.__
_____ | | _____
\__ \ | | / \
/ __ \| |_| Y Y \
(____ /____/__|_| /
\/ \/
UPDATE: Check out the v6 branch for a version of this package to work with the newest Lagotto API.
The alm package is a set of functions to access article level metrics via a RESTful API from the Rails app Lagotto created by the Public Library of Science (PLOS). Lagotto is being used by PLOS, and a number of other publishers:
- PLOS (all their journals) at http://alm.plos.org/
- PLOS test server at http://labs.lagotto.io/
- Copernicus (seems to be down for now)
- Public Knowledge Project (PKP) at http://pkp-alm.lib.sfu.ca/
- Crossref at http://det.labs.crossref.org/
- eLife at http://lagotto.svr.elifesciences.org/
- Pensoft at http://alm.pensoft.net:81/
- Making Dat Count at http://dlm.plos.org
A good place to look for the status of various installations of Lagotto is this status page: http://articlemetrics.github.io/status/ (which also includes what version of Lagotto each is running)
Lagotto has a nice support site at http://discuss.lagotto.io/ for any questions about it.
Glad you asked. The canonical URL for this is perhaps altmetrics.org. Basically it is a metric that measures something about an article. This is in stark contrast to journal level metrics, like the Journal Impact Factor.
Yes indeedy, but see notes
- ImpactStory - No open data (except for your own data)
- Altmetric.com - Some open data
- PlumAnalytics - No open data
You only need an API key for the publishers PKP and Pensoft. You can set the key in your options just for the current session by executing options(PlosApiKey = "YOUalmAPIKEY"), or pass in to each function call with the key parameter, or save in your .Rprofile file.
The default URL is set for the PLOS data sources: http://alm.plos.org/api/v5/articles
You can change this URL. For example, if you want to get data from the Crossref instance, set the api_url parameter to http://alm.labs.crossref.org/api/v5/articles
If R is not your thing, there are Lagotto clients in development for Ruby and Python.
You can get this package from CRAN by:
install.packages("alm")Or the development version by installing with install_github()
install.packages("devtools")
devtools::install_github("ropensci/alm")Load the package
library('alm')The first function we'll look at is alm_ids(), named with ids since we search for data by one of four different choices of identifier.
alm_ids(doi = "10.1371/journal.pone.0029797")
#> $meta
#> total total_pages page error
#> 1 1 1 1 NA
#>
#> $data
#> .id pdf html readers comments likes total
#> 1 citeulike NA NA 1 NA NA 1
#> 2 crossref NA NA NA NA NA 8
#> 3 nature NA NA NA NA NA 4
#> 4 pubmed NA NA NA NA NA 2
#> 5 scopus NA NA NA NA NA 9
#> 6 counter 2583 31976 NA NA NA 34683
