#Seven Bridges Cancer Genomics Cloud -- Deploying your own tools
####This project contains these tools and workflows:
- sonnet-grep (tool)
- dna2protein (workflow)
- transcribe (tool)
- translate (tool)
- GeneExpressionWF (workflow)
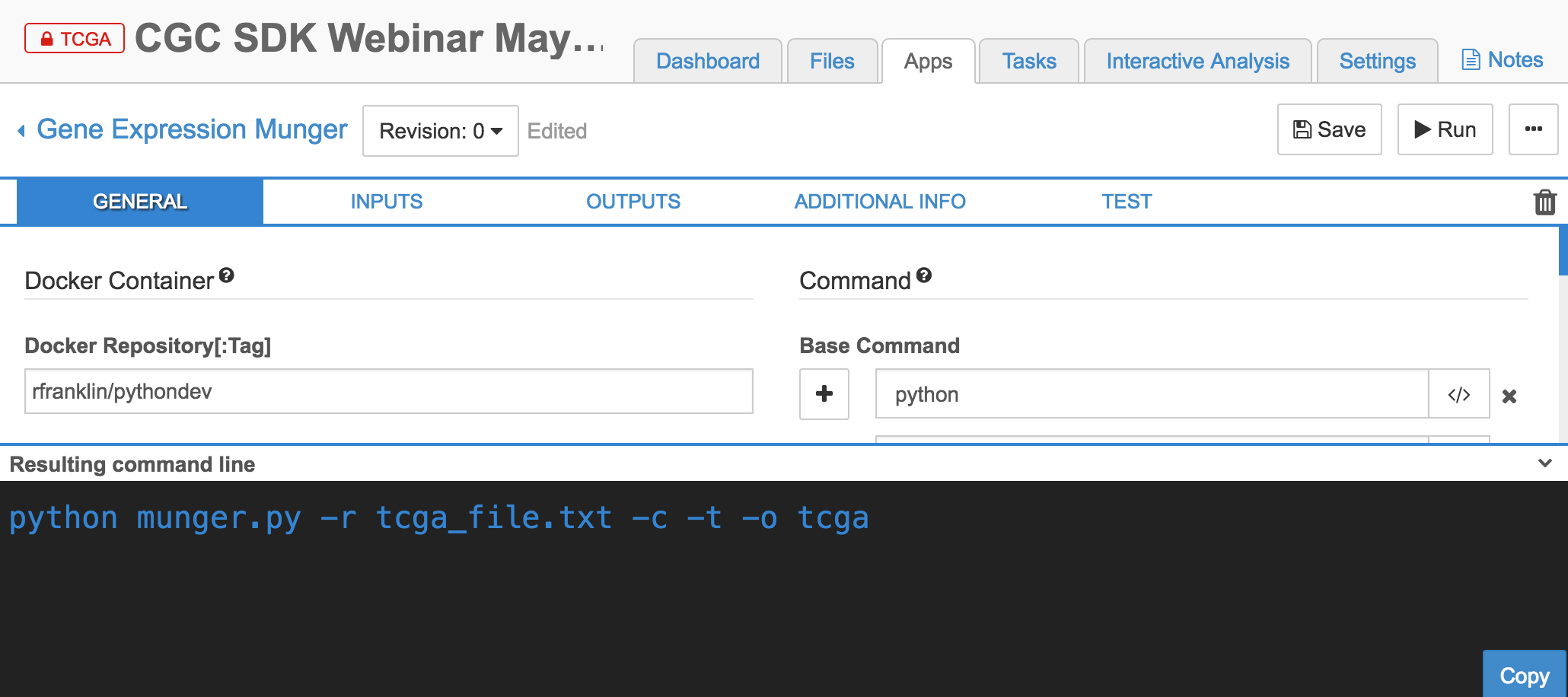
- Gene Expression Munger (tool)
- Differential Expression (tool)
##sonnet-grep This tool looks for patterns in a text file (Shakespeare's 18th Sonnet) and outputs the lines that match. grep is a standard Linux command line tool.
##dna2protein This workflow takes a input text file with a DNA sequence and outputs a text file with a peptide sequence from the first ORF, if possible. dna2protein is a custom workflow written in Python that is composed of transcribe.py (DNA -> mRNA) and translate.py (mRNA -> protein).
##GeneExpressionWF This workflow produces a differential expression analysis from an array of Level 3 TCGA RNASeq Gene Expression Quantification files. It first merges the files into a single matrix and produces a "metadata matrix" from the metadata (e.g. sample type, gender) from each individual case (GeneExpressionMunger, munger.py). It then runs a custom differential expression analysis that produces a matrix of (adj) p-values per gene based on differences among sample-types (e.g. Solid Tissue Normal and Primary Tumor) as well as some plots in PDF (DifferentialExpression, diff.R).
