This package implements various tests, visualizations, and metrics for use with environmental niche models (ENMs) and species distribution models (SDMs).
At present, ENMTools is downloadable from https://github.com/danlwarren/ENMTools. There are multiple ways to download it. The easiest is to use devtools and install from GitHub.
Run the following code from your R console:
install.packages("devtools")
library(devtools)
install_github("danlwarren/ENMTools")
library(ENMTools)
A zipped version of the package is available at https://github.com/danlwarren/ENMTools/archive/master.zip. To install from the zip file, download a copy of it to your system. Once it's finished downloading, type the following (where PATH is the path to the zip file):
install.packages("devtools")
library(devtools)
install_local("PATH")
library(ENMTools)
First we're going to load in some environmental data.
env.files <- list.files(path = "test/testdata/", pattern = "pc", full.names = TRUE)
env <- stack(env.files)
names(env) <- c("pc1", "pc2", "pc3", "pc4")
env <- setMinMax(env)
ENMTools is primarily designed to examine patterns of similarity and difference between ENMs for different species. In order to simplify interactions with the functions in ENMTools, you need to put your data for each of your species into an enmtools.species object. You can create and view an empty enmtools.species object just by typing:
ahli <- enmtools.species()
ahli
##
##
## Range raster not defined.
##
## Presence points not defined.
##
## Background points not defined.
##
## Species name not defined.
You can add data to this object manually:
names(ahli)
## [1] "range" "presence.points" "background.points"
## [4] "models" "species.name"
ahli$species.name <- "ahli"
ahli$presence.points <- read.csv("./test/testdata/ahli.csv")[,2:3]
ahli$range <- background.raster.buffer(ahli$presence.points, 50000, mask = env)
ahli$background.points <- background.points.buffer(points = ahli$presence.points,
radius = 20000, n = 1000, mask = env[[1]])
ahli
##
##
## Range raster:
## class : RasterLayer
## dimensions : 418, 1535, 641630 (nrow, ncol, ncell)
## resolution : 0.008333333, 0.008333333 (x, y)
## extent : -86.90809, -74.11642, 19.80837, 23.2917 (xmin, xmax, ymin, ymax)
## coord. ref. : NA
## data source : in memory
## names : pc1
## values : 1, 1 (min, max)
##
##
##
## Presence points (first ten only):
##
## Longitude Latitude
## ---------- ---------
## -80.0106 21.8744
## -79.9086 21.8095
## -79.8065 21.7631
## -79.8251 21.8095
## -79.8807 21.8374
## -79.9550 21.8374
## -80.3446 22.0136
## -80.2983 21.9951
## -80.1776 21.9023
## -80.1591 21.9673
##
##
## Background points (first ten only):
##
## Longitude Latitude
## ---------- ---------
## -80.14559 22.16254
## -80.35392 22.12087
## -79.87059 22.00420
## -80.05392 22.02087
## -80.40392 22.17087
## -79.97892 21.83754
## -79.97059 22.08754
## -80.08726 22.13754
## -80.25392 22.01254
## -80.31226 21.99587
##
##
## Species name: ahli
Or you can add bits of it when the object is created:
allogus <- enmtools.species(species.name = "allogus",
presence.points = read.csv("./test/testdata/allogus.csv")[,3:4])
allogus$range <- background.raster.buffer(allogus$presence.points, 50000, mask = env)
allogus$background.points <- background.points.buffer(points = allogus$presence.points,
radius = 20000, n = 1000, mask = env[[1]])
allogus
##
##
## Range raster:
## class : RasterLayer
## dimensions : 418, 1535, 641630 (nrow, ncol, ncell)
## resolution : 0.008333333, 0.008333333 (x, y)
## extent : -86.90809, -74.11642, 19.80837, 23.2917 (xmin, xmax, ymin, ymax)
## coord. ref. : NA
## data source : in memory
## names : pc1
## values : 1, 1 (min, max)
##
##
##
## Presence points (first ten only):
##
## Longitude Latitude
## ---------- ---------
## -79.2527 22.2109
## -78.7774 22.2241
## -78.6189 22.2373
## -78.1039 21.1809
## -78.0247 21.1809
## -77.9983 20.9301
## -77.9719 21.7091
## -77.9719 21.5507
## -77.9323 21.6167
## -77.9323 20.7320
##
##
## Background points (first ten only):
##
## Longitude Latitude
## ---------- ---------
## -76.07892 20.12087
## -75.22892 20.49587
## -77.93726 20.93754
## -75.88726 20.23754
## -77.86226 21.17920
## -76.40392 20.37087
## -78.05392 21.28754
## -76.86226 20.72920
## -77.85392 21.72087
## -78.92059 22.25420
##
##
## Species name: allogus
However, ENMTools also contains some sample data. It contains an enmtools.clade object called "iberolacerta.clade", which holds several enmtools.species objects. It also has some low-resolution Worldclim data that we can use to demonstrate functions. We'll pull two of those species out now.
