Development Envronment for the HUC12 sub watershed nutrient loading model.
- R version v3.6.3
- PostgreSQL v12.5
- PostGIS v3.0.2
Note: Use PGAdmin4 to interact directly with the Postgres database named
WRRIrunning onlocalhost:5430with the userpostgres
Install docker to run RStudio in the browser.
Copy wrri_pg_bak from Google Drive (WRRI_FallsJordan) into the top level of this git repository
docker-compose up -d
http://localhost:8787/
docker-compose restart -d
Add or remove R Libraries in install.R and then rebuild the docker container to see changes.
Add or remove linux packages in apt.txt then rebuild the docker container to see changes.
Run the following command from the root directory in the terminal
This command will create a new backup of the database saving any new data or views you added.
docker-compose exec postgis pg_dump -d WRRI -U postgres -w -f /etc/postgresql/wrri_pg_bak -F c -b
Now you need to copy the backup back to your machine using the following command.
docker cp nutrient-loading-model_postgis_1:/etc/postgresql/wrri_pg_bak .
Finally, update the backup in the shared Google Drive to share with others.
-
Install PGAdmin4 if you do not already have it.
-
Create a new server using by right clicking Server -> Create -> Server .
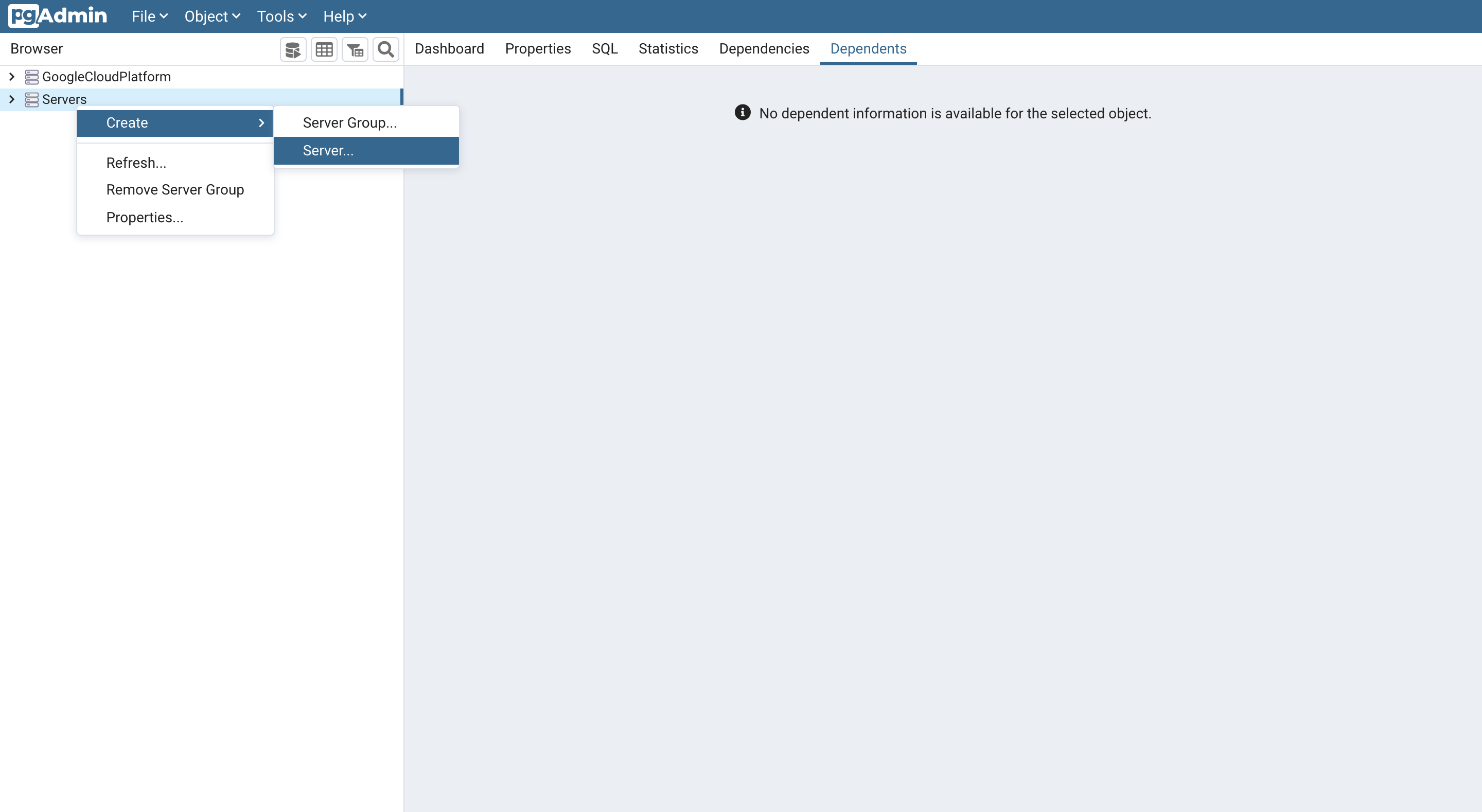
-
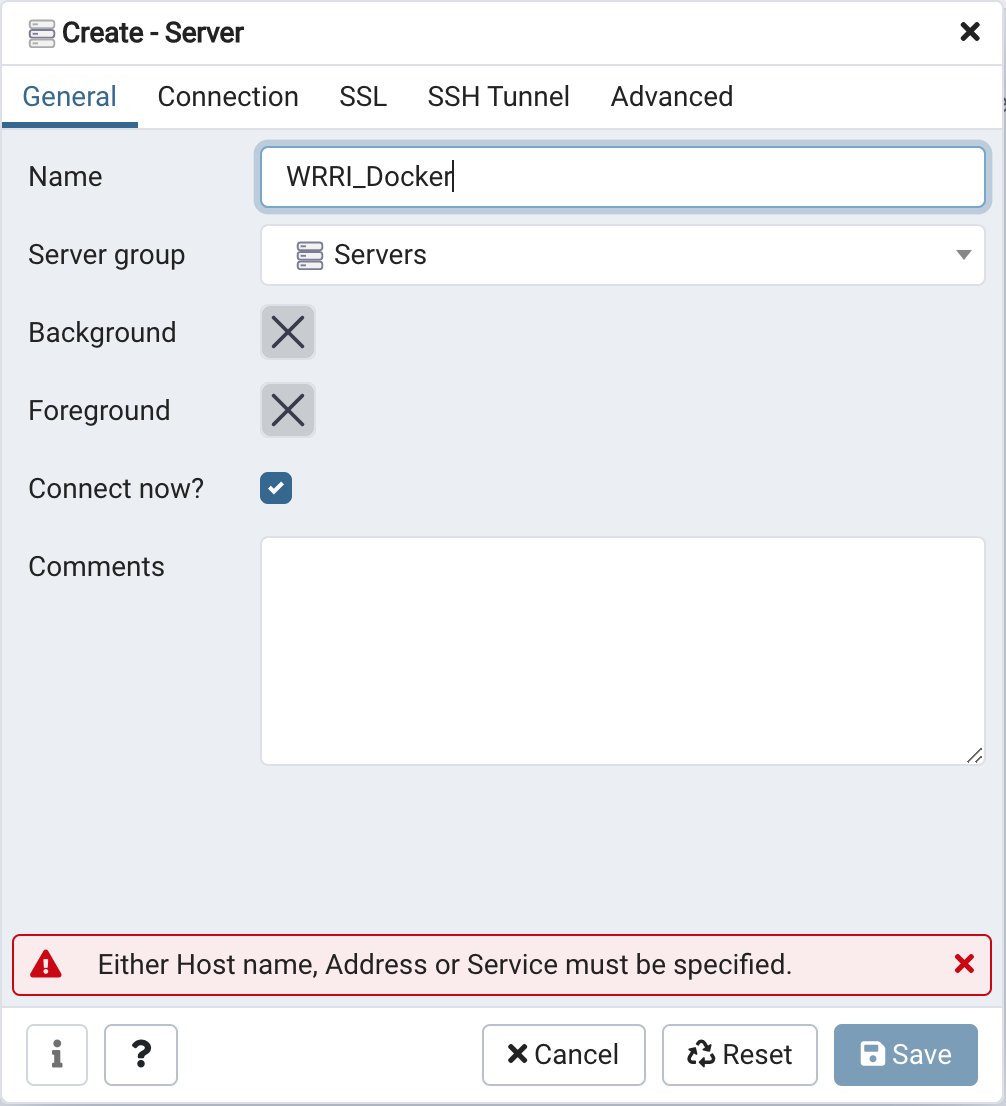
In the create server panel's General tab set the server name to
WRRI_Docker.
