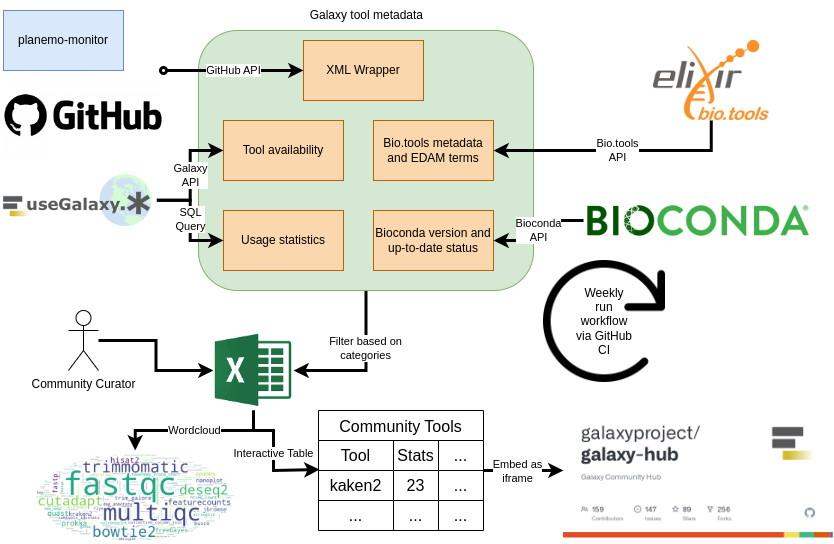
Galaxy Tool Metadata Extractor
What is the tool doing?
This tool automatically collects a table of all available Galaxy tools including their metadata. The created table can be filtered to only show the tools relevant for a specific community. Learn how to add your community.
The tools performs the following steps:
- Parse tool GitHub repository from Planemo monitor listed
- Check in each repo, their
.shed.yamlfile and filter for categories, such as metagenomics - Extract metadata from the
.shed.yaml - Extract the requirements in the macros or xml to get version supported in Galaxy
- Check available against conda version
- Extract bio.tools information if available in the macros or xml
- Check available on the 3 main galaxy instances (usegalaxy.eu, usegalaxy.org, usegalaxy.org.au)
- Get usage statistics form usegalaxy.eu
- Creates an interactive table for all tools: All tools
- Creates an interactive table for all registered communities, e.g. microGalaxy
Usage
Prepare environment
-
Install virtualenv (if not already there)
$ python3 -m pip install --user virtualenv -
Create virtual environment
$ python3 -m venv env -
Activate virtual environment
$ source env/bin/activate -
Install requirements
$ python3 -m pip install -r requirements.txt
Extract all tools
-
Get an API key (personal token) for GitHub
-
Export the GitHub API key as an environment variable:
$ export GITHUB_API_KEY=<your GitHub API key> -
Run the script
$ python bin/extract_all_tools.sh
The script will generate a TSV file with each tool found in the list of GitHub repositories and metadata for these tools:
- Galaxy wrapper id
- Description
- bio.tool id
- bio.tool name
- bio.tool description
- EDAM operation
- EDAM topic
- Status
- Source
- ToolShed categories
- ToolShed id
- Galaxy wrapper owner
- Galaxy wrapper source
- Galaxy wrapper version
- Conda id
- Conda version
Filter tools based on their categories in the ToolShed
-
Run the extraction as explained before
-
(Optional) Create a text file with ToolShed categories for which tools need to be extracted: 1 ToolShed category per row (example for microbial data analysis)
-
(Optional) Create a text file with list of tools to exclude: 1 tool id per row (example for microbial data analysis)
-
(Optional) Create a text file with list of tools to really keep (already reviewed): 1 tool id per row (example for microbial data analysis)
-
Run the tool extractor script
$ python bin/extract_galaxy_tools.py \ --tools <Path to CSV file with all extracted tools> \ --filtered_tools <Path to output CSV file with filtered tools> \ [--categories <Path to ToolShed category file>] \ [--excluded <Path to excluded tool file category file>]\ [--keep <Path to to-keep tool file category file>]
Add your community
In order to add your community you need to:
- Fork this repository.
- Add a folder for your community in
data/communities. - Add at least the file
categories. - Add all
categoriesthat are relevant to initially filter the tools for your community. Possible categories are listed here Galaxy toolshed. - Make a pull request to add your community.
- The workflow will run every sunday, so on the next monday, your community table should be added to
results/<your community name>