data(iberolacerta.clade)
data(euro.worldclim)
monticola <- iberolacerta.clade$species$monticola
cyreni <- iberolacerta.clade$species$cyreni
env <- euro.worldclim
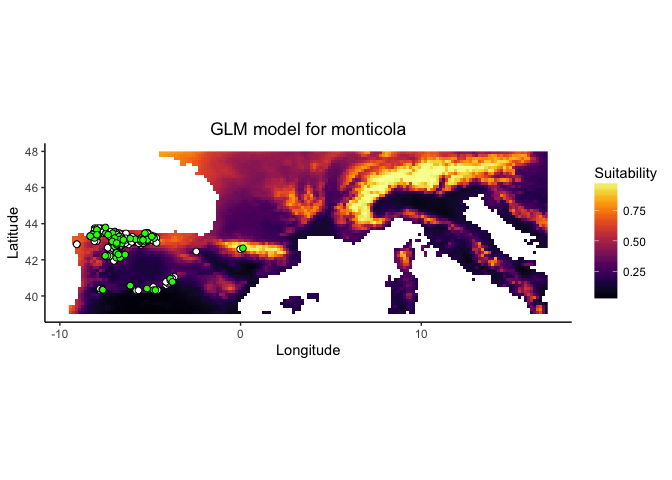
ENMTools contains functions to simplify the ENM construction process. Using enmtools.species objects and the correct modeling commands, we can build models very quickly. These commands are primarily wrappers to dismo model construction and projection functions, and at present are only available for GLM, Maxent, Domain, and Bioclim models. One of the nice bits about this setup is that it allows enmtools to automatically generate suitability maps, do model evaluation, and plot the marginal suitability of habitat for each variable separately.
Before we get started modeling, however, we might want to trim our predictor set so as to reduce collinearity. ENMTools contains a couple of handy functions for this.
raster.cor.matrix(env)
## bio1 bio2 bio3 bio4 bio5
## bio1 1.00000000 0.208715969 0.36766171 -0.21357041 0.8853513
## bio2 0.20871597 1.000000000 0.58362743 0.30918934 0.5417192
## bio3 0.36766171 0.583627431 1.00000000 -0.56601419 0.3052211
## bio4 -0.21357041 0.309189341 -0.56601419 1.00000000 0.1754180
## bio5 0.88535133 0.541719174 0.30522110 0.17541802 1.0000000
## bio6 0.93295069 -0.017231880 0.42521254 -0.52131150 0.6875177
## bio7 -0.05329392 0.709020870 -0.14900115 0.87998888 0.4016817
## bio8 0.24945265 0.068562982 -0.16262988 0.30400935 0.2862188
## bio9 0.73849291 0.183148800 0.42195497 -0.37866158 0.6400464
## bio10 0.96045067 0.311324869 0.21975162 0.06247355 0.9606340
## bio11 0.95921479 0.101719882 0.48107648 -0.47305206 0.7533941
## bio12 -0.60846674 -0.482731496 -0.31488066 -0.07961919 -0.7137513
## bio13 -0.41433559 -0.510317568 -0.25626002 -0.19124801 -0.5722885
## bio14 -0.72871981 -0.338229901 -0.37618609 0.16712793 -0.7297745
## bio15 0.45421260 -0.007931916 0.11838593 -0.21759678 0.3821863
## bio16 -0.44597594 -0.506032477 -0.26995545 -0.16924912 -0.5920746
## bio17 -0.70338945 -0.343806007 -0.33935275 0.10832583 -0.7243068
## bio18 -0.81091065 -0.331213024 -0.47078459 0.28348226 -0.7776879
## bio19 -0.12414066 -0.396445432 0.07348362 -0.48121938 -0.3430162
## bio6 bio7 bio8 bio9 bio10
## bio1 0.93295069 -0.05329392 0.24945265 0.7384929 0.96045067
## bio2 -0.01723188 0.70902087 0.06856298 0.1831488 