#> 7 researchblogging NA NA NA NA NA 1
#> 8 pmc 81 551 NA NA NA 632
#> 9 facebook NA NA 150 22 60 232
#> 10 mendeley NA NA 86 NA NA 86
#> 11 twitter NA NA NA 12 NA 12
#> 12 wikipedia NA NA NA NA NA 50
#> 13 scienceseeker NA NA NA NA NA 0
#> 14 relativemetric NA NA NA NA NA 243342
#> 15 f1000 NA NA NA NA NA 0
#> 16 figshare 0 31 NA NA 0 31
#> 17 pmceurope NA NA NA NA NA 4
#> 18 pmceuropedata NA NA NA NA NA 49
#> 19 openedition NA NA NA NA NA 0
#> 20 wordpress NA NA NA NA NA 0
#> 21 reddit NA NA NA 0 0 0
#> 22 datacite NA NA NA NA NA 0
#> 23 copernicus NA NA NA NA NA 0
#> 24 articlecoverage NA NA NA 0 NA 0
#> 25 articlecoveragecurated NA NA NA 0 NA 0
#> 26 plos_comments NA NA NA 11 NA 16
#> 27 orcid NA NA 0 NA NA 0alm_ids(doi = "10.1371/journal.pone.0029797", info = "detail")
#> $meta
#> total total_pages page error
#> 1 1 1 1 NA
#>
#> $data
#> $data$info
#> id
#> 1 doi/10.1371/journal.pone.0029797
#> title
#> 1 Ecological Guild Evolution and the Discovery of the World's Smallest Vertebrate
#> publisher_id doi
#> 1 340 10.1371/journal.pone.0029797
#> canonical_url
#> 1 http://www.plosone.org/article/info:doi/10.1371/journal.pone.0029797
#> pmid pmcid scp wos update_date
#> 1 22253785 3256195 84855712734 000301355700052 2015-03-13T09:10:19Z
#> issued
#> 1 2012-01-11
#>
#> $data$signposts
#> id viewed saved discussed cited
#> 1 doi/10.1371/journal.pone.0029797 35315 87 244 9
#>
#> $data$totals
#> .id pdf html readers comments likes total
#> 1 citeulike NA NA 1 NA NA 1
#> 2 crossref NA NA NA NA NA 8
#> 3 nature NA NA NA NA NA 4
#> 4 pubmed NA NA NA NA NA 2
#> 5 scopus NA NA NA NA NA 9
#> 6 counter 2583 31976 NA NA NA 34683
#> 7 researchblogging NA NA NA NA NA 1
#> 8 pmc 81 551 NA NA NA 632
#> 9 facebook NA NA 150 22 60 232
#> 10 mendeley NA NA 86 NA NA 86
#> 11 twitter NA NA NA 12 NA 12
#> 12 wikipedia NA NA NA NA NA 50
#> 13 scienceseeker NA NA NA NA NA 0
#> 14 relativemetric NA NA NA NA NA 243342
#> 15 f1000 NA NA NA NA NA 0
#> 16 figshare 0 31 NA NA 0 31
#> 17 pmceurope NA NA NA NA NA 4
#> 18 pmceuropedata NA NA NA NA NA 49
#> 19 openedition NA NA NA NA NA 0
#> 20 wordpress NA NA NA NA NA 0
#> 21 reddit NA NA NA 0 0 0
#> 22 datacite NA NA NA NA NA 0
#> 23 copernicus NA NA NA NA NA 0
#> 24 articlecoverage NA NA NA 0 NA 0
#> 25 articlecoveragecurated NA NA NA 0 NA 0
#> 26 plos_comments NA NA NA 11 NA 16
#> 27 orcid NA NA 0 NA NA 0
#>
#> $data$sum_metrics
#> .id year month day total X[[1L]]
#> 1 citeulike 2012 1 12 1 NA
#> 2 nature 2012 1 11 1 NA
#> 3 nature 2012 1 12 1 NA
#> 4 nature 2012 2 1 1 NA
#> 5 researchblogging NA NA NA NA NA
#> 6 plos_comments 2012 1 11 1 NA
#> 7 plos_comments 2012 1 12 7 NA
#> 8 plos_comments 2012 1 13 1 NA
