sars2pack
The sars2pack R package includes data resources, workflows, and data science tools to understand and interpret the COVID-19 pandemic. Access to data resources is “real-time” to get the most up-to-date information. Use cases and introductory material are available in vignettes and in documentation.
Contributions are welcome. Have an interesting analysis that you’d like to share, write up an R markdown document and contribute it as a vignette. If you are not used to working collaboratively in github, just post a new issue to ask for help.
Thanks to the armies of people providing data for the rest of the world, often on a volunteer basis. Without their tireless work, we would not have these rapidly-developing resources.
Install
BioManager::install('seandavi/sars2pack')
# OR
devtools::install_github('seandavi/sars2pack')
Features
Data resources available
- Johns Hopkins global pandemic time series
- USAFacts county-level US epidemic data
- NYTimes state and county-level US epidemic data
- COVIDTracker.com US state- and county-level epidemic data that includes detailed positive, negative, and pending test data
- US Healthcare Capacity dataset, including details on US hospital capacity and capabilities.
- US County-level metadata, including geospatial data
Capabilities
- Access data from multiple sources: jhu_data(), usa_facts_data(), nytimes_county_data(), nytimes_state_data(), covidtracker_data(), etc.
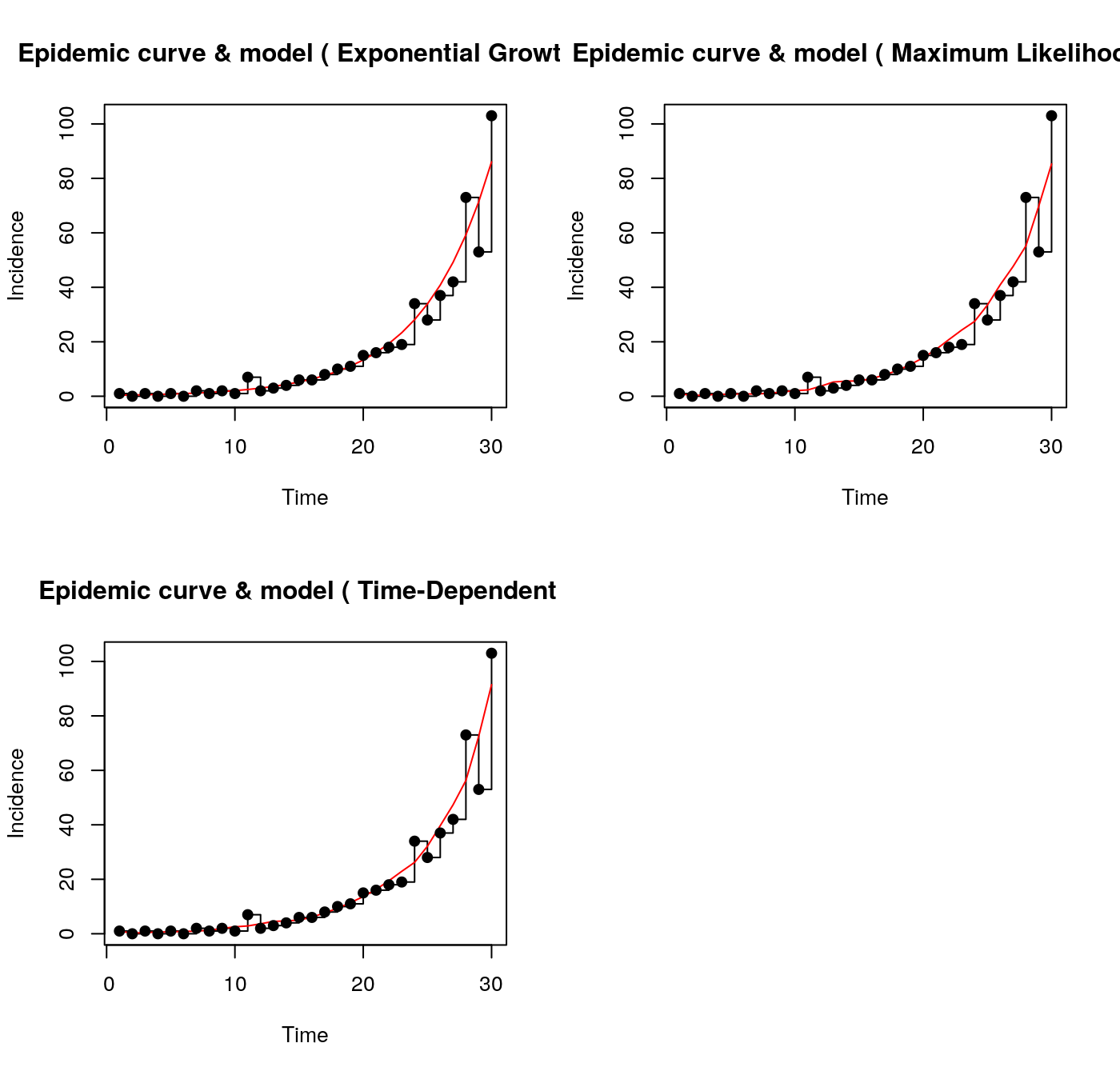
- Estimate R0 for localities or countries.
