Hi, Welcome to the NeuroimaGene project!
The goal of this project is to improve the interpretability of genetic studies relating to neurological and/or psychiatric traits. Analyses such as genome-wide association studies often highlight genetic variants that segregate with disease. Methods such as fine mapping, eQTL colocalization, and transcriptome-wide analyses (TWAS) have been developed to identify the gene level targets of these trait-associated variants. Once a set of gene targets has been identified from GWAS findings, it often remains challenging to identify the biological consequences of variation in these genes. Briefly, the NeuroimaGene R package makes it possible to identify the neuroimaging correlates of TWAS associations.
More specifically, NeuroimaGene can be used to (1) characterize individual genes or gene sets as relevant to the structure and function of the brain, (2) identify the region(s) of the brain or body in which expression of target gene(s) is neurologically relevant, (3) impute the brain features most impacted by user-defined gene sets such as those produced by cohort level gene association studies, and (4) generate publication level, modifiable visual plots of neuroimaGene associations.
We recommend installing the NeuroimaGene package directly from CRAN. Use the following script in the R console:
install.packages("neuroimaGene")You can also install the NeuroimaGene package directly from GitHub using the devtools package. First, ensure you have devtools installed. Then, use the install_github function to install the package from the GitHub repository.
-
Install
devtools:install.packages("devtools") -
Install
NeuroimaGenefrom GitHub: Use theinstall_githubfunction from thedevtoolspackage to installNeuroimaGene:devtools::install_github("xbledsoe/NeuroimaGene_R")
-
Load the Package: Once installed, you can load the
NeuroimaGenepackage as follows:library(neuroimaGene)
Here is a complete example in R:
# Install devtools if not already installed
install.packages("devtools")
# Install the NeuroimaGene package from GitHub
devtools::install_github("xbledsoe/NeuroimaGene_R")
# Load the package
library(neuroimaGene)The NeuroimaGene R package serves to identify, characterize, and visualize the neurological correlates of genetically regulated gene expression (GReX). The following functions serve this goal.
The main package function, neuroimaGene() takes a vector of gene names
(HUGO gene names or ensembl id’s) and returns a data table of GReX-NIDP
associations that fit a number of user-defined parameters described
below:
ng <- neuroimaGene( gene_list, modality='T1', atlas='Desikan', mtc='BH', nidps = NA, filename = NA)
gene_list: a vector of gene names (HUGO gene names or ensembl id’s). Typically these are associated with a single trait of interest.
modality: the MRI modality used to define the NIDP in the initial UK Biobank imaging protocol. (See table below for options)
atlas: the parcellation atlas used to define the NIDPs from the MRI. We use the term “atlas” loosely recognizing that the fMRI data, dMRI data, and T1/T2 data are parcellated in diverse ways. (See table below for options)
mtc: Multiple Testing Correction. Options include Nominal (nom), Benjamini Hochberg (BH), or Bonferroni (BF)
nidps: A user-defined list of NIDPs to test. This option overrides the modality and atlas parameters. Default is NA.
filename: A user-defined pathname/filename for text output of the NeuroimaGene results.
Extended Parameter Descriptions
Modality and Atlas parameters
In the process of using the tool, the user is responsible for selecting
a subset of NIDPs from the resource for analysis using the modality
and atlas parameters. These parameters allow the user to target
specific types of NIDPs such as hippocampal subfields, area and
thickness of named cortical regions, fractional anisotropy of named
white matter tracts etc. It is recommended to identify the type of brain
measure one is interested in prior to performing the gene set analysis.
The input options for the modality and atlas parameters are detailed
in the dropdown table below
Multiple Testing Correction Parameter for statistical significance
In addition to selecting the atlas and modality, neuroimaGene requires a multiple testing threshold correction. Each imaging modality contains a different number of NIDPs (see table above). The Bonferroni correction (‘BF’) treats each of these NIDPs as independent even though we know through significant data analyses that this is not accurate. This is a highly conservative threshold that will yield high confidence associations but is likely to generate many false negatives. Recognizing the correlation of brain measures from the same modality and atlas, we recommend using the less stringent Benjamini Hochberg (‘BH’) False Discovery Rate for discovery analyses. The nominal option (‘nom’) will provide the uncorrected p-value.