0.31132487
## bio3 0.42521254 -0.14900115 -0.16262988 0.4219550 0.21975162
## bio4 -0.52131150 0.87998888 0.30400935 -0.3786616 0.06247355
## bio5 0.68751772 0.40168174 0.28621877 0.6400464 0.96063401
## bio6 1.00000000 -0.38884596 0.09285337 0.7734384 0.80726704
## bio7 -0.38884596 1.00000000 0.24603268 -0.1633540 0.20071994
## bio8 0.09285337 0.24603268 1.00000000 -0.2200483 0.33135353
## bio9 0.77343845 -0.16335400 -0.22004834 1.0000000 0.66175815
## bio10 0.80726704 0.20071994 0.33135353 0.6617581 1.00000000
## bio11 0.98896057 -0.29134540 0.12985480 0.7886188 0.84907884
## bio12 -0.48704454 -0.29133189 -0.30622476 -0.4629735 -0.65829840
## bio13 -0.28138429 -0.37121605 -0.19351156 -0.3074562 -0.48860809
## bio14 -0.67564389 -0.07381576 -0.25699492 -0.6265099 -0.71411350
## bio15 0.47445069 -0.11344874 0.17648853 0.3970693 0.41693023
## bio16 -0.31485467 -0.35410914 -0.21747973 -0.3274923 -0.51511693
## bio17 -0.63769389 -0.11473790 -0.30488093 -0.5552129 -0.70368996
## bio18 -0.80126262 0.02381502 0.03291308 -0.8163353 -0.76592247
## bio19 0.07715578 -0.53249335 -0.51356967 0.1072073 -0.26914814
## bio11 bio12 bio13 bio14 bio15
## bio1 0.95921479 -0.60846674 -0.4143356 -0.72871981 0.454212596
## bio2 0.10171988 -0.48273150 -0.5103176 -0.33822990 -0.007931916
## bio3 0.48107648 -0.31488066 -0.2562600 -0.37618609 0.118385927
## bio4 -0.47305206 -0.07961919 -0.1912480 0.16712793 -0.217596785
## bio5 0.75339408 -0.71375135 -0.5722885 -0.72977455 0.382186338
## bio6 0.98896057 -0.48704454 -0.2813843 -0.67564389 0.474450688
## bio7 -0.29134540 -0.29133189 -0.3712161 -0.07381576 -0.113448735
## bio8 0.12985480 -0.30622476 -0.1935116 -0.25699492 0.176488534
## bio9 0.78861884 -0.46297347 -0.3074562 -0.62650986 0.397069264
## bio10 0.84907884 -0.65829840 -0.4886081 -0.71411350 0.416930231
## bio11 1.00000000 -0.54163405 -0.3297284 -0.72423144 0.492885864
## bio12 -0.54163405 1.00000000 0.9064818 0.77619752 -0.243139096
## bio13 -0.32972845 0.90648180 1.0000000 0.48404051 0.130409758
## bio14 -0.72423144 0.77619752 0.4840405 1.00000000 -0.743674332
## bio15 0.49288586 -0.24313910 0.1304098 -0.74367433 1.000000000
## bio16 -0.36513899 0.91789736 0.9902335 0.50120043 0.124449153
## bio17 -0.68356379 0.80791339 0.5245850 0.98592339 -0.729480228
## bio18 -0.82992875 0.76402046 0.5883523 0.84673991 -0.462267795
## bio19 0.01511498 