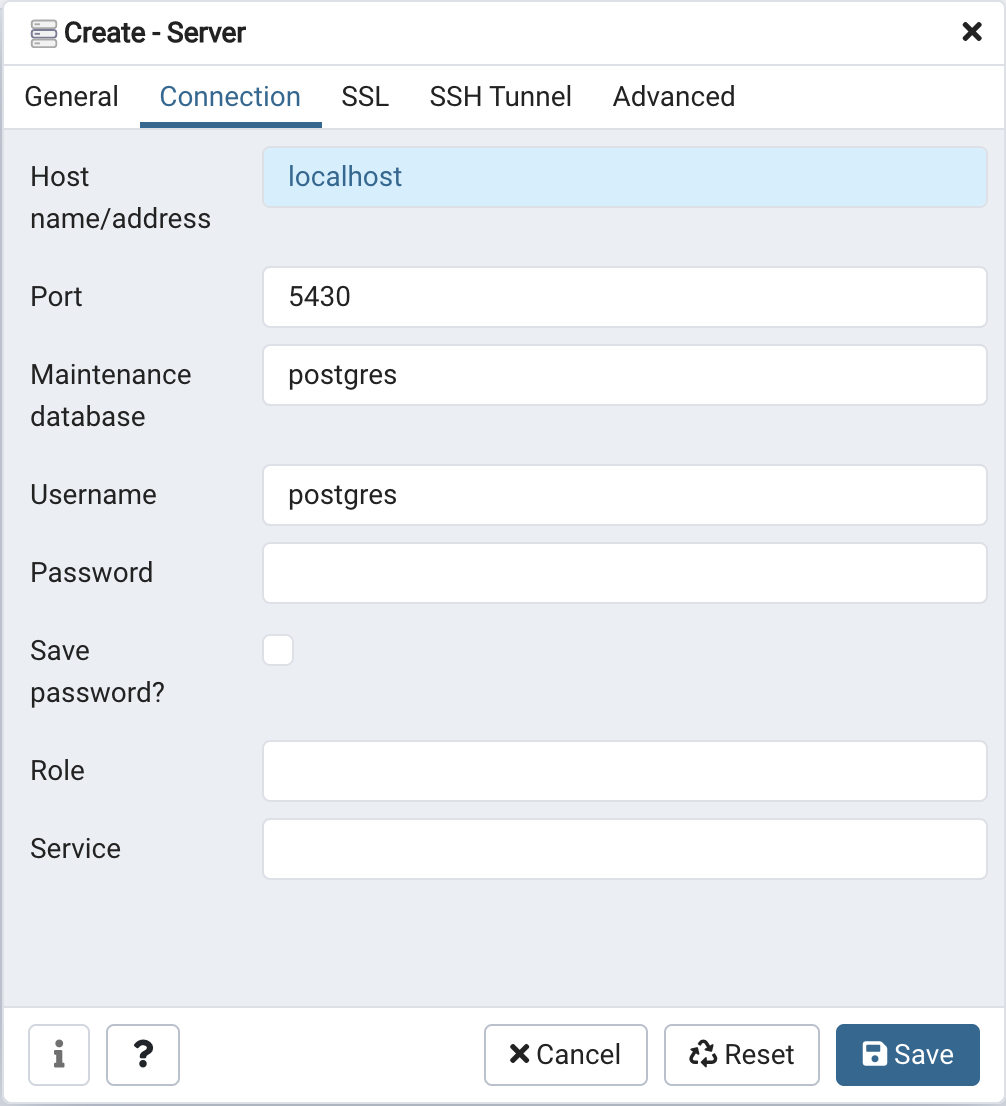
- Select the Connection tab from the create server panel. For the Host name/address enter
localhost, set Port to5430, set the Maintenance database topostgres, and set the username topostgres, leave the password empty.
- Click
saveat the bottom right of the Create - Server panel.
You are now connected to the database and will see your saved connection on the left panel of PGAdmin with the label WRRI_Docker.