#Import tools from this repo into CGC Project You can import any CWL JSON in these projects as a tool into the Cancer Genomics Cloud. First, you must set up a CGC account and create a project.
Steps:
-
In a project on the CGC, navigate to the Apps tab, then 'Add App'
-
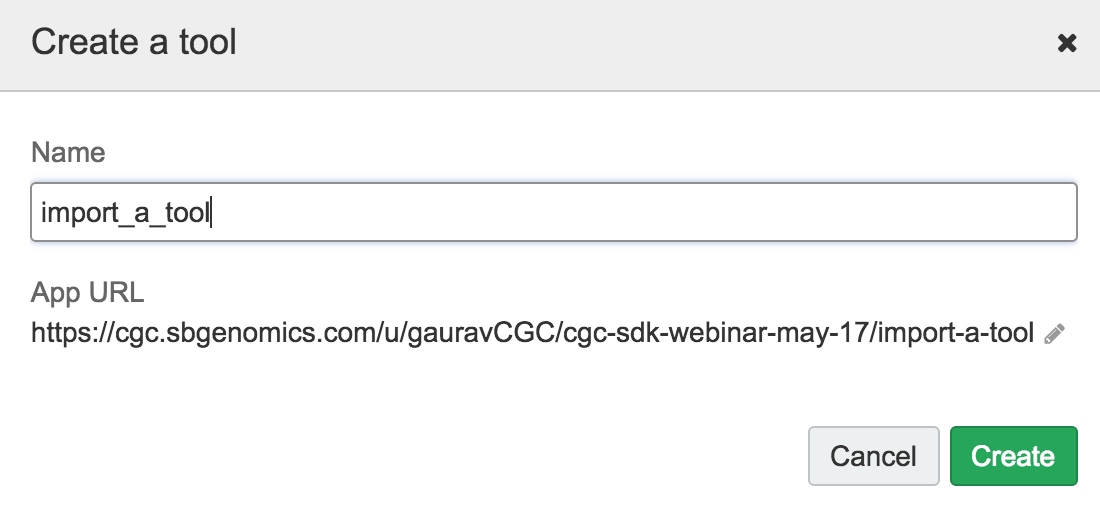
