The mcstatsim package offers an efficient, functional
programming-based approach for statistical simulations, centralizing the
process in a single higher-order function for better manageability.
Besides, it includes ready-to-use functions for well-known simulation
targets.
The core runsim() function processes simulation parameters via
expanded grid, mapping them to the simulation function.
Replication management is performed through the replicater() function,
which estimates the remaining time (ETA) with an option to hide progress
via show_progress = FALSE. Outputs are deliberately structured as
dataframe to simplify analysis and visualization, addressing the
limitations of list outputs in data manipulation.
You can install the latest development version of mcstatsim from
gitHub:
# install.packages("devtools")
#devtools::install_github("ielbadisy/mcstatsim")Here is a basic example to get you started with mcstatsim:
library(mcstatsim)
# Define a simple simulation function
sim_function <- function(a, b) {
Sys.sleep(0.1) # Simulate a time-consuming process
return(data.frame(result = a + b))
}
# Generate a grid of parameters
params <- expand.grid(a = 1:3, b = 4:6)
# Run simulations
results <- runsim(n = 3, grid_params = params, sim_func = sim_function)This example demonstrates how to define a simple simulation function,
create a grid of parameters for the simulation, and run the simulations
in parallel using mcstatsim.
To illustrate the utility of this package in a concrete example, we will
simulate the evaluation of several imputation methods to assess their
effectiveness in preserving the accuracy of coefficient estimates in a
Cox regression model. This will demonstrate the creation of a simulation
function, setting up parameters, and using mcstatsim to run these
simulations in parallel.
Simulation aim: We will to assess the performance of some imputation methods regarding their capacity to preserve the values of coefficients estimates. For this aim, we well set up the following simulation design:
-
Generate fully observed data ->
data_complete -
estimate the beta coefficients values from
data_complete -
Introduce missigness under MCAR to complete dataset generated in (1) ->
data_missing -
impute the dataset generated at (3) ->
data_imputed -
Use the following simulation targets to compute the distortion between beta from
data_completeand betadata_imputed:
- Bias: Bias measures the average deviation of the estimated coefficients from the true coefficients.
- Coverage: Coverage is the proportion of times the true coefficient value falls within the estimated confidence interval across all simulations.
- Mean Squared Error (MSE): MSE combines both the variance of the estimator and its bias, providing a single measure of estimation quality.
NB: All these metrics are already implemented in the mcstatsim
package (see ?calc_bias(), ?calc_coverage(), and ?calc_rmse().
