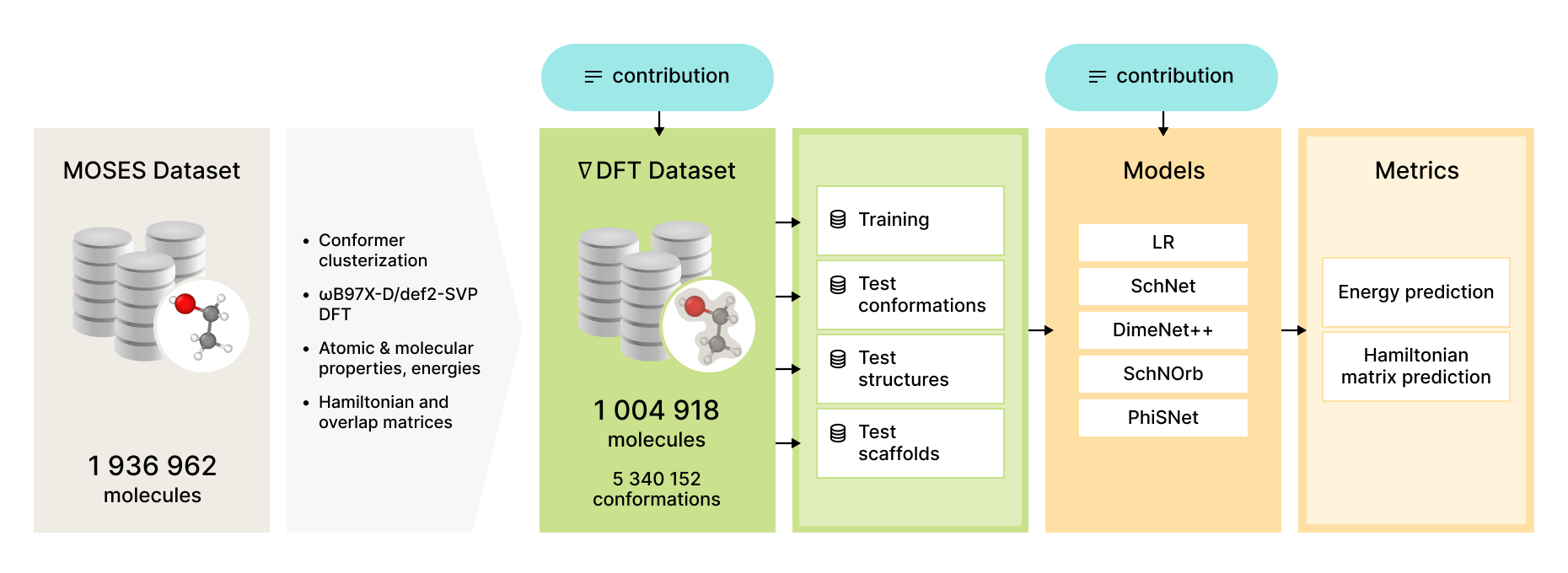
We propose a benchmarking dataset based on a subset of Molecular Sets (MOSES) dataset. Resulting dataset contains 1 004 918 molecules with atoms C, N, S, O, F, Cl, Br, H. It contains 226 424 unique Bemis-Murcko scaffolds and 34 572 unique BRICS fragments.
For each molecule in the dataset we provide from 1 to 62 unique conformations, with 5 340 152 total conformations. For each conformation, we have calculated its electronic properties including the energy (E), DFT Hamiltonian matrix (H), and DFT overlap matrix (S). All properties were calculated using the Kohn-Sham method at ωB97X-D/def2-SVP levels of theory using the quantum-chemical software package Psi4, version 1.5.
We provide several splits of the dataset that can serve as the basis for comparison across different models. First, we fix the training set that consists of 100 000 molecules with 436 581 conformations and its smaller subsets with 10 000, 5 000, and 2 000 molecules and 38 364, 20 349, and 5 768 conformations respectively; these subsets can help determine how much additional data helps various models. We choose another 100 000 random molecules as a structure test set. The scaffold test set has 100 000 molecules containing a Bemis-Murcko scaffold from a random subset of scaffolds which are not present in the training set. Finally, the conformation test set consists of 91 182 (resp., 10 000, 5 000, 2 000) molecules from the training set with new conformations, numbering in total 92 821 (8 892, 4 897, 1 724) conformations; this set can be used for the single-molecule setup.
As part of the benchmark, we provide separate databases for each subset and task and a complete archive with wave function files produced by the Psi4 package that contains quantum chemical properties of the corresponding molecule and can be used in further computations.