#> 9 plos_comments 2012 1 14 1 NAdois <- c("10.1371/journal.pone.0001543", "10.1371/journal.pone.0040117", "10.1371/journal.pone.0029797", "10.1371/journal.pone.0039395")
out <- alm_ids(doi = dois)
lapply(out$data, head)
#> $`10.1371/journal.pone.0040117`
#> .id pdf html readers comments likes total
#> 1 citeulike NA NA 0 NA NA 0
#> 2 crossref NA NA NA NA NA 7
#> 3 nature NA NA NA NA NA 0
#> 4 pubmed NA NA NA NA NA 6
#> 5 scopus NA NA NA NA NA 14
#> 6 counter 439 2312 NA NA NA 2780
#>
#> $`10.1371/journal.pone.0039395`
#> .id pdf html readers comments likes total
#> 1 citeulike NA NA 0 NA NA 0
#> 2 crossref NA NA NA NA NA 1
#> 3 nature NA NA NA NA NA 0
#> 4 pubmed NA NA NA NA NA 1
#> 5 scopus NA NA NA NA NA 4
#> 6 counter 384 1678 NA NA NA 2095
#>
#> $`10.1371/journal.pone.0029797`
#> .id pdf html readers comments likes total
#> 1 citeulike NA NA 1 NA NA 1
#> 2 crossref NA NA NA NA NA 8
#> 3 nature NA NA NA NA NA 4
#> 4 pubmed NA NA NA NA NA 2
#> 5 scopus NA NA NA NA NA 9
#> 6 counter 2583 31976 NA NA NA 34683
#>
#> $`10.1371/journal.pone.0001543`
#> .id pdf html readers comments likes total
#> 1 citeulike NA NA 0 NA NA 0
#> 2 crossref NA NA NA NA NA 7
#> 3 nature NA NA NA NA NA 0
#> 4 pubmed NA NA NA NA NA 7
#> 5 scopus NA NA NA NA NA 12
#> 6 counter 472 2809 NA NA NA 3332res <- alm_ids(doi = "10.1371/journal.pone.0035869", total_details = TRUE)
res$data[, 3:10]
#> citeulike_pdf citeulike_html citeulike_readers citeulike_comments
#> 1 NA NA 27 NA
#> citeulike_likes citeulike_total crossref_pdf crossref_html
#> 1 NA 27 NA NAout <- alm_events(doi = "10.1371/journal.pone.0029797")
names(out) # names of sources
#> [1] "citeulike" "crossref" "nature"
#> [4] "pubmed" "scopus" "counter"
#> [7] "researchblogging" "pmc" "facebook"
#> [10] "mendeley" "twitter" "wikipedia"
#> [13] "relativemetric" "figshare" "pmceuropedata"
#> [16] "plos_comments"out <- out[!out %in% c("sorry, no events content yet", "parser not written yet")] # remove those with no data
out[["pmc"]] # get the results for PubMed Central
#> $events_url
#> [1] "http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3256195"
#>
#> $events
#> scanned.page.browse month cited.by abstract full.text unique.ip pdf
#> 1 0 1 0 1 51 42 8
#> 2 0 2 0 0 15 11 4
#> 3 0 3 0 0 11 12 4
#> 4 0 4 0 1 6 6 2
#> 5 0 5 0 0 5 5 1
#> 6 0 6 0 0 7 9 2
#> 7 0 7 0 1 6 8 3
#> 8 0 8 0 1 5 4 0
#> 9 0 9 0 0 14 13 5
#> 10 0 10 0 1 20 16 4
#> 11 0 12 0 0 13 12 1
#> 12 0 1 0 0 13 14 7
#> 13 0 3 0 1 22 20 2
#> 14 0 2 0 0 13 10 2
#> 15 0 4 1 1 45 24 4
#> 16 0 11 0 1 10 9 1
#> 17 0 5 0 0 18 21 5
#> 18 0 6 0 0 12 11 1
#> 19 0 7 0 0 27 14 1
#> 20 0 8 0 0 21 13 0
#> 21 0 9 0 0 14 13 0
#> 22 0 10 0 0 14 15 3
#> 23 0 11 0 6 18 16 2
#> 24 0 12 0 0 14 9 1
#> 25 0 1 0 0 22 16 1
#> 26 0 2 0 0 13 12 1