In the Tool Editor, click on '...' on the upper-right and then select 'Import':

-
You will see a blank window. Paste the contents of the CWL JSON file (*.cwl.json) from this repo:
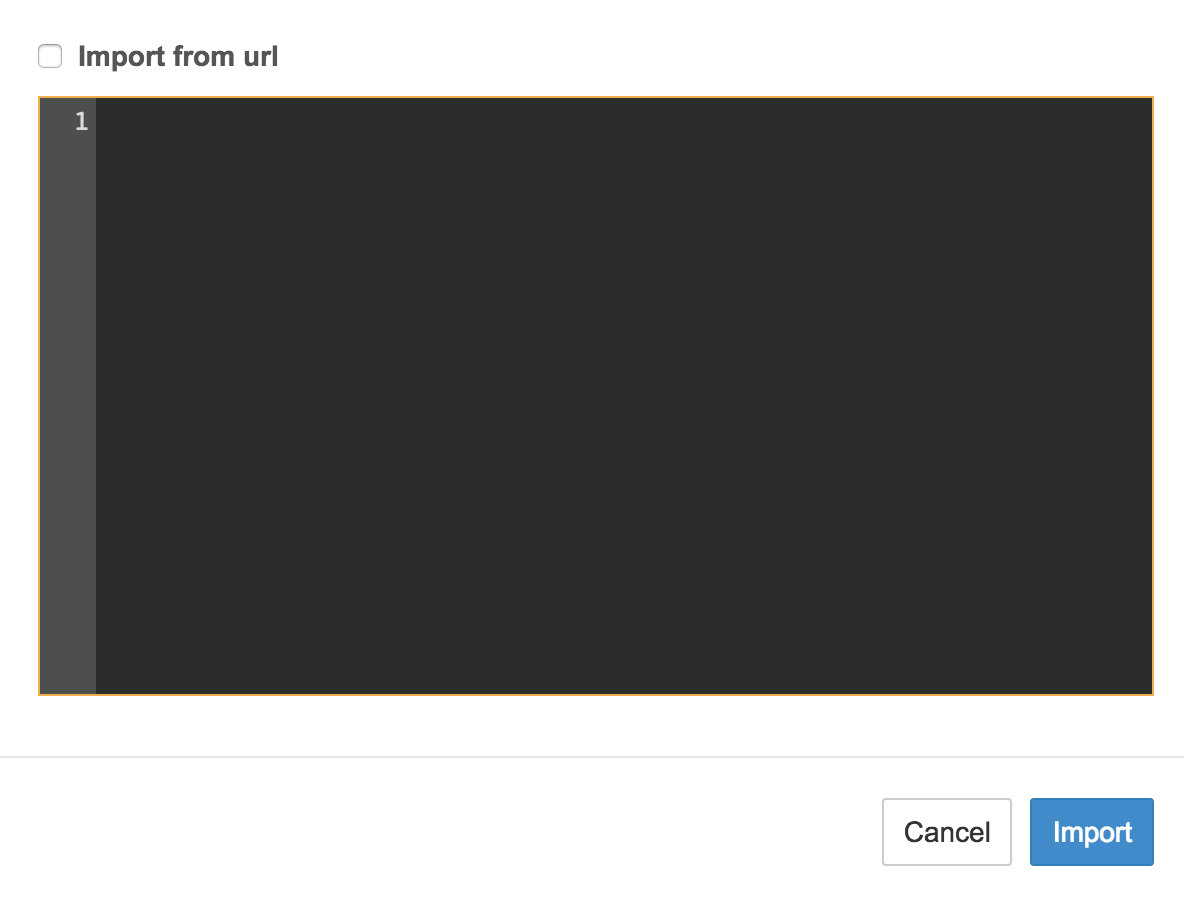
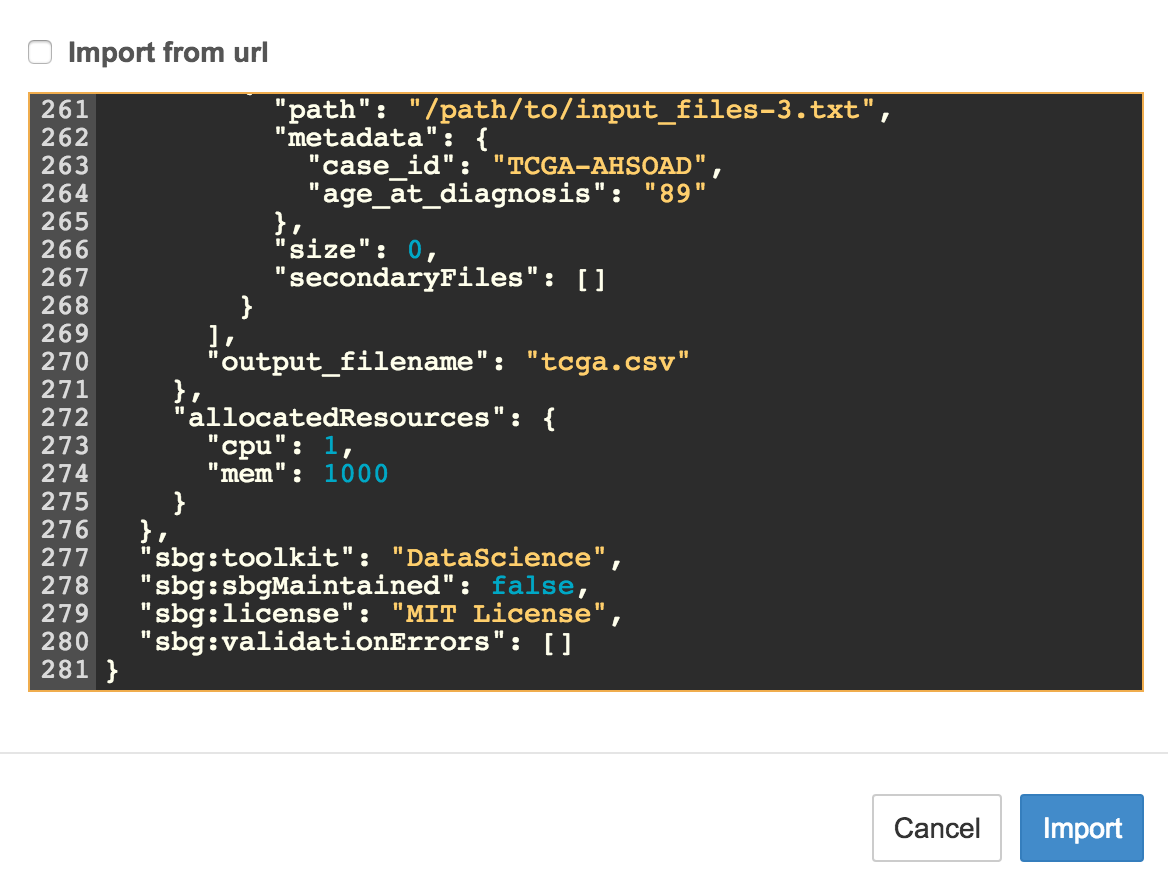
-
Click 'Import'. You will see the appropriate fields in the Tool Editor window: