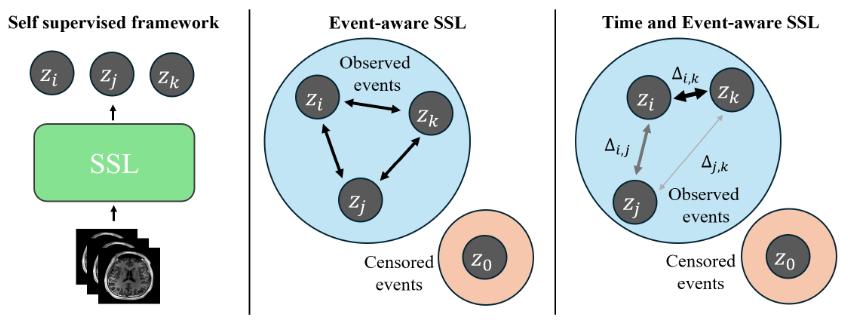
This repository contains supplementary code for Time and Event-Aware Self-Supervised Learning for Alzheimer's Disease Progression Analysis, published in The Medical Image Computing and Computer Assisted Intervention Society (MICCAI) 2024. In this work, we analyze the effect of incorporating time and event labels to a self-supervised pretraining pipeline for survival analysis of Alzheimer's Dementia.
Figure: Schematic diagram of the proposed time- and event-aware SSL, where
Usage:
from lib.Loss import TESSL_Loss
# alpha and beta define maximum and minimum weight
loss_fn = TESSL_loss(alpha=1, beta=0.5)
# features in [batch_size, n_views, dim]
# events, times in [batch_size]
tessl_loss = loss_fn(features, labels=events, times=times)
# Compute SupCon by excluding times
supcon_loss = loss_fn(features, labels=events, times=None)
# Compute SimCLR by exluding events and times
simclr_loss = loss_fn(features, labels=None, times=None)Our data consists of a cohort of 493 unique patients from the ADNI dataset (link). Specific data splits can be found in datasets/files folder. Images were preprocessed via Clinica according to process define in "Generalizable deep learning model for early Alzheimer's Disease detection from structural MRIs". Refer to here for more details
Comparison against regular SSL, Event-Aware SSL and No Pretaining baseline. Results averaged across 3 seeds
| Method | C-td | IBS |
|---|---|---|
| No Pretraining | 0.7329 | 0.2099 |
| SSL | 0.7511 | 0.1985 |
| E-SSL | 0.7720 | 0.1997 |
| TE-SSL | 0.7873 | 0.1889 |
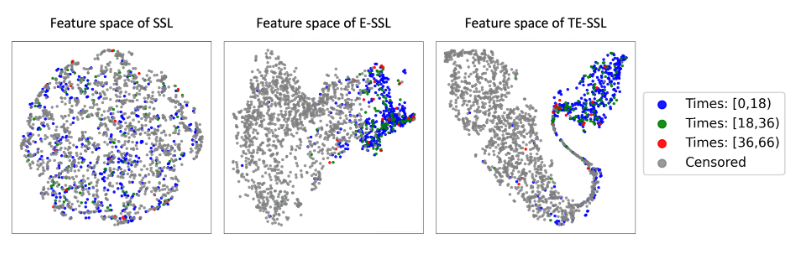
t-SNE visualization demonstrates superior seperability for TE-SSL embeddings compared to SSL and E-SSL.
Figure: t-SNE analysis of feature representations captured by the projection head across different SSL frameworks. Individual points, if nto censored, are labeld with different time-to-event groups