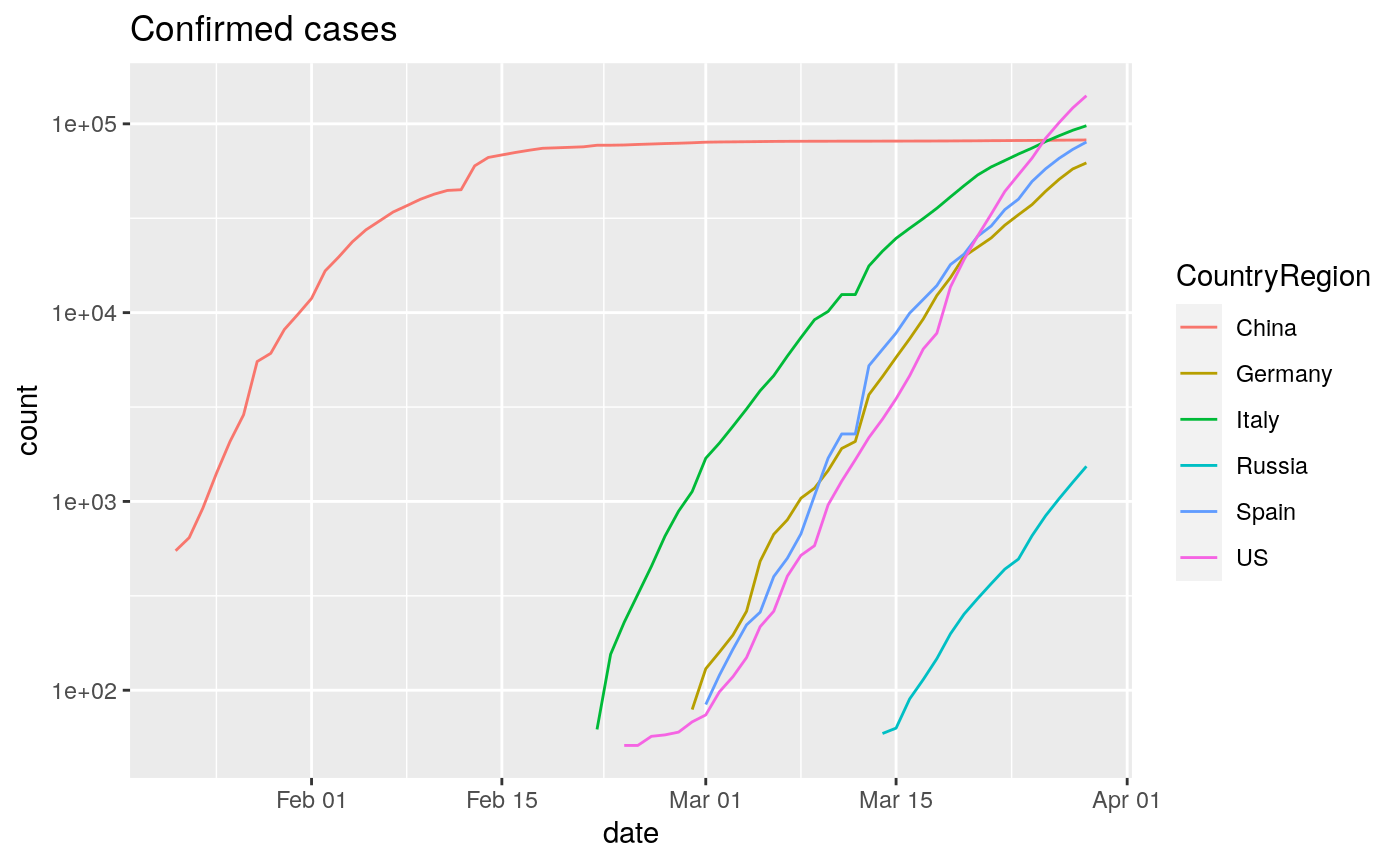
- Visualize time series pandemic case growth
- Perform data mashups between COVID-19 pandemic data and additional geographic, financial, and demographic datasets
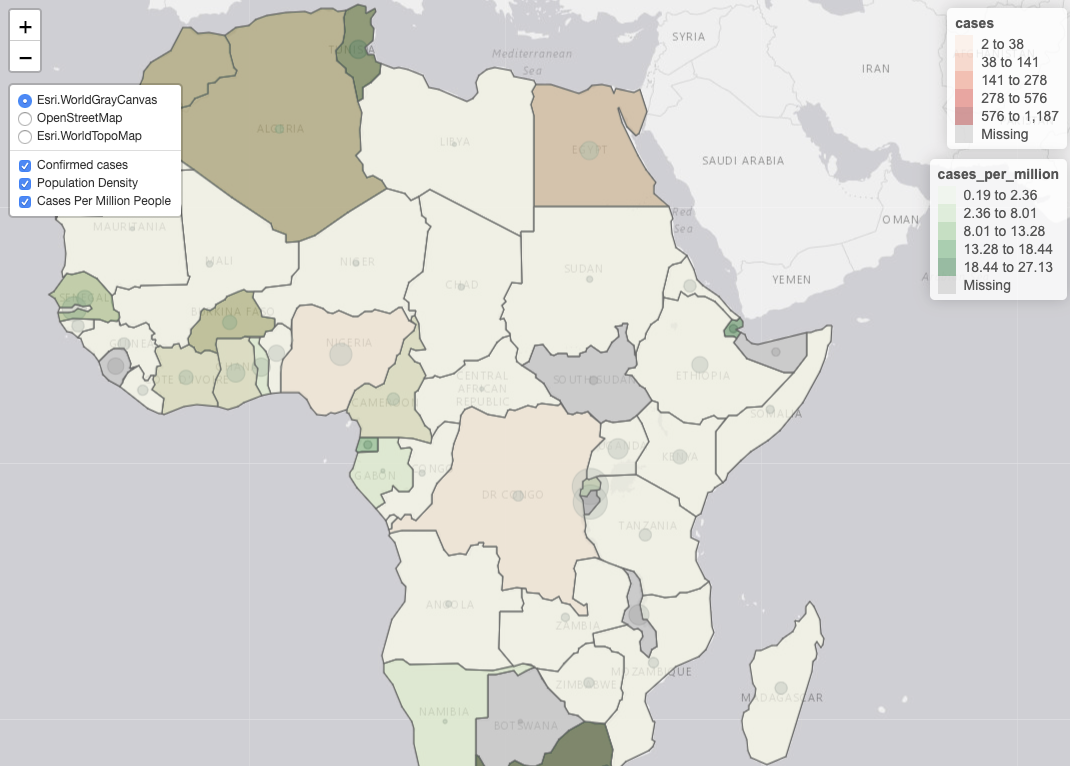
- Create static and interactive maps of COVID-19 data for states, countries, or even counties.
Visualization
Workflow status
Automated workflows and current status.
| Workflow Status | Description |
|---|---|
| Produces regularly updated data resources and products | |
| Prepare and publish pkgdown documentation |
Contribute
To contribute to this package please make a fork and then issue pull requests.