fair-software.nl recommendations:
Code quality checks:

Matchms is an open-source Python package to import, process, clean, and compare mass spectrometry data (MS/MS). It allows to implement and run an easy-to-follow, easy-to-reproduce workflow from raw mass spectra to pre- and post-processed spectral data. Spectral data can be imported from common formats such mzML, mzXML, msp, metabolomics-USI, MGF, or json (e.g. GNPS-syle json files). Matchms then provides filters for metadata cleaning and checking, as well as for basic peak filtering. Finally, matchms was build to import and apply different similarity measures to compare large amounts of spectra. This includes common Cosine scores, but can also easily be extended by custom measures. Example for spectrum similarity measures that were designed to work in matchms are Spec2Vec and MS2DeepScore.
If you use matchms in your research, please cite the following software paper:
F Huber, S. Verhoeven, C. Meijer, H. Spreeuw, E. M. Villanueva Castilla, C. Geng, J.J.J. van der Hooft, S. Rogers, A. Belloum, F. Diblen, J.H. Spaaks, (2020). matchms - processing and similarity evaluation of mass spectrometry data. Journal of Open Source Software, 5(52), 2411, https://doi.org/10.21105/joss.02411
To make typical matchms workflows (data import, processing, score computations) more accessible to users, matchms now offers a Pipeline class to handle complex workflows. This also allows to define, import, export, or modify workflows using yaml files. See code examples below (and soon: updated tutorial).
We realized that many matchms-based workflows aim to compare many-to-many spectra whereby not all pairs and scores are equally important. Often, for instance, it will be about searching similar or related spectra/compounds. This also means that often not all scores need to be stored (or computed). For this reason we now shifted to a sparse handling of scores in matchms (that means: only storing actuallly computed, non-null values).
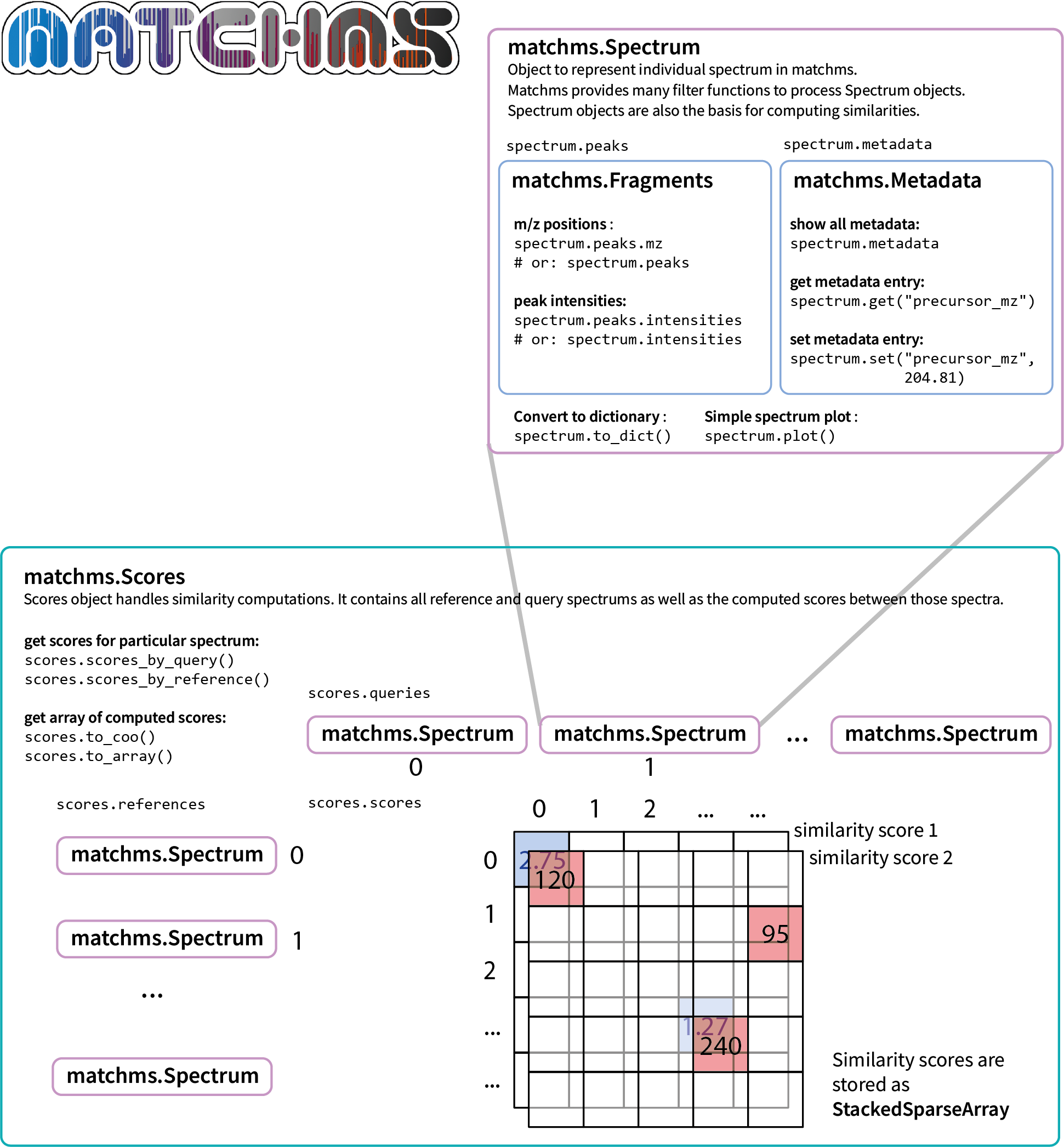

This is the first of a few releases to work our way towards matchms 1.0.0, which also means that a few things in the API will likely change. Here the main change is that Spectrum.metadata is no longer a simple Python dictionary but became a Metadata object. In this context metadata field-names/keys will now be harmonized by default (e.g. "Precursor Mass" will become "precursor_mz). For list of conversions see matchms key conversion table.
- metadata is now stored using new
