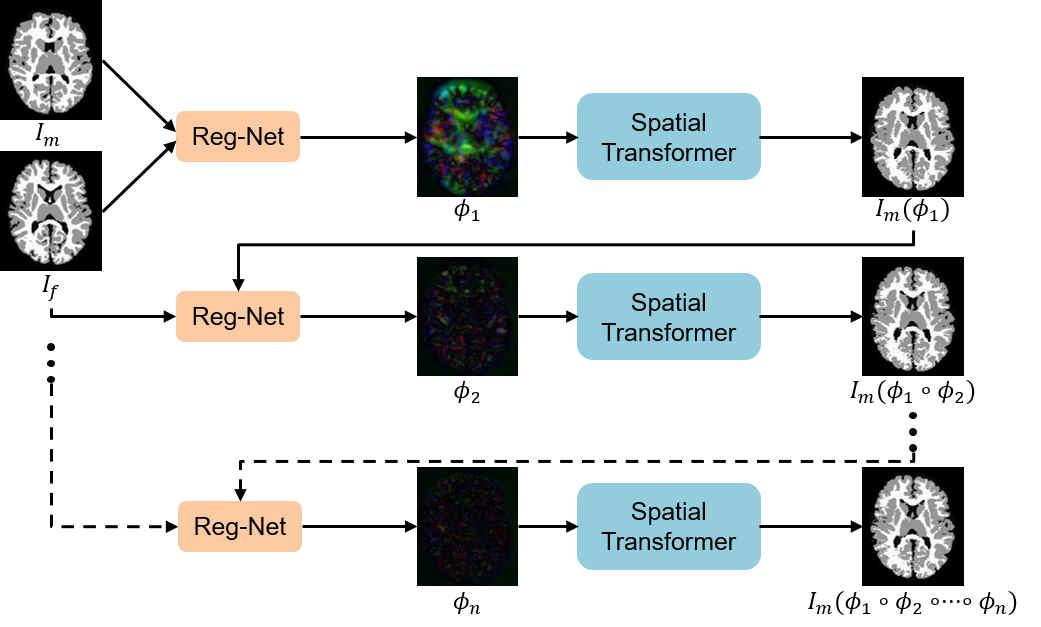
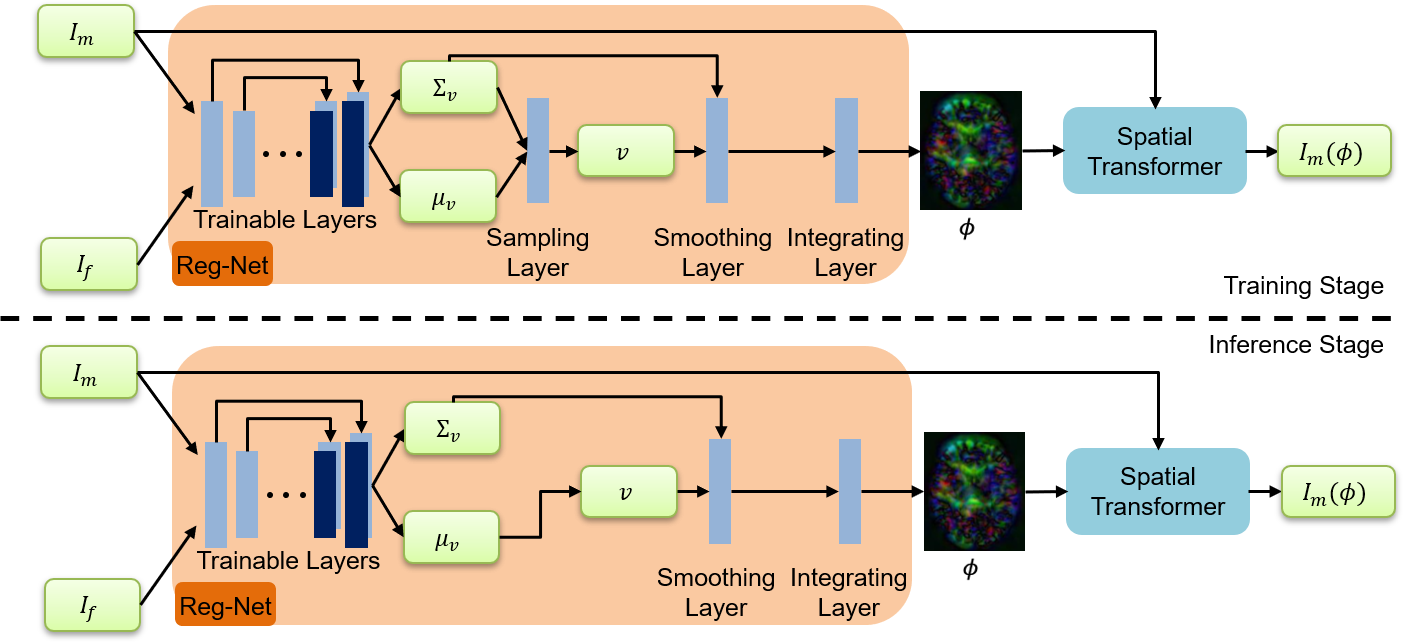
An Auto-Context based Tissue-Aware Deformable Registration Network for Infant Brain MRI
Coming soon! Code review processing. Welcome for any bugs report.
