For detailed documentation and tutorials, please visit our GitHub Pages site. The Articles section provides several detailed vignettes on the use of gmod.
gmod (Grammar of Modelling) is an R package for build Markov models and decision trees for Medical Decision Making and cost-effectiveness analyses. There is a graphical user interface at DecisionTwig to develop with the gmod syntax visually.
To install gmod from GitHub, use the following command in R:
library(devtools)
install_github("hjalal/gmod")gmod builds a model in layers, similar to ggplot2:
mymodel <- gmod() +
decisions() +
states() +
event() +
event() +
...
payoffs()This will generate a gmod object with 1 decisions layer, 1 states layer, one payoff layer, and 1 or more event layers.
Consider a simple Markov model with 2 decisions: StandardofCare, and NewTreatment, and two health states: Healthy and Dead. The cohort starts at the healthy state each year there is a 0.1 probability of death.
First, we define the generic cycle tree, which is at the core of the Grammar of Modeling.
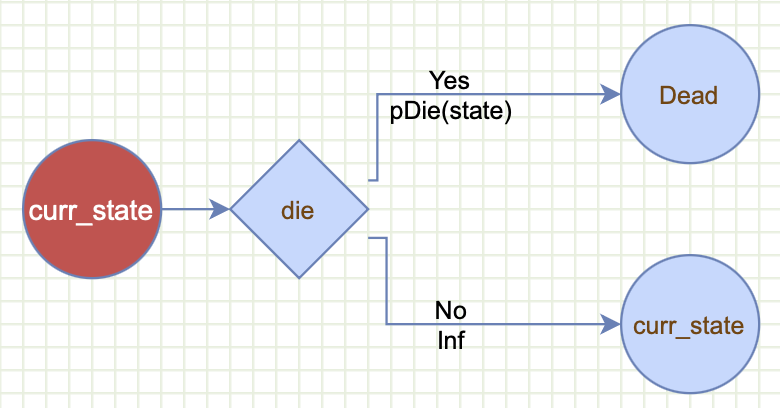
This same generic cycle tree is applied to both health states ("Healthy" and "Dead"). For example, in each cycle a proportion of the cohort that is healthy, will die. This will be determined by the function pDie(state="Healthy"). The rest will stay healthy computed by the infinity Inf placeholder which gmod will translate to 1-pDie(state="Healthy"). The proportion that remains healthy handled by the special health state curr_state. Likewise, this cycle tree will also be applied to the proportion of the cohort that is already dead. But, in this case, none of the cohort will die because pDie(state="Dead") returns 0.
The gmod syntax for this generic cycle tree will consist of a single event layer:
event(name="die",
scenarios=c("Yes","No"),
probs=c(pDie(decision, state),Inf),
outcomes=c("Dead","curr_state"))and we can define pDie like a standard R function. Here we assume that NewTreatment reduces probability of death from 0.2 to 0.1:
pDie <- function(decision, state){
if(state == "Healthy"){
if (decision == "NewTreatment") 0.1 else 0.2
} else {
0
}
}Similarly, we can define a function compute_cost which we will set to return $1000 for NewTreatment, and 0 for StandardOfCare:
compute_cost <- function(decision){
if (decision=="NewTreatment") 1000 else 0
}The code below shows the full gmod syntax with the two functions:
# gmod
mygmod <- gmod() +
decisions("StandardOfCare","NewTreatment") +
states(names=c("Healthy","Dead"),
init_probs=c(1,0)) +
event(name="die",
scenarios=c("Yes","No"),
probs=c(pDie(state),Inf),
outcomes=c("Dead","curr_state")) +
payoffs(cost=compute_cost(decision))
# probabiltiy of death
pDie <- function(decision, state){
if(state == "Healthy"){
if (decision == "NewTreatment") 0.1 else 0.2
} else {
0
}
}
# cost payoff
compute_cost <- function(decision){
if (decision=="NewTreatment") 1000 else 0
}To confirm that the functions are behaving as expected, it is always good to check them. the function gmod_expand_functions(mygmod) iterates through the functions and dependencies and produces a dataset for each function with the values.
gmod_expand_functions(mygmod)
# Note: The dataset df_compute_cost created for function compute_cost .
# Note: The dataset df_pDie created for function pDie .
# df_pDie
# decision state pDie
# 1 StandardOfCare Healthy 0.2
# 2 NewTreatment Healthy 0.1
# 3 StandardOfCare Dead 0.0
# 4 NewTreatment Dead 0.0
# df_compute_cost
# decision compute_cost
# 1 StandardOfCare 0
# 2 NewTreatment 1000pDie and compute_cost are behaving as expected.
First, we convert the gmod syntax to a standard R function, using gmod_gen_model_function(), then we define the n_cycles variable, and run the generated model function, which by default it will be named my_markov_model
model_struc <- gmod_gen_model_function(mygmod)
n_cycles <- 5
my_markov_model(model_struc)
# $summary_payoffs
# cost
# StandardOfCare 0
# NewTreatment 5000As expected for the 5 cycles, cost of NewTreatment is $5000. We can also produce the transition probability matrix P and the Markov trace Trace:
my_markov_model(model_struc, return_transition_prob = T, return_trace = T)
# $P
# , , StandardOfCare
#
# Healthy Dead
# Healthy 0.8 0.2
# Dead 0.0 1.0
#
# , , NewTreatment
#
# Healthy Dead
# Healthy 0.9 0.1
# Dead 0.0 1.0
#
#
# $Trace
# , , StandardOfCare
#
# Healthy Dead
# 1 1.0000 0.0000
# 2 0.8000 0.2000
# 3 0.6400 0.3600
# 4 0.5120 0.4880
# 5 0.4096 0.5904
#
# , , NewTreatment
#
# Healthy Dead
# 1 1.0000 0.0000
# 2 0.9000 0.1000
# 3 0.8100 0.1900
# 4 0.7290 0.2710
# 5 0.6561 0.3439
#
#
# $summary_payoffs
# cost
# StandardOfCare 0
# NewTreatment 5000currently, gmod can support these features:
-
simulation time dependency (e.g., age dependency)
-
state residency dependency (e.g., tunnel states)
-
multiple payoffs (e.g., cost, effectiveness, life expectancy, ... etc)
-
multiple events in each cycle
-
transition rewards (e.g., transitional cost or disutility)
-
discounting
Explore gmod capabilities by reviewing the vignettes in the Articles section. Currently, there are 8 vignettes, 4 for decision trees D0-D3, and 4 for Markov models M0-M3. The vignettes are labelled from 0 to 3 going from beginner to advanced.
Please note that both Decision Twigs and gmod are still under active development and are provided as-is without any warranty.
Jalal, H. (2024). Grammar of Modelling, gmod R package. Retrieved from https://github.com/hjalal/gmod
Jalal, H. (2024). DecisionTwig. Retrieved from https://www.dashlab.ca/projects/decision_twig/