0.74717071 0.7906993 0.31690226 0.128299899
## bio16 bio17 bio18 bio19
## bio1 -0.4459759 -0.7033895 -0.81091065 -0.12414066
## bio2 -0.5060325 -0.3438060 -0.33121302 -0.39644543
## bio3 -0.2699555 -0.3393528 -0.47078459 0.07348362
## bio4 -0.1692491 0.1083258 0.28348226 -0.48121938
## bio5 -0.5920746 -0.7243068 -0.77768793 -0.34301616
## bio6 -0.3148547 -0.6376939 -0.80126262 0.07715578
## bio7 -0.3541091 -0.1147379 0.02381502 -0.53249335
## bio8 -0.2174797 -0.3048809 0.03291308 -0.51356967
## bio9 -0.3274923 -0.5552129 -0.81633533 0.10720732
## bio10 -0.5151169 -0.7036900 -0.76592247 -0.26914814
## bio11 -0.3651390 -0.6835638 -0.82992875 0.01511498
## bio12 0.9178974 0.8079134 0.76402046 0.74717071
## bio13 0.9902335 0.5245850 0.58835234 0.79069926
## bio14 0.5012004 0.9859234 0.84673991 0.31690226
## bio15 0.1244492 -0.7294802 -0.46226779 0.12829990
## bio16 1.0000000 0.5362200 0.61586400 0.79209065
## bio17 0.5362200 1.0000000 0.81764699 0.38396995
## bio18 0.6158640 0.8176470 1.00000000 0.17654879
## bio19 0.7920907 0.3839700 0.17654879 1.00000000
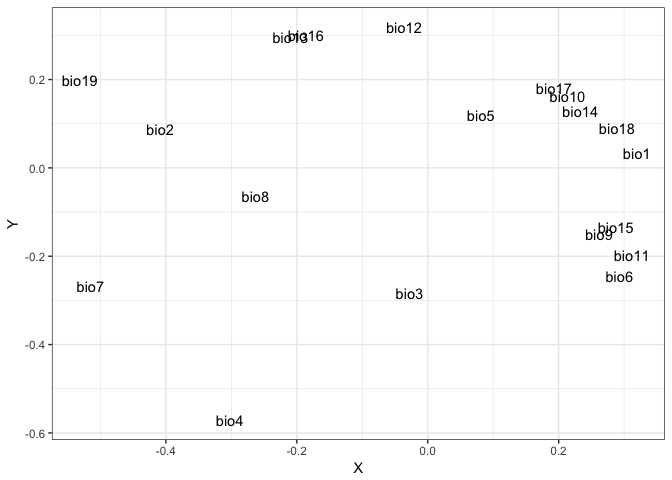
That's great, but it's a bit hard to pick variables this way. Let's try it visually instead.
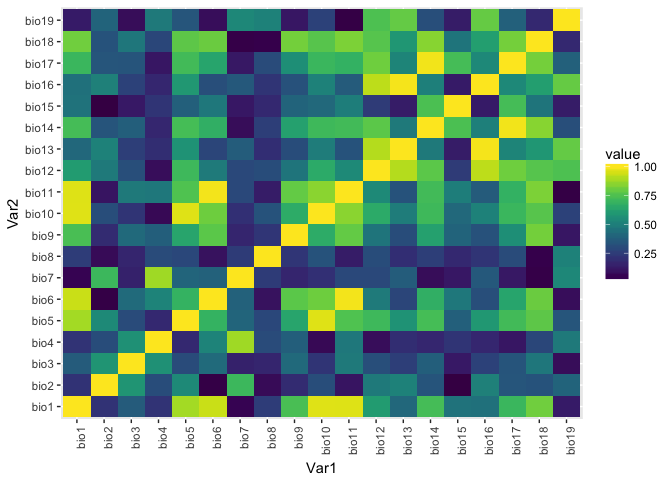
raster.cor.plot(env)
## $cor.mds.plot
##
## $cor.heatmap
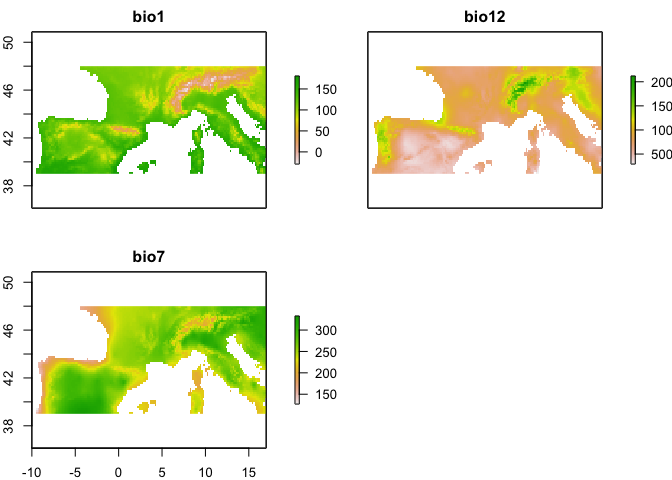
The raster.cor.plot function gives us two visualizations. One heatmap that colors pairs of predictors by their Pearson correlation coefficient, and one cluster plot that does mds scaling of the predictor variables and then plots them in a two dimensional space so that more correlated predictors are closer to each other. We're going to make an arbitrary decision to just use three predictors, and to keep those predictors relatively uncorrelated we'll select predictors that are far apart in this mds plot. Here we'll choose bio1, bio12, and bio7.
env <- env[[c("bio1", "bio12", "bio7")]]
plot(env)
raster.cor.matrix(env)
## bio1 bio12 bio7
## bio1 1.00000000 -0.6084667 -0.05329392
## bio12 -0.60846674 1.0000000 -0.29133189
## bio7 -0.05329392 -0.2913319 1.00000000
GLMs usually require the user to supply a formula, an enmtools.species object, and some environmental data. If your formula is a strictly additive function of all of the environmental layers in env, though, enmtools.glm will build a formula automatically.