-
If you use ACTA-Reg-Net, please cite:
An Auto-Context Deformable Registration Network for Infant Brain MRI
Dongming Wei, Sahar Ahmad, Yunzhi Huang, Lei Ma, Zhengwang Wu, Gang Li, Li Wang, Qian Wang, Pew-Thian Yap, Dinggang Shen
eprint arXiv:2005.09230
Instruction
This code is for registering infant brain MR images. The registration is based on the segmentation map (gray matter, white matter). You can try to modify the directory in train.py to train your model. Or you can directly use test.py to test your data, where a trained model is given in ./models.
For obtaining an ACTA-Reg-Net, you need to work on ./src/demo.sh to run all the experiments.
Requirements
- Python 3.6 (3.7 should work well)
- Tensorflow 1.10 (any 1.xx version should work well)
- Keras 2.2.4
- Bash
You can choose to run
pip install -r requirements.txtOr you can perform
conda create -n tf10-py36 python=3.6
conda activate tf10-py36
conda install tensorflow-gpu==1.10Train
python train.pyFor training your dataset, you need to modify the data directory in the trian.py. For our task, we save the infant brian images into the ../data/MAPS_DATASET/Train_Set. After this step, you have obtained your Reg-Net, which is supposed to generate smooth deformation fields. Then, you can execute the demo.sh to perform an 'auto-context' manner to boost the registration performance.
Test
python test.py gpu_id ../models/ iteration_num fixed.nii.gz moving.nii.gz moving_label.nii.gzDemo
cd ./src
./demo.sh -m moving.nii.gz -l moving_label.nii.gz -n save_dir -f fixed.nii.gzThe results are saved into ../data/results/*, including warped moving image, moving label, deformation field, and displacement uncertainty map.
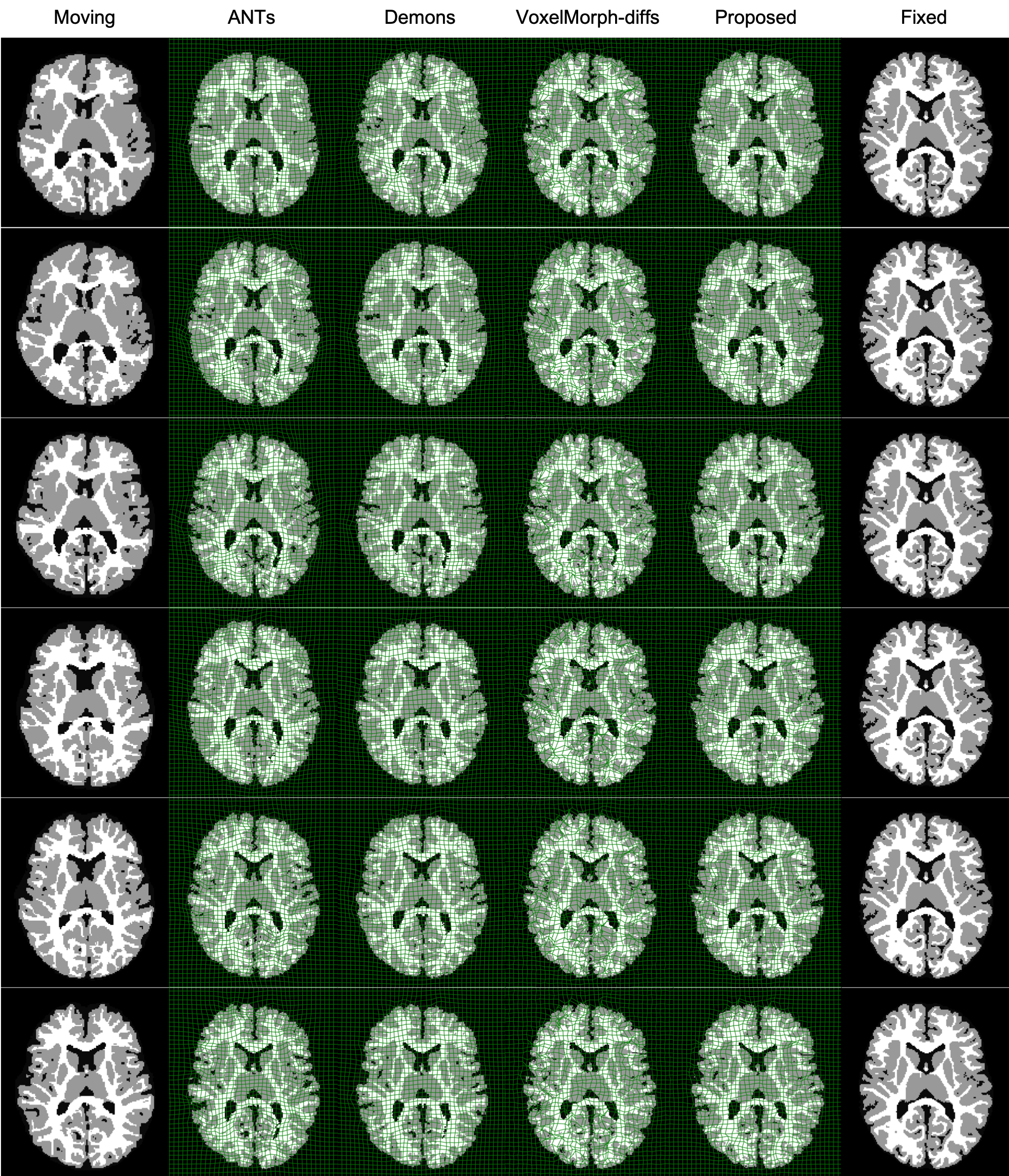
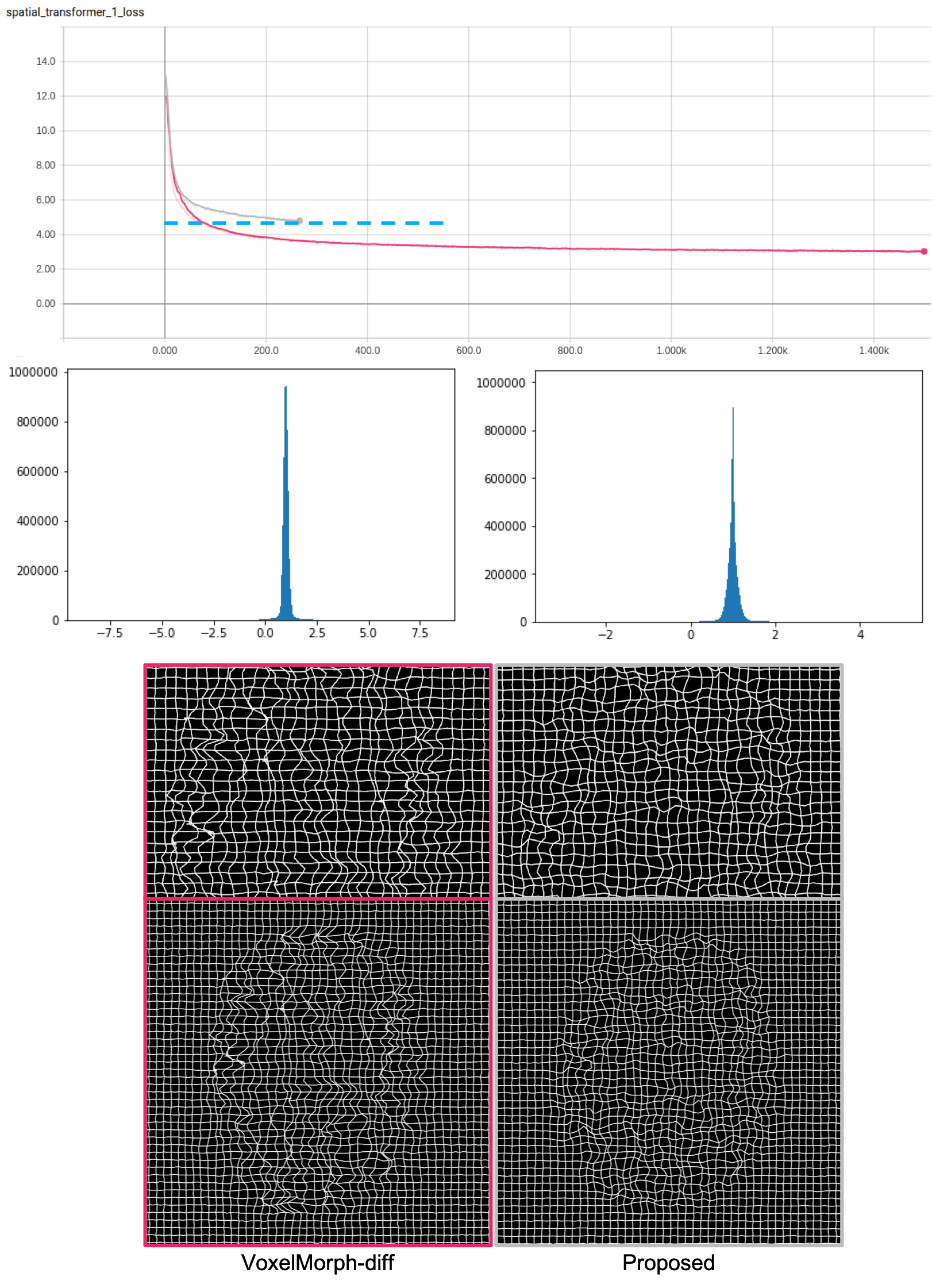
Result
Acknowledgement:
We would like to acknowledge the contribution of VoxelMorph.
Contact:
For any problems, please open an issue.