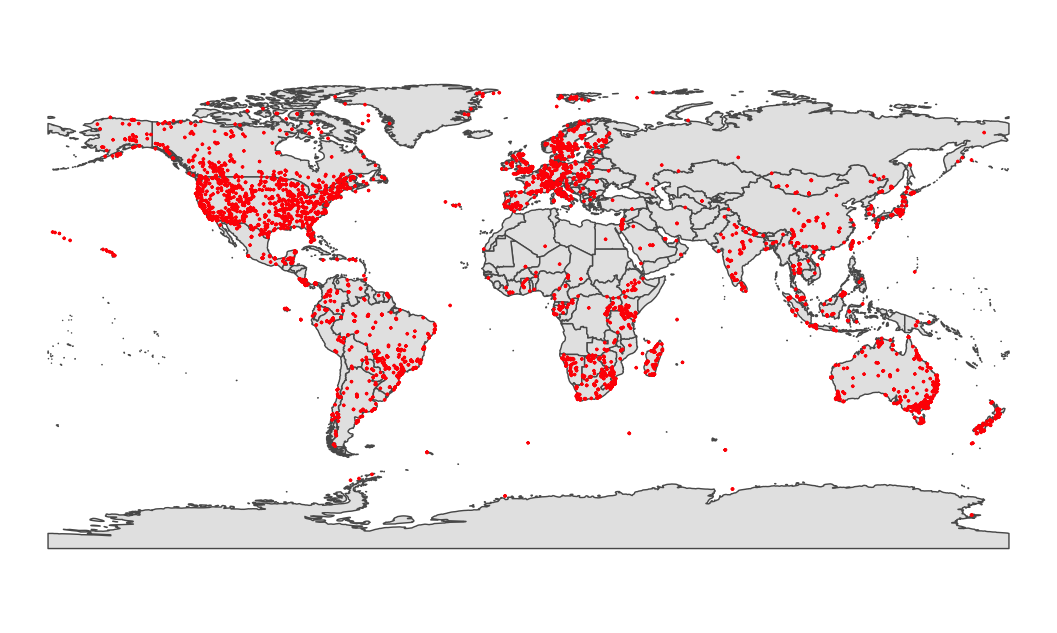
The HomeRange data contains mammal home-ranges estimates, species names, methodological information on data collection, home-range estimation method, period of data collection, study coordinates and name of location, as well as species traits derived from the studies, such as body mass, life stage, reproductive status and locomotor habit. The data paper describes the contents of HomeRange in more detail:
Broekman, M.J.E., Hoeks, S., Freriks, R., Langendoen, M.M., Runge, K.M., Savenco, E., ter Harmsel, R., Huijbregts, M.A.J. & Tucker, M.A. (2022). HomeRange: A global database of mammalian home ranges. Global Ecology and Biogeography. https://doi.org/10.1111/geb.13625.
- The latest version of the HomeRange data is available from this repository (see below).
- The HomeRange data is archived on Dryad: https://doi.org/10.5061/dryad.d2547d85x.
- An interactive map of the HomeRange data can be accessed here.
-
HomeRange data: the latest HomeRange data (formatted as a CSV) are contained within the
HomeRangeData_2023_11_28_1.zip, which can be downloaded using this link -
Metadata: the metadata PDF for the HomeRage data can be accessed here.
-
Reference list: all references for the home-range values contained in the HomeRange dataset can be found in the
HomeRangeReferences_2023_11_28.csvfile included in theHomeRangeData_2023_11_28_1.zipavailable from this repository (link).
-
HomeRange data: the R package can be used to download and import the HomeRange data all from within R using a single function call
GetHomeRangeData()(see example code below). -
Metadata: the metadata can be viewed from the R package using
ViewMetaData()orvignette(package="HomeRange","Metadata")(see example code below). -
Reference list: by setting the
IncludeReferencesagrument toTRUEin theGetHomeRangeData()function of the R package (GetHomeRangeData(IncludeReferences = TRUE)) all references are downloaded and merged with the HomeRange dataset directly.