monticola.glm <- enmtools.glm(species = monticola, env = env, f = pres ~ bio1 + bio12 + bio7, test.prop = 0.2)
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
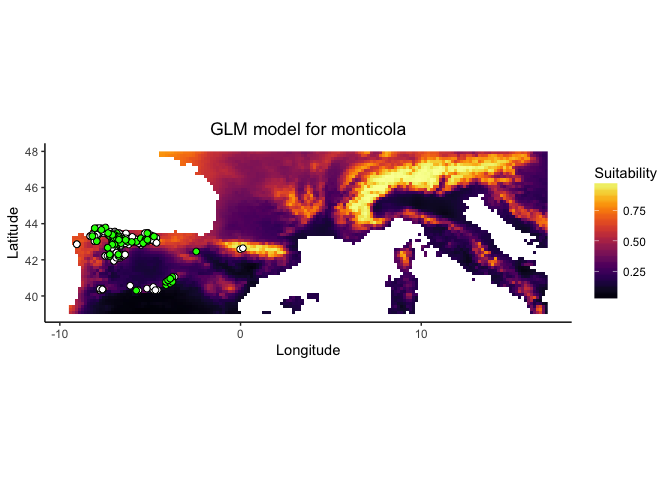
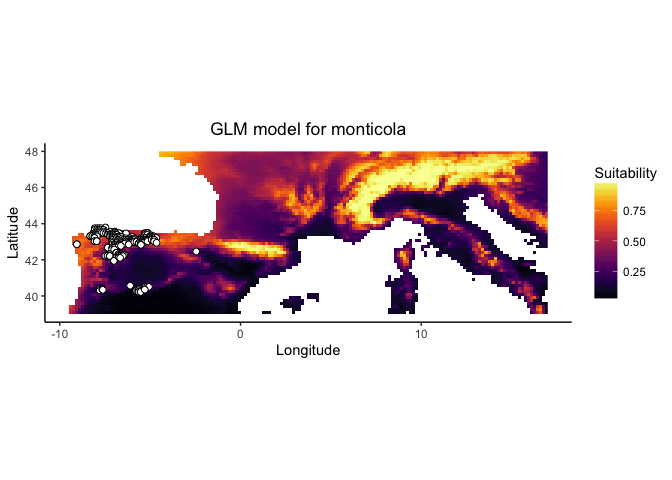
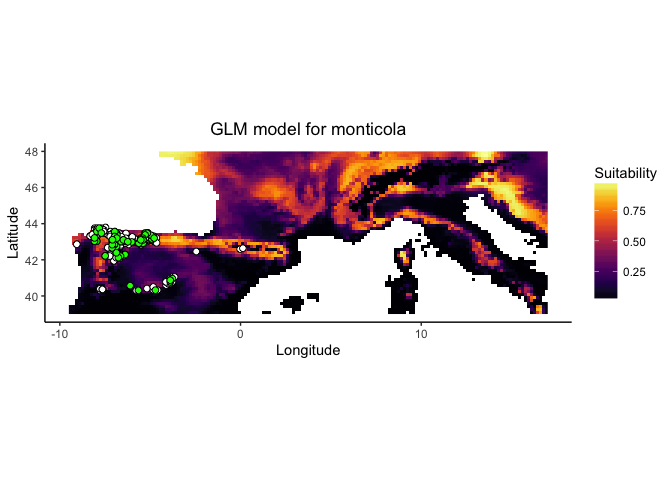
monticola.glm
##
##
## Formula: presence ~ bio1 + bio12 + bio7
## <environment: 0x7fdc2203aa70>
##
##
## Data table (top ten lines):
##
## Longitude Latitude bio1 bio12 bio7 presence
## --- ---------- --------- ----- ------ ----- ---------
## 1 -5.171215 43.06957 78 917 249 1
## 2 -6.036635 43.02531 76 1012 246 1
## 3 -7.679727 40.38852 137 1143 247 1
## 4 -7.790437 40.30959 129 1231 242 1
## 6 -6.575039 42.91070 84 1012 247 1
## 7 -5.132756 43.49572 133 822 190 1
## 8 -7.787378 40.39362 137 1143 247 1
## 9 -4.941888 43.35310 128 843 194 1
## 10 -7.621731 40.34170 101 1514 229 1
## 11 -7.645674 40.36543 101 1514 229 1
##
##
## Model:
## Call:
## glm(formula = f, family = "binomial", data = analysis.df[, -c(1,
## 2)], weights = weights)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -1.6439 -0.7903 -0.4959 0.8448 2.2629
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) 8.3089037 2.1931131 3.789 0.000151 ***
## bio1 -0.0342866 0.0065015 -5.274 1.34e-07 ***
## bio12 0.0003411 0.0007222 0.472 0.636701
## bio7 -0.0205783 0.0049710 -4.140 3.48e-05 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 576.70 on 719 degrees of freedom
## Residual deviance: 507.23 on 716 degrees of freedom
## AIC: 249.05
##
## Number of Fisher Scoring iterations: 4
##
##
##
## Model fit (training data): class : ModelEvaluation
## n presences : 208
## n absences : 512
## AUC : 0.7393705
## cor : 0.3210289
## max TPR+TNR at : -0.01700272
##
##
## Environment space model fit (training data): class : ModelEvaluation
## n presences : 208
## n absences : 10000
## AUC : 0.4276125
## cor : -0.01447189
## max TPR+TNR at : 0.4956744
##
##
## Proportion of data wittheld for model fitting: 0.2
##
## Model fit (test data): class : ModelEvaluation
## n presences : 52
## n absences : 512
## AUC : 0.6924579
## cor : 0.162511
## max TPR+TNR at : -0.00580997
##
##
## Environment space model fit (test data): class : ModelEvaluation
## n presences : 52
## n absences : 10000
## AUC : 0.4033269
## cor : -0.01488643
## max TPR+TNR at : 0.4984725
##
##