The full hamiltonian database is available at nablaDFT Hamiltonian database (7 TB)
Links to other hamiltonian databases including different train and test subsets are in file Hamiltonian databases
An archive with numpy indexes: splits indexes
Links to energy databases including different train and test subsets are in file Energy databases
Links to tarballs: wave functions
The csv file with conformations index, SMILES, atomic DFT properties and wfn archive names: summary.csv
Tar archive with xyz files archive
from nablaDFT.dataset import HamiltonianDatabase
train = HamiltonianDatabase("dataset_train_2k.db")
Z, R, E, F, H, S, C = train[0] # atoms numbers, atoms positions, energy, forces, core hamiltonian, overlap matrix, coefficients matrixfrom ase.db import connect
train = connect("train_2k_energy.db")
atoms_data = connect.get(1)A variety of properties can also be loaded directly from the wavefunctions. See main paper for more details. Properties include DFT matrices:
import numpy as np
import psi4
wfn = np.load(<PATH_TO_WFN>, allow_pickle=True).tolist()
orbital_matrix_a = wfn["matrix"]["Ca"] # alpha orbital coefficients
orbital_matrix_b = wfn["matrix"]["Cb"] # betta orbital coefficients
density_matrix_a = wfn["matrix"]["Da"] # alpha electonic density
density_matrix_b = wfn["matrix"]["Db"] # betta electonic density
aotoso_matrix = wfn["matrix"]["aotoso"] # atomic orbital to symmetry orbital transformation matrix
core_hamiltonian_matrix = wfn["matrix"]["H"] # core Hamiltonian matrix
fock_matrix_a = wfn["matrix"]["Fa"] # DFT alpha Fock matrix
fock_matrix_b = wfn["matrix"]["Fb"] # DFT betta Fock matrix and bond orders for covalent and non-covalent interactions and atomic charges:
wfn = psi4.core.Wavefunction.from_file(<PATH_TO_WFN>)
psi4.oeprop(wfn, "MAYER_INDICES")
psi4.oeprop(wfn, "WIBERG_LOWDIN_INDICES")
psi4.oeprop(wfn, "MULLIKEN_CHARGES")
psi4.oeprop(wfn, "LOWDIN_CHARGES")
meyer_bos = wfn.array_variables()["MAYER INDICES"] # Mayer bond indices
lodwin_bos = wfn.array_variables()["WIBERG LOWDIN INDICES"] # Wiberg bond indices
mulliken_charges = wfn.array_variables()["MULLIKEN CHARGES"] # Mulliken atomic charges
lowdin_charges = wfn.array_variables()["LOWDIN CHARGES"] # Löwdin atomic charges- Unifying machine learning and quantum chemistry with a deep neural network for molecular wavefunctions (SchNOrb)
- SE(3)-equivariant prediction of molecular wavefunctions and electronic densities (PhiSNet)
- A continuous-filter convolutional neural network for modeling quantum interactions (SchNet)
- Equivariant message passing for the prediction of tensorial properties and molecular spectra (PaiNN)
- Fast and Uncertainty-Aware Directional Message Passing for Non-Equilibrium Molecules (DimeNet++)
Several checkpoints for each model is available here: checkpoints links
In the tables below ST, SF, CF denote structures test set, scaffolds test set and conformations test set correspondingly.
| Model | MAE for energy prediction |
||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Test ST | Test SF | Test CF | |||||||||||||
| 2k | 5k | 10k | 100k | full train | 2k | 5k | 10k | 100k | full train | 2k | 5k | 10k | 100k | full train | |
| LR | 4.6 | 4.7 | 4.7 | 4.7 | - | 4.6 | 4.7 | 4.7 | 4.7 | - | 4.0 | 4.2 | 4.0 | 4.0 | - |
| SchNet | 151.8 | 66.1 | 29.6 | - | - | 126.5 | 68.3 | 27.4 | - | - | 79.1 | 67.3 | 21.4 | - | - |
| SchNOrb | 5.9 | 3.7 | 13.3(*) | - | - | 5.9 | 3.4 | 14.8(*) | - | - | 5.0 | 3.6 | 14.5(*) | - | - |
| DimeNet++ | 24.1 | 21.1 | 10.6 | 3.2 | - | 21.6 | 20.9 | 10.1 | 3.0 | - | 18.3 | 33.7 | 5.2 | 2.5 | - |
| PAINN | 137.2 | 62.8 | 13.1 | 7.0 | - | 131.1 | 53.0 | 12.6 | 6.7 | - | 134.4 | 50.0 | 12.1 | 7.0 | - |
| Model | MAE for Hamiltonian matrix prediction |
MAE for overlap matrix prediction |
||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Test ST | Test SF | Test CF | Test ST | Test SF | Test CF | |||||||||||||||||||||||||
| 2k | 5k | 10k | 100k | full train | 2k | 5k | 10k | 100k | full train | 2k | 5k | 10k | 100k | full train | 2k | 5k | 10k | 100k | full train | 2k | 5k | 10k | 100k | full train | 2k | 5k | 10k | 100k | full train | |
| SchNOrb | 386.5 | 383.4 | 382.0(*) | - | - | 385.3 | 380.7 | 383.6(*) | - | - | 385.0 | 384.8 | 392.0(*) | - | - | 1550 | 1455 | 1493(*) | - | - | 1543 | 1440 | 1496(*) | - | - | 1544 | 1480 | 1536(*) | - | - |
| PhiSNet | 7.4 | 3.2 | 2.9 | - | - | 7.2 | 3.2 | 2.9 | - | - | 6.5 | 3.2 | 2.8 | - | - | 5.1 | 4.3 | 3.5 | - | - | 5.0 | 4.3 | 3.5 | - | - | 5.1 | 4.6 | 3.6 | - | - |
Fields with - or * symbols correspond to the models, which haven't converged and will be updated in the future.
If you are using nablaDFT in your research paper, please cite us as
@article{10.1039/D2CP03966D,
author ="Khrabrov, Kuzma and Shenbin, Ilya and Ryabov, Alexander and Tsypin, Artem and Telepov, Alexander and Alekseev, Anton and Grishin, Alexander and Strashnov, Pavel and Zhilyaev, Petr and Nikolenko, Sergey and Kadurin, Artur",
title ="nablaDFT: Large-Scale Conformational Energy and Hamiltonian Prediction benchmark and dataset",
journal ="Phys. Chem. Chem. Phys.",
year ="2022",
volume ="24",
issue ="42",
pages ="25853-25863",
publisher ="The Royal Society of Chemistry",
doi ="10.1039/D2CP03966D",
url ="http://dx.doi.org/10.1039/D2CP03966D"}