Note that the mtc parameter represents a study-wide is dynamic,
depending on the modality and atlas parameters provided in the initial
neuroimaGene query.
-
If a user provides an atlas, multiple testing correction will be calculated at an atlas-wide level.
-
If the user provides a modality but sets the atlas as NA or ‘all’, multiple testing correction will be calculated at a modality-wide level.
-
If the user sets the both the modality and atlas to NA or ‘all’, the correction will be applied to the entire data set.
The above bullet points apply to both the Bonferroni (BF) and Benjamini Hochberg (BH) corrections. Alpha is set to 0.05 in either case.
Providing a user selected set of NIDP names
If the user has a specific set of NIDPs that they would like to assess,
the nidps parameter receives a vector of NIDPs by name. The NIDP names
all include data about the modality, atlas, region, and hemisphere. The
names have to be provided in character form and matching must be
perfect. To aid with the user-provision of NIDPs, we include the helper
utility listNIDPs(). This function takes as parameters modality and
atlas and returns a list of all NIDP names that satisfy these
criteria. The user can then manually curate a list of desired NIDPs.
Note that when providing a user-defined set of NIDPs, this option overrides the modality and atlas parameters. This applies both to the set of NIDPs analyze as well as the multiple testing correction. The nominal p-value will be reported as both the Bonferroni and Benjamini-Hochberg corrections that are encoded rely on knowing the set of NIDPs being tested. The user NIDP input violates this assumption and requires the user to identify their own multiple testing threshold.
See dropdown table for NIDP descriptions and source links
NIDP atlas descriptions and source links
| Modality | Atlas name | # of NIDPs | Description | source |
|---|---|---|---|---|
| T1 | all | 1319 | All measures recorded by the UKB neuroimaging study derived from T1 imaging | see note* |
| T1 | Destrieux | 444 | Destrieux atlas parcellation of cortical sulci and gyri | Destrieux |
| T1 | AmygNuclei | 20 | morphology of Nuclei of the amygdala | Amygdala nuclei |
| T1 | Subcortex | 52 | subcortical volumetric segmentation | aseg_volume |
| T1 | Broadmann | 84 | cortical morphology via Broadmann Areas | Broadmann |
| T1 | Desikan | 202 | Desikan Killiany atlas parcellation of cortical morphology | Desikan |
| T1 | DKT | 186 | DKT atlas parcellation of cortical morphology | DKTatlas |
| T1 | FAST | 139 | cortical morphology via FMRIB’s Automatic Segmentation Tool | FAST |
| T1 | FIRST | 15 | Subcortical morphology via FIRST | FIRST |
| T1 | HippSubfield | 44 | morphology of Hippocampal subfields | HippSubfield |
| T1 | pial | 66 | structure: Desikan Killiany atlas of the pial surface | Desikan |
| T1 | Brainstem | 5 | structure: Freesurfer brainstem parcellation | Brainstem |
| T1 | SIENAX | 10 | structure: Structural Image Evaluation of whole brain measures | SIENAX |
| T1 | ThalamNuclei | 52 | morphology of the Nuclei of the thalamus | ThalamNuclei |
| dMRI | all | 675 | All measures recorded by the UKB neuroimaging study derived from DWI imaging | see note* |
| dMRI | ProbtrackX | 243 | white matter mapping obtained via probabilistic tractography | ProbtrackX* |
| dMRI | TBSS | 432 | white matter mapping obtained via tract-based spatial statistics | TBSS* |
| rfMRI | ICA100 | 1485 | functional connectivity using 100 cortical seeds | see note* |
| rfMRI | ICA25 | 210 | functional connectivity using 25 cortical seeds | see note* |
| rfMRI | ICA-features | 6 | summary of functional connectivity components | see note* |
| T2_FLAIR | BIANCA | 1 | white matter hyperintensity classification algorithm | BIANCA |
| T2star | SWI | 14 | susceptibility-weighted imaging: microhemorrhage and hemosiderin deposits | see note* |
* original publication for details here (Alfaro-Almagro, Fidel, et al. “Image processing and Quality Control for the first 10,000 brain imaging datasets from UK Biobank.” Neuroimage 166 (2018): 400-424.)