Since we want to preserve the spirit of the functional prgramming style, all our simulation design step will be decomposed as helper functions. In others words, the simulation steps will be translated to functions as follow:
pacman::p_load(mcstatsim, survival, dplyr, ggplot2)
## (1) generate fully observed data -> `data_complete`
gencox <- function(n = 300, maxTime = 7, logHR = 0.5) {
lambda <- 0.1
rho <- 1.6
rateC <- 0.09
# covariates
x1 <- rnorm(n)
x2 <- x1^2 + x1 + runif(n)
x3 <- rbinom(n, 1, 0.5)
# estimated survival times
U <- runif(n)
Tlat <- (-log(U) / (lambda * exp(logHR * (x1 + x2 + x3))))^(1 / rho)
Ctimes <- rexp(n, rate = rateC)
# follow-up times and event indicators
time <- pmin(Tlat, Ctimes)
status <- as.numeric(Tlat <= Ctimes)
time <- ifelse(time > maxTime, maxTime, time)
status <- ifelse(time >= maxTime, 1, status)
data <- data.frame(time, status, x1, x2, x3)
data$x3 <- as.factor(data$x3)
return(data)
}
## (2) estimate the beta coefficients values from `data_complete`
estimate_coxest <- function(data) {
myFormula <- survival::Surv(time, status) ~ x1 + x2 + x3
coefs <- summary(survival::coxph(myFormula, data = data))$coef
coefs[, 1]
}
## (3) introduce missigness under MCAR to complete dataset generated in (1) -> `data_missing`
introduce_MCAR <- function(x, covariates = names(x), p = 0.3) {
stopifnot(is.data.frame(x), p >= 0 && p <= 1, all(covariates %in% names(x)))
x[covariates] <- lapply(x[covariates], function(z) {
z[sample(length(z), floor(p * length(z)))] <- NA
z
})
return(x)
}
## (4) impute the dataset generated at (3) -> `data_imputed`
imputer <- function(data, method) {
stopifnot(is.data.frame(data))
if (is.factor(method)) {method <- as.character(method)}
supported_methods <- c("knn", "cart", "missforest", "missranger", "misscforest", "complete")
stopifnot(method %in% supported_methods)
data_imputed <- switch(method,
knn = VIM::kNN(data)[names(data)],
cart = simputation::impute_cart(data, .~.),
missforest = missForest::missForest(data, xtrue = data, verbose = FALSE)$ximp,
missranger = missRanger::missRanger(data, pmm.k = 5, num.trees = 100, verbose = 0),
misscforest = suppressWarnings(missCforest::missCforest(data, ntree = 10L)),
complete = data[stats::complete.cases(data), ])
return(data_imputed)
}
## (5) compute the simulation targets
evaluate_coxest <- function(data, truelogHR) {
myFormula <- survival::Surv(time, status) ~ x1 + x2 + x3
coefs <- summary(survival::coxph(myFormula, data = data))$coef
estimates <- as.data.frame(coefs)
out <- data.frame(
estimates = estimates$coef,
ci_lower = estimates$coef - 1.96 * estimates$`se(coef)`,
ci_upper = estimates$coef + 1.96 * estimates$`se(coef)`
)
out$bias <- mcstatsim::calc_bias(out$estimates, truelogHR)$bias
out$coverage <- mcstatsim::calc_coverage(out$ci_lower, out$ci_upper, truelogHR)$coverage
out$rmse <- mcstatsim::calc_rmse(out$estimates, truelogHR)$rmse