#> 27 0 3 0 0 11 10 1
#> 28 0 5 0 0 15 12 0
#> 29 0 6 0 0 12 12 2
#> 30 0 4 0 0 8 9 1
#> 31 0 8 0 0 18 14 3
#> 32 0 9 0 0 18 20 2
#> 33 0 10 0 0 23 18 4
#> 34 0 11 0 0 17 16 3
#> year figure scanned.summary supp.data
#> 1 2012 9 0 0
#> 2 2012 11 0 2
#> 3 2012 0 0 0
#> 4 2012 0 0 0
#> 5 2012 0 0 0
#> 6 2012 2 0 0
#> 7 2012 3 0 0
#> 8 2012 0 0 0
#> 9 2012 3 0 0
#> 10 2012 1 0 0
#> 11 2012 1 0 0
#> 12 2013 0 0 0
#> 13 2013 0 0 0
#> 14 2013 0 0 0
#> 15 2013 3 0 1
#> 16 2012 0 0 0
#> 17 2013 0 0 1
#> 18 2013 0 0 1
#> 19 2013 0 0 0
#> 20 2013 0 0 0
#> 21 2013 0 0 1
#> 22 2013 2 0 0
#> 23 2013 0 0 0
#> 24 2013 0 0 0
#> 25 2014 0 0 0
#> 26 2014 0 0 0
#> 27 2014 2 0 0
#> 28 2014 0 0 1
#> 29 2014 0 0 0
#> 30 2014 0 0 1
#> 31 2014 0 0 0
#> 32 2014 0 0 0
#> 33 2014 0 0 0
#> 34 2014 2 0 0
#>
#> $csl
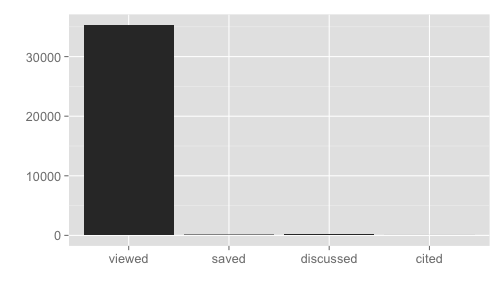
#> list()dat <- alm_signposts(doi = "10.1371/journal.pone.0029797")
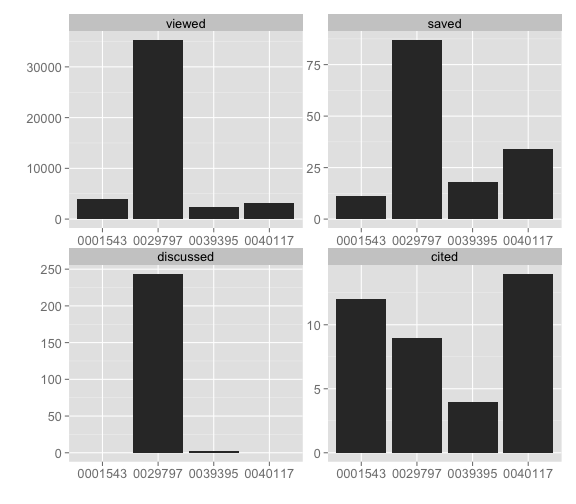
plot_signposts(dat)Or plot many identifiers get a faceted bar chart, note the tick labels have just the last part of the DOI in this case to help you identify each DOI - printing the entire DOI would make lables overlap one another.
dois <- c("10.1371/journal.pone.0001543", "10.1371/journal.pone.0040117", "10.1371/journal.pone.0029797", "10.1371/journal.pone.0039395")
dat <- alm_signposts(doi = dois)
plot_signposts(input = dat)Or make an interactive chart by doing plot_signposts(input=dat, type="multiBarChart"). Try it out! It should open in your browser and you can interact with it.
Note: Do you the key below in the searchplos call in this example, but if you plan to use rplos more, get your own API key here.
library('rplos')
library('plyr')
dois <- searchplos(q = "science", fl = "id", fq = list("-article_type:correction", "cross_published_journal_key:PLoSONE", "doc_type:full", "publication_date:[2010-01-01T00:00:00Z TO 2010-12-31T23:59:59Z]"), limit = 50)
dois <- dois$data$idCollect altmetrics data and combine to a data.frame with ldply
alm <- alm_ids(doi = dois, total_details = TRUE)
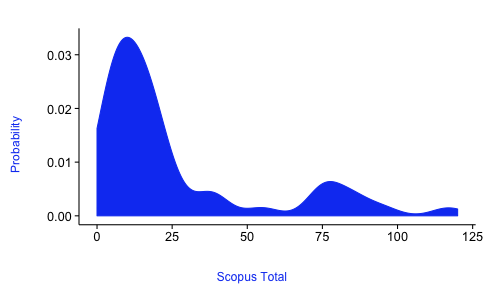
alm <- ldply(alm$data)The default plot
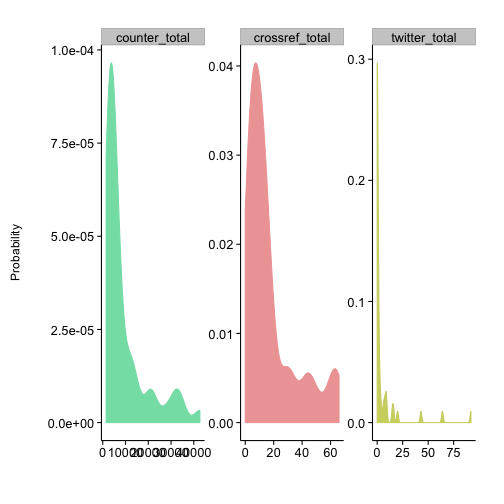
plot_density(alm)Plot many sources in different panels in the same plot, and pass in colors just for fun
plot_density(alm,
source = c("counter_total", "crossref_total", "twitter_total"),
color = c("#83DFB4", "#EFA5A5", "#CFD470"))Crossref
crurl <- "http://det.labs.crossref.org/api/v5/articles"
crkey <- getOption("crossrefalmkey")
alm_ids(doi='10.1371/journal.pone.0086859', api_url = crurl, key = crkey)eLife
elifeurl <- "http://alm.svr.elifesciences.org/api/v5/articles"
elifekey <- getOption("elifealmkey")
alm_ids(doi='10.7554/eLife.00471', api_url = elifeurl, key = elifekey)
