miloR
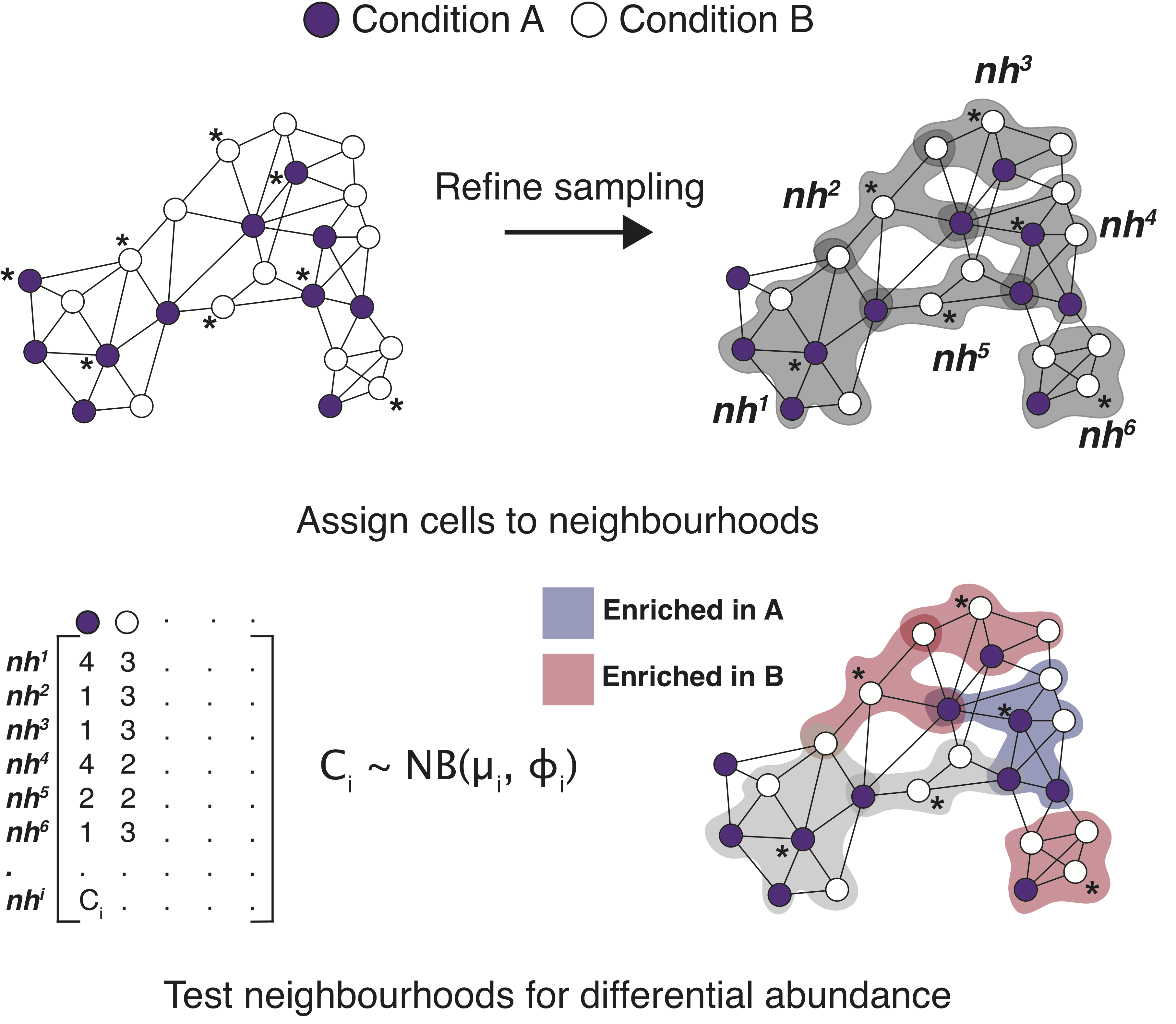
Milo is a method for differential abundance analysis on KNN graph from single-cell datasets. For more details, read our preprint.
Installation
## Milo is available from Bioconductor (preferred stable installation)
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("miloR")
## Install development version
devtools::install_github("MarioniLab/miloR", ref="devel")
Tutorials
- Basic Milo example on simulated dataset
- Milo example on mouse gastrulation dataset: this includes a demo for downstream analysis functions.
- Integrating Milo in scanpy/anndata workflow
Example work flow
An example of the Milo work flow to get started:
data(sim_trajectory)
milo.meta <- sim_trajectory$meta
milo.obj <- Milo(sim_trajectory$SCE)
milo.obj
Build a graph and neighbourhoods.
milo.obj <- buildGraph(milo.obj, k=20, d=30)
milo.obj <- makeNhoods(milo.obj, k=20, d=30, refined=TRUE, prop=0.2)
Calculate distances, count cells according to an experimental design and perform DA testing.
milo.obj <- calcNhoodDistance(milo.obj, d=30)
milo.obj <- countCells(milo.obj, samples="Sample", meta.data=milo.meta)
milo.design <- as.data.frame(xtabs(~ Condition + Sample, data=milo.meta))
milo.design <- milo.design[milo.design$Freq > 0, ]
milo.res <- testNhoods(milo.obj, design=~Condition, design.df=milo.design)
head(milo.res)
Support
For any question, feature request or bug report please create a new issue in this repository.
Contributions
We welcome contributions and suggestions from the community (though we may not take them onboard if they don't align with our development roadmap - please don't be offended). Please submit the initial idea as an issue, which we will discuss and ask for refinements/clarifications. If we approve the idea, then please open a pull request onto the devel branch, from which we will begin a review process. To smooth the process, please note that code changes must be backwards compatible, and must include all relevant unit tests.