Metadataclass which automatically applied restrictions to used field names/keys to avoid confusion between different format styles - all metadata keys must be lower-case, spaces will be changed to underscores.
- Known key conversions are applied to metadata entries using a matchms key conversion table.
- new
MetadataMatchsimilarity measure in matchms.similarity. This can be used to find matches between metadata entries and currently supports either full string matches or matches of numerical entries within a specified tolerance - new
interpret_pepmass()filter to handle different pepmass entries found in data Spikesclass has becomeFragmentsclass
For more extensive documentation see our readthedocs and our matchms introduction tutorial.
Prerequisites:
- Python 3.7, 3.8 or 3.9, (3.10 and 3.11 should work as well, but are not yet tested systematically)
- Anaconda (recommended)
We recommend installing matchms in a new virtual environment to avoid dependency clashes
conda create --name matchms python=3.9
conda activate matchms
conda install --channel bioconda --channel conda-forge matchmsAlternatively, matchms can also be installed using pip but users will then either have to install rdkit on their own or won't be able to use the entire functionality. Without rdkit installed several filter functions related to processing and cleaning chemical metadata will not run.
To install matchms with pip simply run
pip install matchmsMatchms functionalities can be complemented by additional packages. To date we are aware of:
- Spec2Vec an alternative machine-learning spectral similarity score that can simply be installed by pip install spec2vec and be imported as from spec2vec import Spec2Vec following the same API as the scores in matchms.similarity.
- MS2DeepScore a supervised, deep-learning based spectral similarity score that can simply be installed by pip install ms2deepscore and be imported as from ms2deepscore import MS2DeepScore following the same API as the scores in matchms.similarity.
- matchmsextras which contains additional functions to create networks based on spectral similarities, to run spectrum searchers against PubChem, or additional plotting methods.
- memo a method allowing a Retention Time (RT) agnostic alignment of metabolomics samples using the fragmentation spectra (MS2) of their consituents.
(if you know of any other packages that are fully compatible with matchms, let us know!)
To get started with matchms, we recommend following our matchms introduction tutorial.
Below is a small example of using matchms to calculate the Cosine score between mass Spectrums in the tests/pesticides.mgf file.
