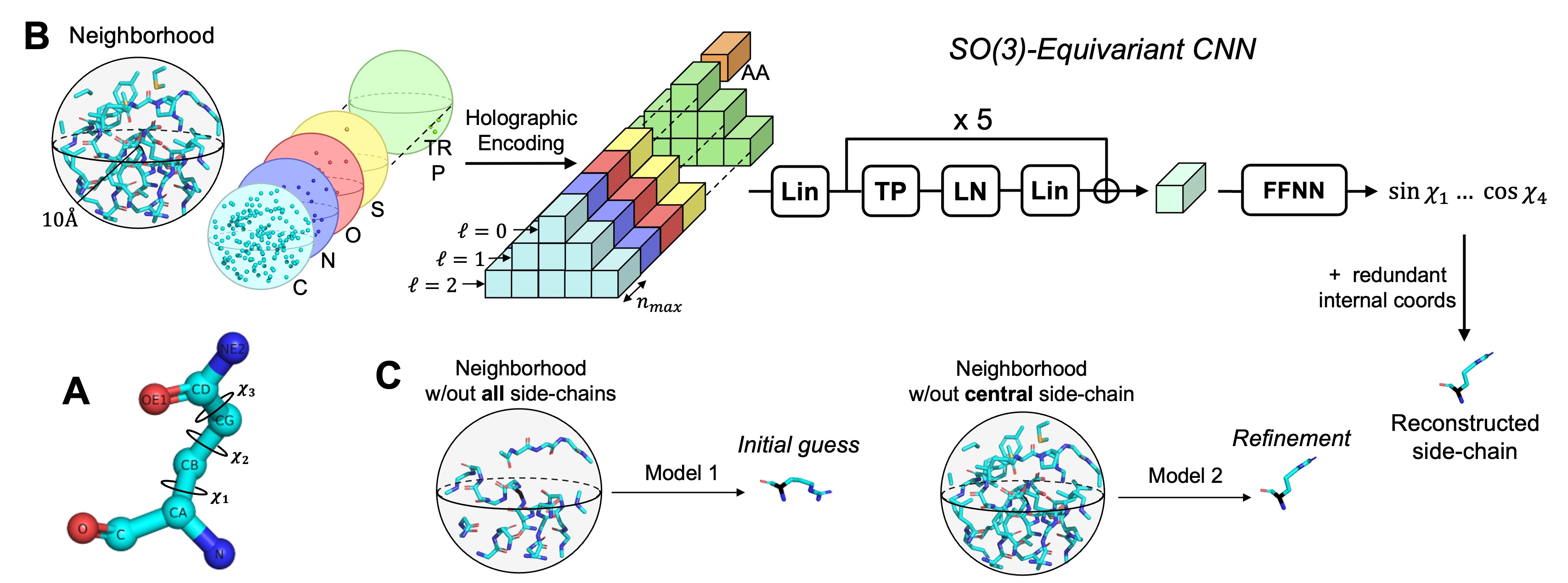
H-Packer: Holographic Rotationally Equivariant Convolutional Neural Network for Protein Side-Chain Packing
This repo contains code for H-Packer, a method for side-chain packing based upon rotationally equivariant convolutional neural networks.
- Packing side-chain conformations of a full structure, providing a backbone structure and desired sequence information
- Refining side-chain conformations of a full structure
- Add and pack side-chains in parts of a structure (keeping some of the structure constant)
- Apply mutations and selectively pack the surrounding side-chains
- Training new HPacker models
Create the hpacker conda environment by running the following
conda env create -f env.ymlto install the necessary dependencies.
Then run
pip install .to install the code in this repo as a package.
If you're going to make edits to the code, run
pip install -e .so you can test your changes.
As simple as a few lines of code:
from hpacker import HPacker
# Initialize HPacker object by passing it a tutple of paths to the pre-trained models, and the backbone-only structure that you want to add side-chains to
hpacker = HPacker(['pretrained_models/initial_guess','pretrained_models/refinement','pretrained_models/initial_guess_conditioned'], 'T0950_bb_only.pdb')
hpacker.reconstruct_sidechains(num_refinement_iterations=5)
hpacker.write_pdb('reconstructed_from_bb_only_T0950.pdb')See the provided hpacker.ipynb notebook for more examples, as well as explanations of the inner workings of H-Packer.
Coming soon
- Cannot process hetero residues, since they do not play nice with BioPython's
internal_coordsmodule.
If you used H-Packer or learned something from it, please cite us:
@misc{visani2023hpacker,
title={H-Packer: Holographic Rotationally Equivariant Convolutional Neural Network for Protein Side-Chain Packing},
author={Gian Marco Visani and William Galvin and Michael Neal Pun and Armita Nourmohammad},
year={2023},
eprint={2311.09312},
archivePrefix={arXiv},
primaryClass={q-bio.BM}
}