# install the HomeRange R package
install.packages("https://github.com/SHoeks/HomeRange/raw/main/HomeRange_1.05.tar.gz",
repos=NULL,
method="libcurl")
# alternatively, install the HomeRange R package using install_github:
# remotes::install_github("SHoeks/HomeRange", subdir='pkg')
# load package into R
library('HomeRange')
# package information
?HomeRange
# view HomeRange metadata directly as PDF in the browser
ViewMetaData()
# Or access the metadata from the HomeRange vignettes
vignette(package="HomeRange","Metadata")
# get the dataset, this function automatically downloads and imports the data
HomeRangeData <- GetHomeRangeData() # by default IncludeReferences is set to FALSE
# get data with the references attached
HomeRangeDataWithRefs <- GetHomeRangeData(IncludeReferences = TRUE)
# some information on the HomeRange data
head(HomeRangeData)
head(HomeRangeDataWithRefs)
summary(HomeRangeData)
str(HomeRangeData)# plotting data
PlotMap(HomeRangeData)
PlotHistogram(HomeRangeData)# get more information
MakeStatTable(HomeRangeData)# match with the COMBINE imputed dataset
# https://esajournals.onlinelibrary.wiley.com/doi/10.1002/ecy.3344
COMBINE <- read.csv("/path/to/combine/trait_data_imputed.csv")
merged_data = MergeWithCOMBINE(HomeRangeData, COMBINE)
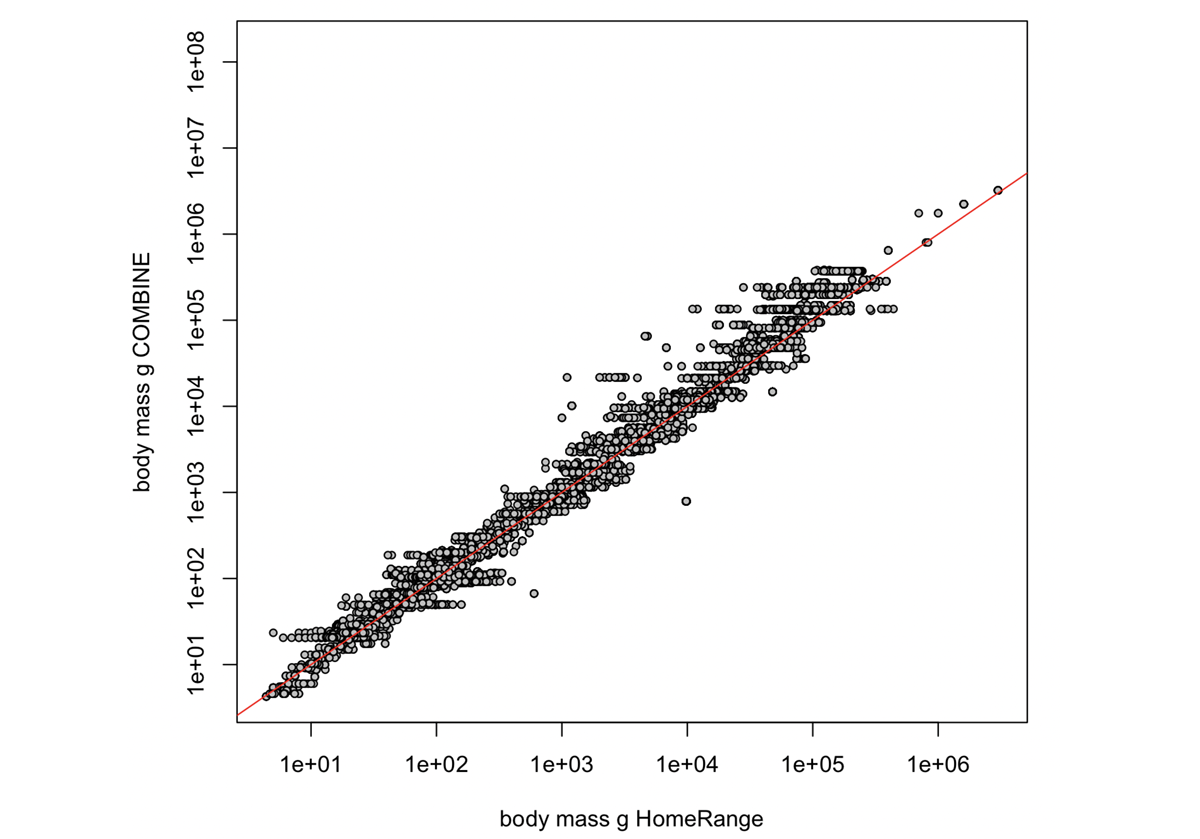
# example plot of the merged data
plot(merged_data$Body_mass_kg*1000,
merged_data$COMBINE_adult_mass_g,
log = "xy", pch=21,
cex=0.7, bg="grey",
xlim=c(10^0,10^7), ylim=c(10^0,10^7),
xlab="body mass g HomeRange",
ylab="body mass g COMBINE")
abline(0,1,col="red")