#> $meta
#> total total_pages page error
#> 1 1 1 1 NA
#>
#> $data
#> .id pdf html shares groups comments likes citations total
#> 1 pmc 667 1520 NA NA NA NA NA 2187
#> 2 crossref NA NA NA NA NA NA 154 154
#> 3 scopus NA NA NA NA NA NA 154 154
#> 4 facebook NA NA 3 NA 0 0 NA 3
#> 5 mendeley NA NA 0 0 NA NA NA 0
#> 6 twitter_search NA NA NA NA 0 NA NA 0
#> 7 citeulike NA NA 1 NA NA NA NA 1
#> 8 pubmed NA NA NA NA NA NA 111 111
#> 9 wordpress NA NA NA NA NA NA 0 0
#> 10 reddit NA NA NA NA 0 0 NA 0
#> 11 wikipedia NA NA NA NA NA NA 1 1
#> 12 datacite NA NA NA NA NA NA 0 0
#> 13 pmceurope NA NA NA NA NA NA 151 151
#> 14 pmceuropedata NA NA NA NA NA NA 1 1
#> 15 scienceseeker NA NA NA NA NA NA 0 0
#> 16 nature NA NA NA NA NA NA 0 0
#> 17 openedition NA NA NA NA NA NA 0 0
#> 18 f1000 NA NA NA NA NA NA 4 4
#> 19 figshare NA NA NA NA NA NA 0 0
#> 20 plos_comments NA NA NA NA NA NA 0 0
#> 21 connotea NA NA NA NA NA NA 0 0
#> 22 postgenomic NA NA NA NA NA NA 0 0
#> 23 bloglines NA NA NA NA NA NA 0 0
#> 24 biod NA NA NA NA NA NA NA 0Pensoft
psurl <- 'http://alm.pensoft.net:81/api/v5/articles'
pskey <- getOption("pensoftalmkey")
alm_ids(doi='10.3897/zookeys.88.807', api_url = psurl, key = pskey)
#> $meta
#> total total_pages page error
#> 1 1 1 1 NA
#>
#> $data
#> .id pdf html readers comments likes total
#> 1 facebook NA NA NA NA NA 0
#> 2 citeulike NA NA 1 NA NA 1
#> 3 mendeley NA NA 123 NA NA 123
#> 4 wordpress NA NA NA NA NA 0
#> 5 datacite NA NA NA NA NA 0
#> 6 wikipedia NA NA NA NA NA 137
#> 7 copernicus 0 0 NA NA NA 0
#> 8 nature NA NA NA NA NA 0
#> 9 f1000 NA NA NA NA NA 0
#> 10 scopus NA NA NA NA NA 192
#> 11 pmceuropedata NA NA NA NA NA 0
#> 12 figshare NA NA NA NA NA 0
#> 13 counter NA NA NA NA NA 0
#> 14 crossref NA NA NA NA NA 81
#> 15 reddit NA NA NA 0 0 0
#> 16 twitter_search NA NA NA 0 NA 0
#> 17 pmceurope NA NA NA NA NA 52
#> 18 pubmed NA NA NA NA NA 51Making Data Count - Dataset altmetrics
dlmurl <- 'http://dlm.plos.org/api/v5/articles'
alm_ids('10.5061/dryad.7fj1k', api_url = dlmurl)
#> $meta
#> total total_pages page error
#> 1 1 1 1 NA
#>
#> $data
#> .id pdf html readers comments likes total
#> 1 citeulike NA NA 0 NA NA 0
#> 2 wordpress NA NA NA NA NA 0
#> 3 wikipedia NA NA NA NA NA 0
#> 4 datacite NA NA NA NA NA 96
#> 5 pmceuropedata NA NA NA NA NA 0
#> 6 mendeley NA NA 0 NA NA 0
#> 7 facebook NA NA 0 0 0 0
#> 8 twitter_search NA NA NA NA NA 0
#> 9 europe_pmc_fulltext NA NA NA NA NA 1
#> 10 plos_fulltext NA NA NA NA NA 1
#> 11 orcid NA NA 0 NA NA 0
#> 12 bmc_fulltext NA NA NA NA NA 0
#> 13 nature_opensearch NA NA NA NA NA 0- Please report any issues or bugs.
- License: MIT
- Get citation information for
almin R doingcitation(package = 'alm')