 0.377 0.710
#> 5 critical 0.0312 0.117 0.267 0.792
#> 6 advance -0.183 0.147 -1.24 0.225
# Specify prior distribution and approximate Bayes factor
attitude_jab <- jab(
attitude_lm
, prior = dcauchy
, location = 0
, scale = sqrt(2) / 4
)
attitude_jab
#> complaints privileges learning raises critical advance
#> 0.03751703 2.98754241 0.88364001 2.31936704 3.68709775 1.82951234Now compare this with the Jeffreys-Zellner-Siow (JZS) Bayes factor from
BayesFactor::regressionBF() with the same prior distribution.
# Calculate JZS-Bayes factor
attitude_jzs <- BayesFactor::regressionBF(
rating ~ .
, data = attitude
, rscaleCont = sqrt(2) / 4
, whichModels = "top"
, progress = FALSE
)
# Compare results
tibble::tibble(
predictor = attitude_tidy_lm$term
# Frequentist p-values
, p = attitude_tidy_lm$p.value
# Bayes factors in favor of the null hypothesis
, jab = attitude_jab
, jzs = rev(as.vector(attitude_jzs))
# Naive posterior probabilities
, jab_pp = jab / (jab + 1)
, jzs_pp = jzs / (jzs + 1)
)
#> # A tibble: 6 × 6
#> predictor p jab jzs jab_pp jzs_pp
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 complaints 0.000694 0.0375 0.0231 0.0362 0.0225
#> 2 privileges 0.588 2.99 2.92 0.749 0.745
#> 3 learning 0.0640 0.884 0.727 0.469 0.421
#> 4 raises 0.710 2.32 3.13 0.699 0.758
#> 5 critical 0.792 3.69 3.23 0.787 0.764
#> 6 advance 0.225 1.83 1.73 0.647 0.634Pretty close!
To vary the scale of the prior distribution, simply pass a vector of scaling parameters, one scale for each coefficient.
jab(
attitude_lm
, prior = dcauchy
, location = 0
, scale = c(rep(0.5, 3), rep(sqrt(2) / 4, 3))
)
#> complaints privileges learning raises critical advance
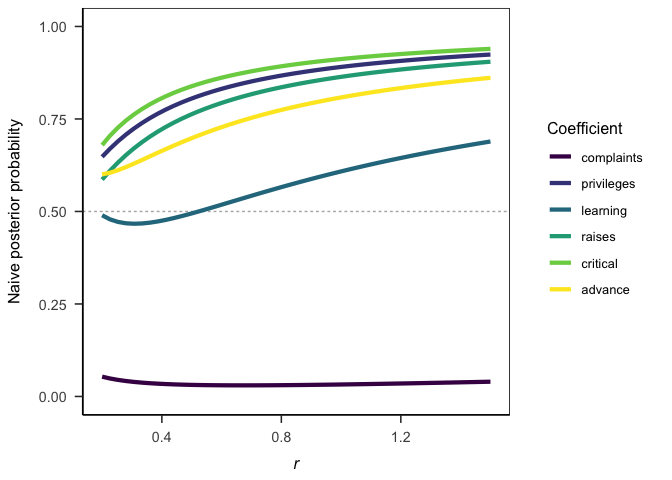
#> 0.0322969 4.1376723 0.9791980 2.3193670 3.6870978 1.8295123Similarly, performing a prior sensitivity analysis is straight forward and fast.
# Specify design
jab_sensitivity <- expand.grid(
coef = names(coef(attitude_lm))
, r = seq(0.2, 1.5, length.out = 50)
) |>
# Calculate Bayes factors for each prior setting
dplyr::group_by(r) |>
dplyr::mutate(
jab = jab(
attitude_lm
, prior = dcauchy
, location = 0
, scale = r
)
)
# Plot results
ggplot(jab_sensitivity) +
aes(x = r, y = jab / (1 + jab), color = coef) +
geom_hline(
yintercept = 0.5
, linetype = "22"
, color = grey(0.7)
) +
geom_line(linewidth = 1.5) +
scale_color_viridis_d() +
lims(y = c(0, 1)) +
labs(
x = bquote(italic(r))
, y = "Naive posterior probability"
, color = "Coefficient"
) +
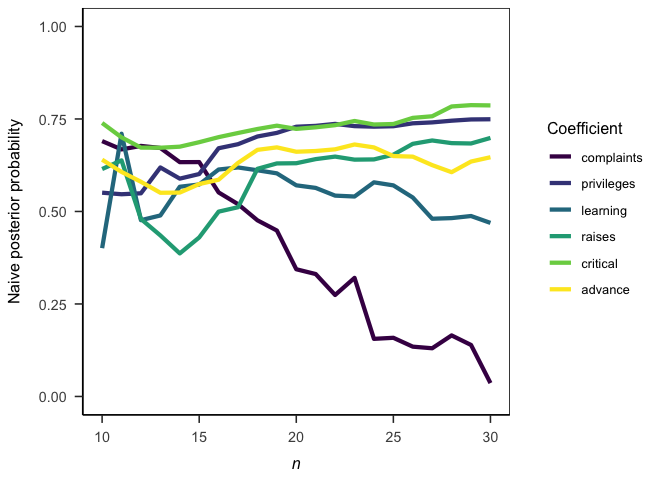
papaja::theme_apa(box = TRUE)Sequential analyses are also a breeze.