from matchms import Pipeline
pipeline = Pipeline()
# Read spectrums from a MGF formatted file, for other formats see https://matchms.readthedocs.io/en/latest/api/matchms.importing.html
pipeline.query_files = "tests/pesticides.mgf"
pipeline.filter_steps_queries = [
["default_filters"],
["add_parent_mass"],
["normalize_intensities"],
["select_by_intensity", {"intensity_from": 0.001, "intensity_to": 1.0}],
["select_by_mz", {"mz_from": 0, "mz_to": 1000}],
["require_minimum_number_of_peaks", {"n_required": 5}]
]
pipeline.score_computations = [["precursormzmatch", {"tolerance": 100.0}],
["cosinegreedy", {"tolerance": 1.0}],
["filter_by_range", {"name": "CosineGreedy_score", "low": 0.2}]]
pipeline.logging_file = "my_pipeline.log" # for pipeline and logging message
pipeline.logging_level = "INFO"
pipeline.run()Alternatively, in particular if you need more room to add custom functions and steps, the individual
steps can run without using the matchms Pipeline:
from matchms.importing import load_from_mgf
from matchms.filtering import default_filters, normalize_intensities
from matchms import calculate_scores
from matchms.similarity import CosineGreedy
# Read spectrums from a MGF formatted file, for other formats see https://matchms.readthedocs.io/en/latest/api/matchms.importing.html
file = load_from_mgf("tests/pesticides.mgf")
# Apply filters to clean and enhance each spectrum
spectrums = []
for spectrum in file:
# Apply default filter to standardize ion mode, correct charge and more.
# Default filter is fully explained at https://matchms.readthedocs.io/en/latest/api/matchms.filtering.html .
spectrum = default_filters(spectrum)
# Scale peak intensities to maximum of 1
spectrum = normalize_intensities(spectrum)
spectrums.append(spectrum)
# Calculate Cosine similarity scores between all spectrums
# For other similarity score methods see https://matchms.readthedocs.io/en/latest/api/matchms.similarity.html .
scores = calculate_scores(references=spectrums,
queries=spectrums,
similarity_function=CosineGreedy())
# Matchms allows to get the best matches for any query using scores_by_query
query = spectrums[15] # just an example
best_matches = scores.scores_by_query(query, 'CosineGreedy_score', sort=True)
# Print the calculated scores for each spectrum pair
for (reference, score) in best_matches[:10]
# Ignore scores between same spectrum
if reference is not query:
print(f"Reference scan id: {reference.metadata['scans']}")
print(f"Query scan id: {query.metadata['scans']}")
print(f"Score: {score[0]:.4f}")
print(f"Number of matching peaks: {score[1]}")
print("----------------------------")Matchms comes with numerous different scoring methods in matchms.similarity and can furthe seemlessly work with Spec2Vec or MS2DeepScore.
Code example:
from matchms.importing import load_from_usi
import matchms.filtering as msfilters
import matchms.similarity as mssim
usi1 = "mzspec:GNPS:GNPS-LIBRARY:accession:CCMSLIB00000424840"
usi2 = "mzspec:MSV000086109:BD5_dil2x_BD5_01_57213:scan:760"
mz_tolerance = 0.1
spectrum1 = load_from_usi(usi1)
spectrum1 = msfilters.select_by_mz(spectrum1, 0, spectrum1.get("precursor_mz"))
spectrum1 = msfilters.remove_peaks_around_precursor_mz(spectrum1,
mz_tolerance=0.1)
spectrum2 = load_from_usi(usi2)
spectrum2 = msfilters.select_by_mz(spectrum2, 0, spectrum1.get("precursor_mz"))
spectrum2 = msfilters.remove_peaks_around_precursor_mz(spectrum2,
mz_tolerance=0.1)
# Compute scores:
similarity_cosine = mssim.CosineGreedy(tolerance=mz_tolerance).pair(spectrum1, spectrum2)
similarity_modified_cosine = mssim.ModifiedCosine(tolerance=mz_tolerance).pair(spectrum1, spectrum2)
similarity_neutral_losses = mssim.NeutralLossesCosine(tolerance=mz_tolerance).pair(spectrum1, spectrum2)
print(f"similarity_cosine: {similarity_cosine}")
print(f"similarity_modified_cosine: {similarity_modified_cosine}")
print(f"similarity_neutral_losses: {similarity_neutral_losses}")
spectrum1.plot_against(spectrum2)To install matchms, do:
git clone https://github.com/matchms/matchms.git
cd matchms
conda create --name matchms-dev python=3.8
conda activate matchms-dev
# Install rdkit using conda, rest of dependencies can be installed with pip
conda install -c conda-forge rdkit
python -m pip install --upgrade pip
pip install --editable .[dev]Run the linter with:
prospectorAutomatically fix incorrectly sorted imports:
isort .Files will be changed in place and need to be committed manually. If you only want to inspect the isort suggestions then simply run:
isort --check-only --diff .Run tests (including coverage) with:
pytestThe conda packaging is handled by a recipe at Bioconda.
