The official code implementation for DREAMwalk from our paper, Biomedical knowledge graph learning for drug repurposing by extending guilt-by-association to multiple layers.
We also provide codes for calculating semantic similarities of entities given hierarchy data, along with example heterogeneous network file and notebook file for generating node embedding and predicting associations.
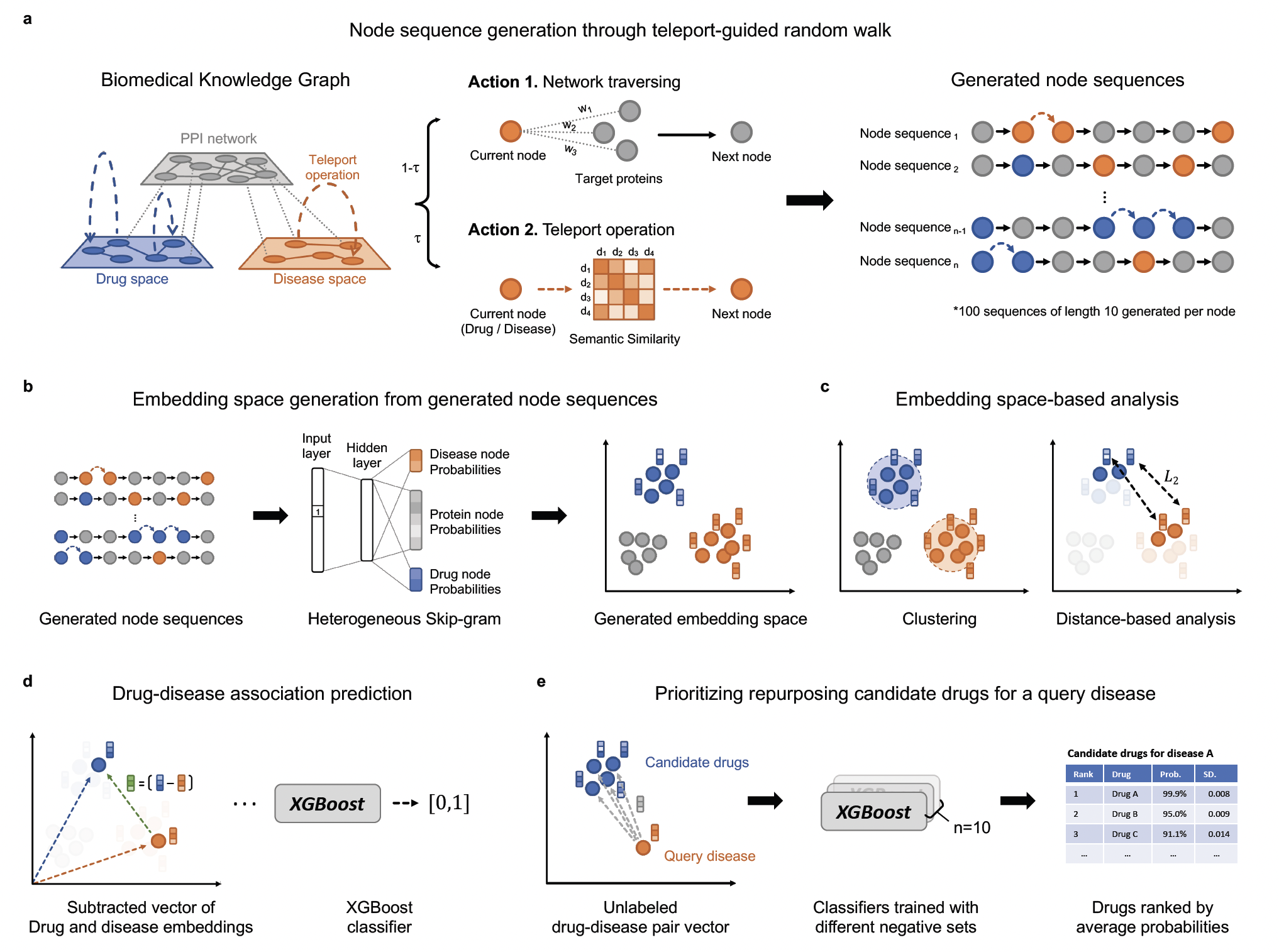
The full model architecture is provided below. DREAMwalk's drug-disease association prediction pipeline is consisted of three steps;
Step 1. Create semantic similarity network from semantic hierarchies
Step 2. Node embedding generation through teleport-guided random walk
Step 3. Drug-disease association prediction (or prediction of any other links)
First, clone this repository and move to the directory.
git clone https://github.com/eugenebang/DREAMwalk.git
To install the appropriate environment for DREAMwalk, you should install conda package manager.
After installing conda and placing the conda executable in PATH, the following command will create conda environment named dreamwalk. It will take up to 10 minutes to setup the environment, but may vary upon the Internet connection and package cache states.
conda env create -f environment.yaml && \
conda activate dreamwalk
To check whether DREAMwalk works properly, please refer to the Example codes section below.
Sample code to generate the embedding space and predict drug-disease associations are provided in run_demo.ipynb.
- The file formats for each input file can be found in here.
- Detailed instructions in running the codes can be found here.
Operating system
DREAMwalk training and evaluation were tested for Linux (Ubuntu 18.04) operating systems.
Prerequisites
DREAMwalk training and evaluation were tested for the following python packages and versions.
- For embedding generation
python= 3.8networkx= 2.8.8numpy=1.23.3pandas=1.4.4scikit-learn=1.1.3scipy=1.9.1tqdm=4.64.1parmap=1.6.0xgboost=1.7.4
Bang, D., Lim, S., Lee, S., & Kim, S. Biomedical knowledge graph learning for drug repurposing by extending guilt-by-association to multiple layers. Nature Communications (2023)
@article{bang2023biomedical,
title={Biomedical knowledge graph learning for drug repurposing by extending guilt-by-association to multiple layers},
author={Bang, Dongmin and Lim, Sangsoo and Lee, Sangseon and Kim, Sun},
journal={Nature Communications},
volume={14},
number={1},
pages={3570},
year={2023},
publisher={Nature Publishing Group UK London}
}