# Specify design
sequential_jab <- expand.grid(
coef = names(coef(attitude_lm))
, n = 10:nrow(attitude_z)
) |>
# Calculate Bayes factors for each subsample
dplyr::group_by(n) |>
dplyr::mutate(
jab = jab(
update(attitude_lm, data = attitude_z[1:unique(n), ])
, dcauchy
, location = 0
, scale = sqrt(2) / 4
)
, jab_pp = jab / (jab + 1)
)
# Plot results
ggplot(sequential_jab) +
aes(x = n, y = jab_pp, color = coef) +
geom_line(linewidth = 1.5) +
scale_color_viridis_d() +
lims(y = c(0, 1)) +
labs(
x = bquote(italic(n))
, y = "Naive posterior probability"
, color = "Coefficient"
) +
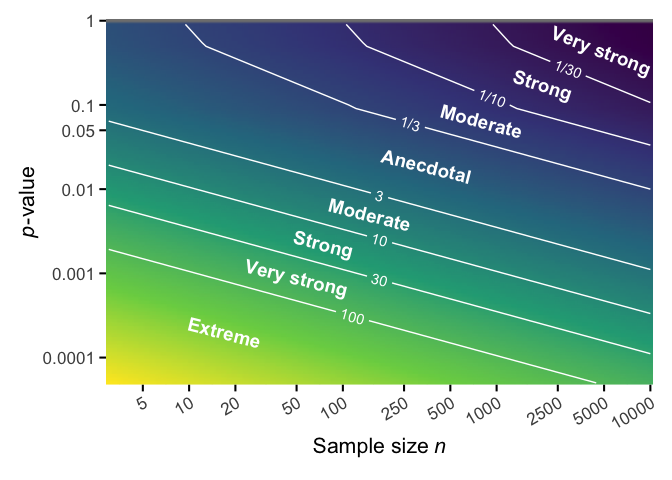
papaja::theme_apa(box = TRUE)By calculating JAB from p-values, we can explore approximately how much evidence a p-value provides for the alternative (or null) hypothesis for a given sample size. Here I use the precise piecewise approximation suggested by Wagenmakers (2022), Eq. 9. Note that both axes are on a log-scale.
library("geomtextpath")
p_boundaries <- c(0.0001, 0.001, 0.01, 0.05, 0.1, 1)
dat <- expand.grid(
p = exp(seq(log(0.00005), log(1), length.out = 100))
, n = exp(seq(log(3), log(10000), length.out = 100))
) |>
transform(jab_p = 1 / jab::jab_p(p, n))
evidence_labels <- data.frame(
n = c(17, 50, 75, 150, 350, 800, 2000, 4800)
, p = c(0.0002, 0.0009, 0.00225, 0.005, 0.019, 0.065, 0.175, 0.45)
, label = c("Extreme", "Very strong", "Strong", "Moderate", "Anecdotal", "Moderate", "Strong", "Very strong")
, angle = -c(17, 17, 17, 17, 17, 18, 21, 24) + 3
)
plot_settings <- list(
scale_x_continuous(
expand = expansion(0, 0)
, breaks = c(5, 10, 20, 50, 100, 250, 500, 1000, 2500, 5000, 10000)
, trans = "log"
, name = bquote("Sample size" ~ italic(n))
)
, scale_y_continuous(
expand = expansion(0, 0)
, breaks = p_boundaries
, labels = format(p_boundaries, scientific = FALSE, drop0trailing = TRUE)
, trans = "log"
, name = bquote(italic(p)*"-value")
)
, scale_fill_viridis_c(guide = "none")
, theme_minimal(base_size = 16)
, theme(
axis.ticks.length = unit(5, "pt")
, axis.ticks.x = element_line()
, axis.ticks.y = element_line()
, plot.margin = margin(0.5, 0.5, 0.5, 0.5, "cm")
, axis.title.x = element_text(margin = margin(t = 0.1, unit = "cm"))
, axis.title.y = element_text(margin = margin(r = -0, unit = "cm"))
, axis.text.x = element_text(angle = 30, hjust = 1)
)
)
to_reciprocal <- function(x) {
ifelse(
x > 1
, as.character(round(x))
, paste0("1/", round(1/x))
)
}
ggplot(dat) +
aes(x = n, y = p) +
geom_raster(aes(fill = log(jab_p)), interpolate = TRUE) +
# geom_hline(yintercept = p_boundaries, color = "white", alpha = 0.2) +
geom_textcontour(
aes(z = jab_p, label = to_reciprocal(after_stat(level)))
, color = "white"
, breaks = c(1/30, 1/10, 1/3, 3, 10, 30, 100)
) +
geom_text(
aes(x = n, y = p, label = label, angle = angle)
, data = evidence_labels
, color = "white"
, fontface = "bold"
, size = 5
) +
plot_settings
#> Warning: Removed 100 rows containing non-finite values (`stat_textcontour()`).