Members:
- Trouvain, Mira
- Paluca, Denis
- Höbenreich, Jonas
- Link: https://gitlab.lrz.de/denis/psemoldyn_groupb
- Branch: master
- commit: 5515b42b
- log4cxx
-
Compile with
cmake -DDISABLE_DOXYGEN=ON -DCMAKE_BUILD_TYPE=Debug -G "CodeBlocks - Unix Makefiles" {PATH_TO_PROJECT} make MolSim -j <1.5 * number of cores> -
run main program with parameters specified in assignment:
./MolSim ./inputs/a4_task2_big.xml -
Compile on Cluster (Deactivate log4cxx)
cmake -DCMAKE_BUILD_TYPE=Release -DCLUSTER=ON -G "CodeBlocks - Unix Makefiles" {PATH_TO_PROJECT} module load xerces-c cmake --build . --target MolSim -- -j 20 -
Activate Parallelisation with -DOPENMP=ON
cmake -DCMAKE_BUILD_TYPE=Release -DOPENMP=ON -G "CodeBlocks - Unix Makefiles" {PATH_TO_PROJECT} cmake --build . --target MolSim -- -j 20
./MolSim <PATH_TO_INPUT_FILE>
or for performance testing:
./MolSim <PATH_TO_INPUT_FILE> -pt
-
Compile with
cmake -DDISABLE_DOXYGEN=ON -DCMAKE_BUILD_TYPE=Debug -G "CodeBlocks - Unix Makefiles" {PATH_TO_PROJECT} make Tests_run -j <1.5 * number of cores> -
run
ctestor
cd tests/ ./Tests_run
-
Build with
cmake -DCMAKE_BUILD_TYPE=Debug -G "CodeBlocks - Unix Makefiles" {PATH_TO_PROJECT} make doc_doxygen
see XSD file
-
Additional Membrane Simulations:
*The OS/hardware the measurements were made on Linux Cluster (COOL-MUC 2)
*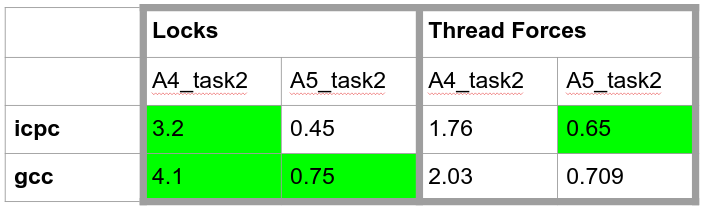
- Compiler used: gcc 9.3.0
- CMake version: 3.16.3
- Google Tests version: 1.10.0
- modules used on cluster:
- admin/1.0 4) spack/staging/20.2.2 7) intel-mpi/2019-intel
- tempdir/1.0 5) intel/19.0.5 8) xerces-c/3.2.1
- lrz/1.0 6) intel-mkl/2019 9) gcc/8.4.0