library(neuroimaGene, quietly = TRUE)
gene_list <- c('TRIM35', 'PROSER3', 'EXOSC6', 'PICK1', 'UPK1A', 'ESPNL', 'ZIC4')
ng <- neuroimaGene(gene_list, atlas = NA, mtc = 'BH', vignette = TRUE)
| gene | gene_name | gwas_phenotype | training_model | zscore | mod_BHpval |
|---|---|---|---|---|---|
| ENSG00000223496 | EXOSC6 | aparc-a2009s_lh_thickness_G-front-middle | JTI_Adrenal_Gland | 4.160209 | 0.0357891 |
| ENSG00000223496 | EXOSC6 | aparc-a2009s_lh_thickness_G-front-sup | JTI_Adrenal_Gland | 5.163521 | 0.0012963 |
| ENSG00000223496 | EXOSC6 | aparc-a2009s_lh_thickness_G-parietal-sup | JTI_Adrenal_Gland | 5.099214 | 0.0016672 |
| ENSG00000144488 | ESPNL | aparc-a2009s_lh_thickness_G-parietal-sup | JTI_Adrenal_Gland | -4.306351 | 0.0241802 |
| ENSG00000144488 | ESPNL | aparc-a2009s_lh_thickness_G-pariet-inf-Angular | JTI_Adrenal_Gland | -6.103620 | 0.0000168 |
| ENSG00000223496 | EXOSC6 | aparc-a2009s_lh_thickness_G-pariet-inf-Angular | JTI_Adrenal_Gland | 4.362468 | 0.0205838 |
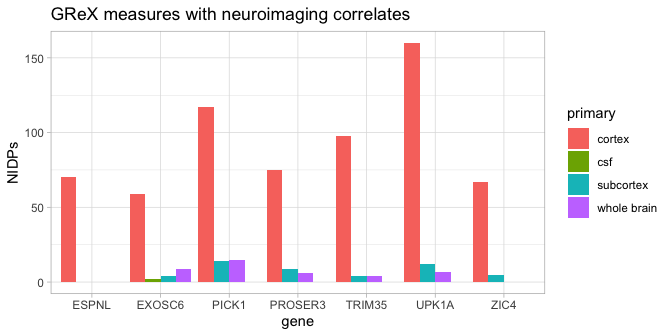
Users may wish to know the comparative contribution of each gene from the provided list to the NeuroimaGene results. We include a function to visualize the number of NIDPs per gene in the neuroimaGene package.
plot_gns(ng, maxGns = 15)
The legend describes the primary measurement of the NIDPs For structural
data, this is the cortex, subcortex, CSF, or whole brain measurements.
The parameter maxGns describes the maximum number of genes to display.
It is set to a default of 15. Genes are selected by ranking the
neuroimagene genes by their zscore magnitudes and selecting the top N
genes.
Note that this plot does not reflect any information on the number of
tissue contexts in which the GReX-NIDP association is significant. This
data is better captured by the plot_gnNIDP() function.
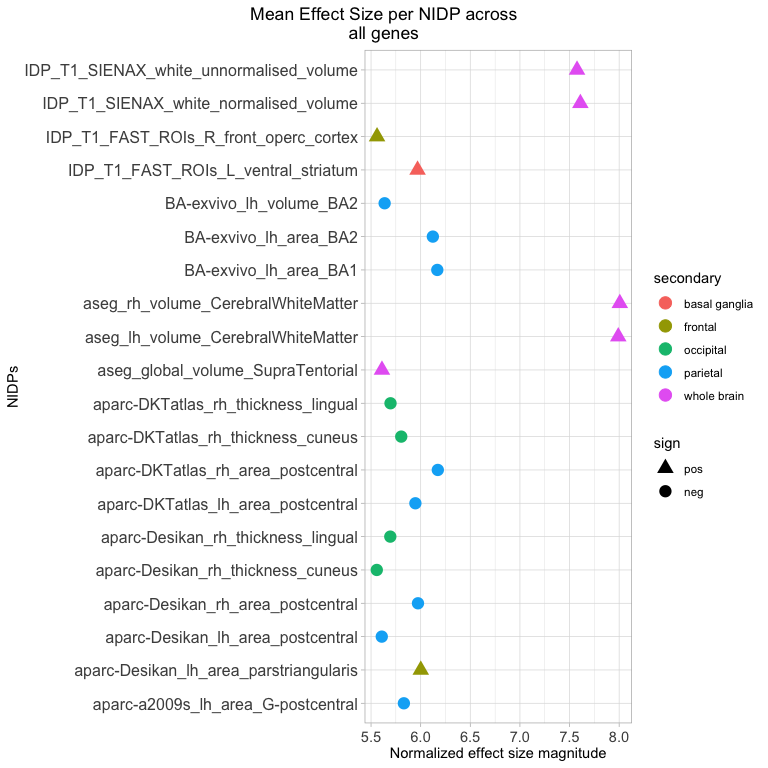
It is also useful to see what NIDPs are most impacted by the genes
provided. The plot_nidps() function calculates the mean zscore for
each NIDP across all tissue models and genes. It amounts to an
aggregated effect of the genes on the region of interest. This linear
aggregation is an approximation best understood to show convergence in
effect size magnitude and direction.