return(out)
}simcox <- function(n, logHR, pmiss, covariates = c("x2"), method = method){
data_complete <- gencox(n = n)
truelogHR <- estimate_coxest(data_complete)
data_missing <- introduce_MCAR(data_complete, covariates = covariates, p = pmiss)
data_imputed <- imputer(data_missing, method = method)
res_est <- evaluate_coxest(data_missing, truelogHR = truelogHR)
res <- cbind(n, pmiss, method, covariates, res_est, row.names = NULL)
return(res)
}Now, to link our simulation function to the simulation parameters, we will generate a grid of paremeters with the same names as the argument of simulation function:
params <- expand.grid(n = c(200, 500),
logHR = 0.5,
pmiss = c(0.2, 0.5),
method = c("knn", "cart", "missforest", "missranger",
"misscforest", "complete"))In one line of code, we can lunch our simulation:
pacman::p_load(survival, missRanger, missCforest, missForest, VIM, simputation, ggplot2, tidyr)
sim_res <- mcstatsim::runsim(n = 3, grid_params = params, sim_func = simcox)Finaly, we retreive all the results in one single dataset (i.e
sim_res) for ploting and table production:
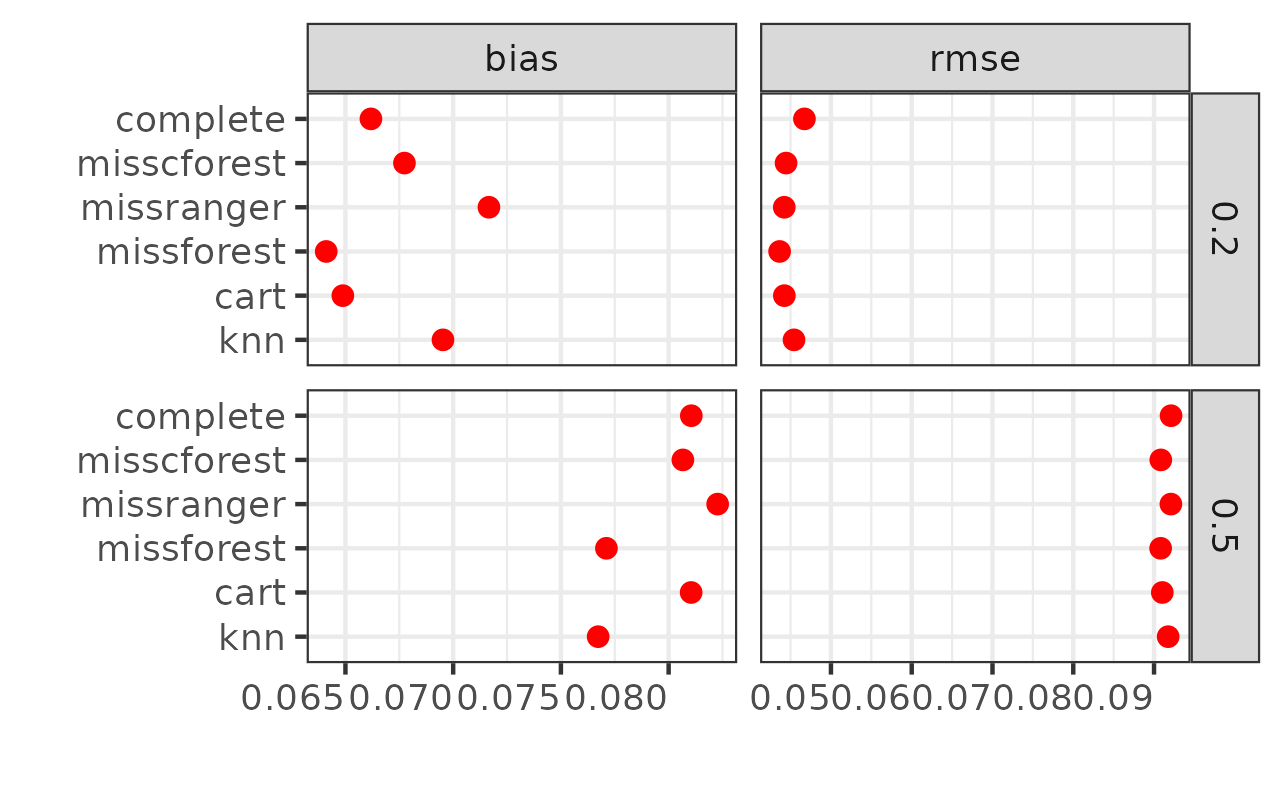
sim_res$bias <- abs(sim_res$bias)
sim_res2 <- gather(sim_res, metric, value, c(bias, rmse))
ggplot(sim_res2, aes(x=value, y=method, fill=method)) +
stat_summary(fun=mean, geom="point", shape=20, size=3, color="red") +
theme_bw() +
theme(axis.text.x = element_text(hjust = 1),
legend.position = "none") +
labs(x = "", y = "") +
facet_grid(pmiss~metric, scales="free")
#ggsave("quick_res.png")library(dplyr)
summary_table <- sim_res %>%
group_by(n, pmiss, method) %>% # Group by number of observations, missingness, and method
summarise(
Mean_Estimate = mean(estimates),
Mean_CI_Lower = mean(ci_lower),
Mean_CI_Upper = mean(ci_upper),
Mean_Bias = mean(bias),
Coverage = mean(coverage),
Mean_RMSE = mean(rmse),
.groups = 'drop'
)
knitr::kable(summary_table, format = "markdown", caption = "Summary of Simulation Results")Let’s skip interpreting the results since the simulation design isn’t
complete yet—we need to add more simulation targets and assess different
hyperparameter values. However, this provides a good demonstration of
how the mcstatsim package can efficiently organize Monte Carlo
simulations without the complexities of for-loops and managing numerous
parameters.
-
No Dependencies:
mcstatsimis designed to be lightweight and standalone, requiring no additional packages for its core functionalities, simplifying installation and usage. -
Functional programming approach: Streamlines the process of setting up and running simulations.
-
Parallel execution: Leverages multiple cores to speed up the execution of simulations.
-
Structured output: Returns simulation results in a dataframe, facilitating quick analysis and visualization.
Contributions are welcome! If you’d like to help improve mcstatsim,
please feel free to submit a pull request.