I usually take dinner with something to watch, here it comes your video, really interesting and I personally wanted to test out some ideas on how to improve it. More than a pull request, this is an abstract on my journey testing out a couple of ideas based in the base code:
So the first idea is very simple: what if we use a cache system, to store all
the distances we are calculating so there is no repetition and reuse the
function calls?
cache = {}
def wagner_fischer_dict(s1, s2):
len_s1, len_s2 = len(s1), len(s2)
if len_s1 > len_s2:
s1, s2 = s2, s1
len_s1, len_s2 = len_s2, len_s1
# Check if the distance is already calculated
if (s1, s2) in cache:
return cache[(s1, s2)]
current_row = range(len_s1 + 1)
for i in range(1, len_s2 + 1):
previous_row, current_row = current_row, [i] + [0] * len_s1
for j in range(1, len_s1 + 1):
add, delete, change = previous_row[j] + \
1, current_row[j-1] + 1, previous_row[j-1]
if s1[j-1] != s2[i-1]:
change += 1
current_row[j] = min(add, delete, change)
# Store the distance in the cache
cache[(s1, s2)] = current_row[len_s1]
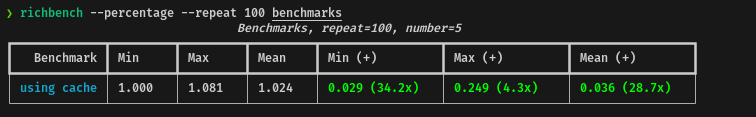
return current_row[len_s1]Using the richbench tool, I've tested the performance of the cache system
against the base code, these are the results:
In average, using the cache system, the performance will be almost 29x times faster, which makes sense. However, I will not call this a win, because there is a trade-off: the memory usage, depending on the input, can be considerable and in case of a large input this can be a problem.
The second approach is taken from an article of Fujimoto Seiji
called Can We Optimize the Wagner-Fischer Algorithm? where he proposes a
couple of ideas to improve the Wagner-Fischer algorithm, for example, using a
one-dimensional array instead of a two-dimensional array which is already
implemented in the base code.
The technique is called Ukkonen’s optimization and it's based on the idea of
cutting off branches using upper bounds. The idea is to stop the computation of
a row if the minimum possible distance is greater than the threshold.
def wagner_fischer_branch(s1, s2):
len_s1, len_s2 = len(s1), len(s2)
if len_s1 > len_s2:
s1, s2 = s2, s1
len_s1, len_s2 = len_s2, len_s1
if len_s2 == 0:
return len_s1
if len_s2 == 1:
return len_s1 - (s2[0] in s1)
if len_s2 == 2:
p1 = s1.find(s2[0])
if p1 != -1:
return len_s1 - (s2[1] in s1[p1+1:]) - 1
else:
return len_s1 - (s2[1] in s1[1:])
buf = [i for i in range(len_s2 + 1)]
Mx = (len_s2 - 1) // 2
mx = 1 - Mx - (len_s1 - len_s2)
for j in range(Mx + 1):
buf[j] = j
for i in range(1, len_s1 + 1):
buf[0] = i - 1
m = max(mx, 1)
M = min(Mx, len_s2)
mx += 1
Mx += 1
dia = buf[m - 1]
top = buf[m]
if s1[i - 1] != s2[m - 1]:
dia = min(dia, top) + 1
buf[m] = dia
left = dia
dia = top
for j in range(m + 1, M + 1):
top = buf[j]
if s1[i - 1] != s2[j - 1]:
dia = min(min(dia, top), left) + 1
buf[j] = dia
left = dia
dia = top
if len_s2 == M:
continue
if s1[i - 1] != s2[M]:
dia = min(dia, left) + 1
buf[M + 1] = dia
dia = buf[len_s2]
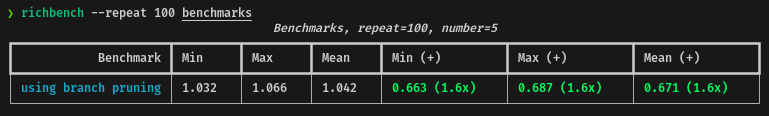
return diaTesting the performance of the branch pruning against the base code, these are the results:
In average, using the branch pruning, the performance will be 1.6x times faster for the current dataset.
Finally, I wanted to test both approaches together. My naive idea was to use the same approach added in the base code, but using a different cache for this method itself since we don't want to give any advantage in the performance test.
# 3. Use the cache and branch pruning together
cache_for_branch = {}
def wagner_fischer_branch_cache(s1, s2):
if (s1, s2) in cache_for_branch:
return cache_for_branch[(s1, s2)]
len_s1, len_s2 = len(s1), len(s2)
if len_s1 > len_s2:
s1, s2 = s2, s1
len_s1, len_s2 = len_s2, len_s1
if len_s2 == 0:
return len_s1
if len_s2 == 1:
return len_s1 - (s2[0] in s1)
if len_s2 == 2:
p1 = s1.find(s2[0])
if p1 != -1:
return len_s1 - (s2[1] in s1[p1+1:]) - 1
else:
return len_s1 - (s2[1] in s1[1:])
buf = [i for i in range(len_s2 + 1)]
Mx = (len_s2 - 1) // 2
mx = 1 - Mx - (len_s1 - len_s2)
for j in range(Mx + 1):
buf[j] = j
for i in range(1, len_s1 + 1):
buf[0] = i - 1
m = max(mx, 1)
M = min(Mx, len_s2)
mx += 1
Mx += 1
dia = buf[m - 1]
top = buf[m]
if s1[i - 1] != s2[m - 1]:
dia = min(dia, top) + 1
buf[m] = dia
left = dia
dia = top
for j in range(m + 1, M + 1):
top = buf[j]
if s1[i - 1] != s2[j - 1]:
dia = min(min(dia, top), left) + 1
buf[j] = dia
left = dia
dia = top
if len_s2 == M:
continue
if s1[i - 1] != s2[M]:
dia = min(dia, left) + 1
buf[M + 1] = dia
dia = buf[len_s2]
cache_for_branch[(s1, s2)] = dia
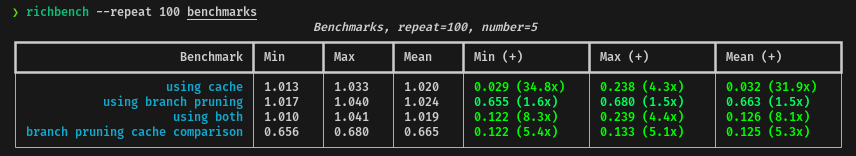
return diaI tested the performance of the branch pruning with the cache system against the base code and the branch pruning itself. Here are the final results:
In average, using the branch pruning with the cache system, the performance will
be up to 4.4x times faster for the current dataset, which is great but not so
much.
We can also check that the cache system improves up to 5x times the
performance of the branch pruning approach, but, as I said before, the memory
usage is something to be aware of.