Separating a 2D kernel (matrix M) into two 1D kernels (column vector C and row vector R) allows for
faster execution of the convolution operation.
Here is an example of a 2D kernel separated into two 1D kernels:
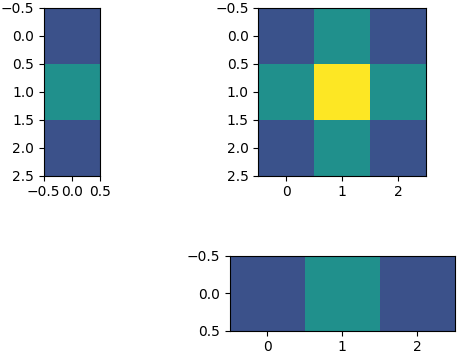
- Clone the repository
- Go to the root level of the repository (where setup.py is located)
- Execute
pip install .
Example of how to use the package:
import numpy as np
from separate_kernel import separate_kernel
M = np.array([[1, 2, 1], [2, 4, 2], [1, 2, 1]])
res = separate_kernel(M, symmetric_kernel=True)
print(f'1st 1D filter: {res.col_vec.flatten()}.T')
print(f'2nd 1D filter: {res.row_vec.flatten()}')
print(f'Multiplying 1st and 2nd 1D filter gives original 2D filter:')
print(res.col_vec @ res.row_vec)This outputs:
1st 1D filter: [1. 2. 1.].T
2nd 1D filter: [1. 2. 1.]
Multiplying 1st and 2nd 1D filter gives original 2D filter:
[[1. 2. 1.]
[2. 4. 2.]
[1. 2. 1.]]
For a 2D kernel to be separable the rank of the matrix M must be 1.
For a separable kernel the equation M = C * R holds.
The method uses nonlinear least squares to minimize the residuals C * R - M.
Therefore, even if a kernel can not be perfectly separated, still an approximation will be output.
Also, SVD could be used to compute C and R, as in the case of separability there is only one nonzero singular value
and the corresponding singular vectors represent (scaled versions of) C and R.