Publishing to PyPI will trigger the creation of a pull request on the bioconda recipes repository Once the PR is merged the new version of matchms will appear on https://anaconda.org/bioconda/matchms
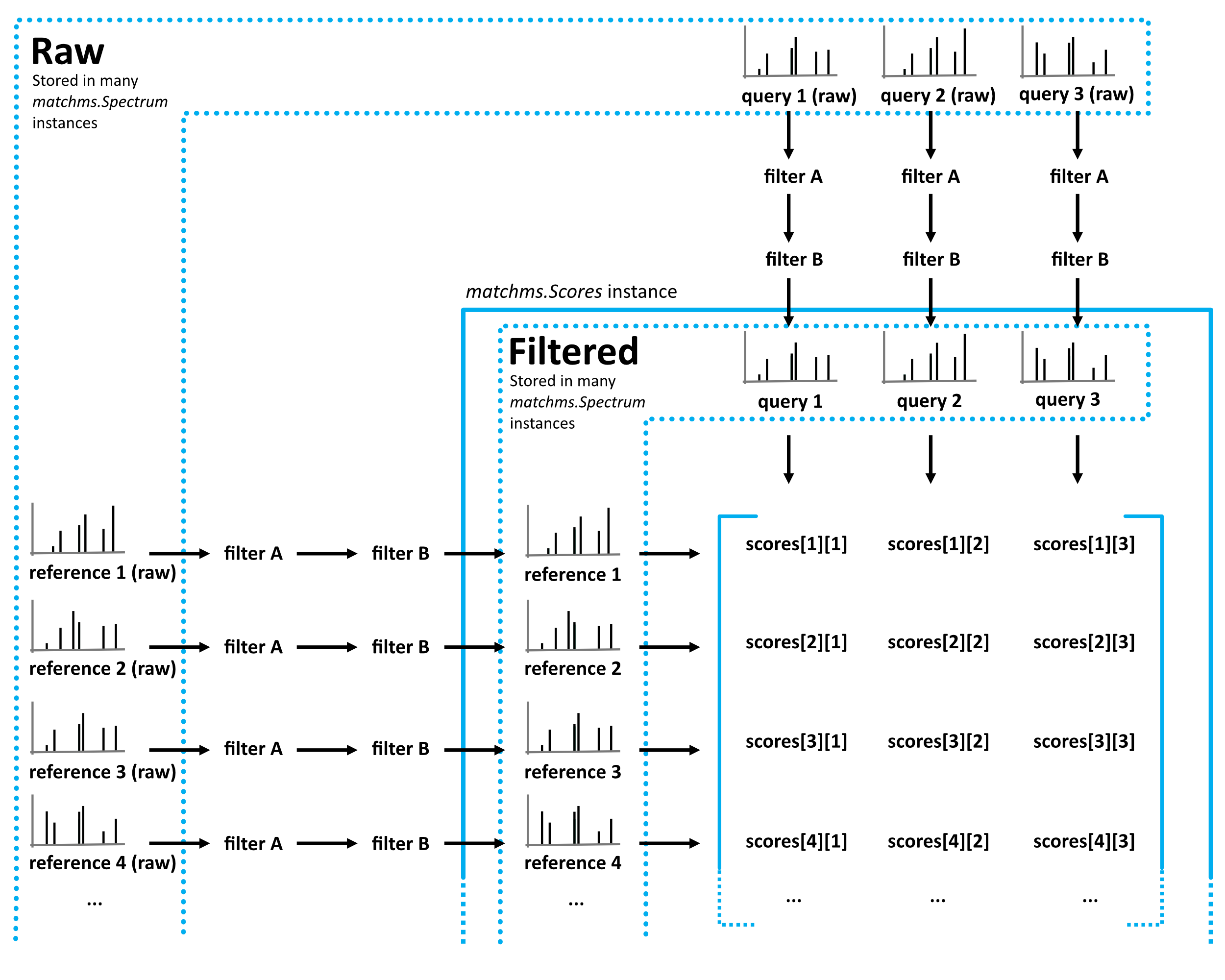
Flowchart of matchms workflow. Reference and query spectrums are filtered using the same set of set filters (here: filter A and filter B). Once filtered, every reference spectrum is compared to every query spectrum using the matchms.Scores object.
If you want to contribute to the development of matchms, have a look at the contribution guidelines.
Copyright (c) 2021, Netherlands eScience Center
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
This package was created with Cookiecutter and the NLeSC/python-template.