plot_nidps(ng, maxNidps = 20)
The legend describes a narrower descriptive profile of the NIDPs than
the plot_gns() tool. The parameter maxNidps describes the maximum
number of NIDPs to display. It is set to a default of 30. NIDPs are
selected by ranking the NIDPs by their zscore magnitudes and selecting
the top N NIDPs.
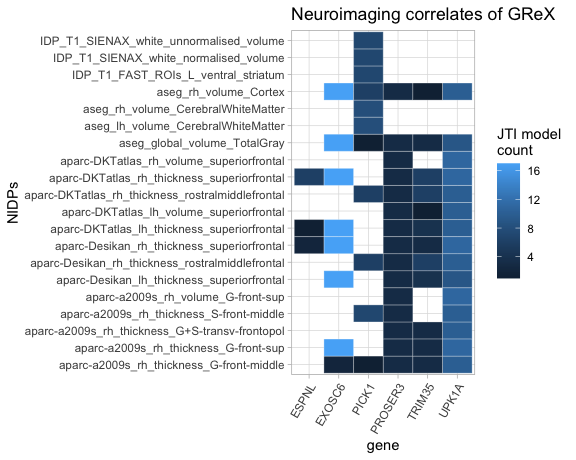
It is also useful to see the intersection of genes and NIDPs. The
plot_gnNIDP() function shows the number of JTI models in which each
GReX-NIDP association is significant.
plot_gnNIDP(ng, maxNidps = 20, maxGns = 15)
The parameter maxNidps describes the maximum number of NIDPs to
display. It is set to a default of 20. NIDPs are selected by ranking the
NIDPs by their zscore magnitudes and selecting the top N NIDPs. The
parameter maxGns describes the maximum number of genes to display. It
is set to a default of 15. Genes are selected by ranking the
neuroimagene genes by their zscore magnitudes and selecting the top N
genes.
It is often useful to provide a visual representation of neuroimaging
features in their 2-dimensional biological context for presentation
purposes. We include the neuro_vis() function for this purpose. This
function calculates the mean normalized effect size for each NIDP in the
neuroimaGene object. The user must provide a number of parameters listed
below.
ng_obj - NeuroimaGene object produced by neuroimaGene() function
atlas - desired atlas for visualization. (Desikan[default], Subcortex, DKT, Destrieux)
lowcol - color for low end of Zscore spectrum. Default is darkred
midcol - color for middle of Zscore spectrum. Default is white
hicol - color for top end of Zscore spectrum. Default is blue4
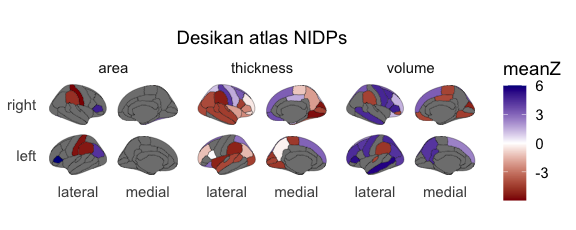
The function only reflects cortical and subcortical measures. It will
automatically separate the NIDPs according to volume, thickness, and
surface areas and plot each as its own facet. Please note that this
function uses the ggseg package and select atlases from the
ggsegExtra package. Please be sure to cite the creators of ggseg if
using this feature in publications.
Mowinckel AM, Vidal-Piñeiro D (2019). “Visualisation of Brain Statistics with R-packages ggseg and ggseg3d.” 1912.08200.
The neuro_vis() function can be used as described below:
neuro_vis(ng, atlas = 'Desikan', lowcol = 'darkred', midcol = 'white', highcol = 'blue4')
#> merging atlas and data by 'label', 'atlas', 'roi'
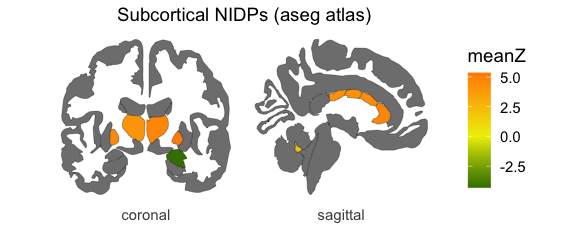
neuro_vis(ng, atlas = 'Subcortex', lowcol = 'darkgreen', midcol = 'yellow2', highcol = 'darkorange')
#> merging atlas and data by 'label', 'atlas', 'side'
There are several key limitations to this resource that we wish to highlight here.
-
First, neuroimaGene describes associations between NIDPs and predicted genetically regulated gene expression (GReX). GReX represents the proportion of gene expression that is genetically determined. Genes that affect a phenotype through protein dysfunction will not show up in neuroimaGene so long as the underlying mRNA expression remains unchanged.
-
Second, the tissue context of the GReX-NIDP association is not causal. GReX-NIDP associations are frequently significant in multiple tissue contexts. In these circumstances, we do not yet have sufficient methods to identify the causal tissue. Even when the GReX measure is only associated with the NIDP in a single tissue, it is important to note that GReX cannot be said to be causal in this tissue without further analyses.
-
Third, while GReX associations are derived from SNP data which is not affected by environment, the S-PrediXcan methodology is still susceptible to confounding from linkage disequilibrium of input SNPs. For causal inference, we recommend applying methods such as MR-JTI (Zhou et al 2020) to the GReX-NIDP associations as a post-hoc test to adjust the association statistics by quantifying and accounting for genetic heterogeneity.
Please see our publication for a more extended discussion on limitations.
GWAS - Genome Wide Association Study
TWAS - Transcriptome Wide Association Study
GReX - Genetically regulated gene expression
JTI - Joint Tissue Imputation (link)
NIDP - Neuroimaging Derived Phenotype
MRI - Magnetic resonance imaging
T1 - MRI modality classically used for structural characterization of the brain
dMRI - diffusion weighted MRI (used in our data for white matter tractography)
fMRI - functional MRI used for examining coordinated activity across regions of the brain
UKB - United Kingdom Biobank
eQTL - expression Quantitative Trait Locus
GTEx - Genotype Tissue Expression Consortium
Barbeira, Alvaro N., et al. “Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics.” Nature communications 9.1 (2018): 1-20.
Zhou, Dan, et al. “A unified framework for joint-tissue transcriptome-wide association and Mendelian randomization analysis.” Nature genetics 52.11 (2020): 1239-1246.
Miller, Karla L., et al. “Multimodal population brain imaging in the UK Biobank prospective epidemiological study.” Nature neuroscience 19.11 (2016): 1523-1536.
Smith, Stephen M., et al. “An expanded set of genome-wide association studies of brain imaging phenotypes in UK Biobank.” Nature neuroscience 24.5 (2021): 737-745.
Elliott, Lloyd T., et al. “Genome-wide association studies of brain imaging phenotypes in UK Biobank.” Nature 562.7726 (2018): 210-216.
Gamazon, Eric R., et al. “Multi-tissue transcriptome analyses identify genetic mechanisms underlying neuropsychiatric traits.” Nature genetics 51.6 (2019): 933-940.
Mowinckel, Athanasia M., and Didac Vidal-Piñeiro. “Visualization of brain statistics with R packages ggseg and ggseg3d.” Advances in Methods and Practices in Psychological Science 3.4 (2020): 466-483.
Please direct all questions to me at the following email: xbledsoe22@gmail.com