## Suitability:
## class : RasterLayer
## dimensions : 54, 162, 8748 (nrow, ncol, ncell)
## resolution : 0.1666667, 0.1666667 (x, y)
## extent : -10, 17, 39, 48 (xmin, xmax, ymin, ymax)
## coord. ref. : +proj=longlat +datum=WGS84 +ellps=WGS84 +towgs84=0,0,0
## data source : in memory
## names : layer
## values : 0.02397329, 0.9972282 (min, max)
##
##
##
## Notes:
Notice this produces the same formula as:
monticola.glm <- enmtools.glm(species = monticola, env = env, test.prop = 0.2)
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
monticola.glm
##
##
## Formula: presence ~ bio1 + bio12 + bio7
## <environment: 0x7fdc379fe9c8>
##
##
## Data table (top ten lines):
##
## Longitude Latitude bio1 bio12 bio7 presence
## --- ---------- --------- ----- ------ ----- ---------
## 1 -5.171215 43.06957 78 917 249 1
## 2 -6.036635 43.02531 76 1012 246 1
## 3 -7.679727 40.38852 137 1143 247 1
## 4 -7.790437 40.30959 129 1231 242 1
## 7 -5.132756 43.49572 133 822 190 1
## 8 -7.787378 40.39362 137 1143 247 1
## 9 -4.941888 43.35310 128 843 194 1
## 11 -7.645674 40.36543 101 1514 229 1
## 12 -7.642539 40.36317 101 1514 229 1
## 13 -6.990000 42.57000 107 893 253 1
##
##
## Model:
## Call:
## glm(formula = f, family = "binomial", data = analysis.df[, -c(1,
## 2)], weights = weights)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -1.6668 -0.7985 -0.4761 0.8251 2.2069
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) 9.7242692 2.2707882 4.282 1.85e-05 ***
## bio1 -0.0371990 0.0066838 -5.566 2.61e-08 ***
## bio12 -0.0000931 0.0007383 -0.126 0.9
## bio7 -0.0235477 0.0051508 -4.572 4.84e-06 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 576.70 on 719 degrees of freedom
## Residual deviance: 505.19 on 716 degrees of freedom
## AIC: 245.69
##
## Number of Fisher Scoring iterations: 4
##
##
##
## Model fit (training data): class : ModelEvaluation
## n presences : 208
## n absences : 512
## AUC : 0.7367131
## cor : 0.3226584
## max TPR+TNR at : 0.06883724
##
##
## Environment space model fit (training data): class : ModelEvaluation
## n presences : 208
## n absences : 10000
## AUC : 0.4251981
## cor : -0.01381147
## max TPR+TNR at : 0.3610308
##
##
## Proportion of data wittheld for model fitting: 0.2
##
## Model fit (test data): class : ModelEvaluation
## n presences : 52
## n absences : 512
## AUC : 0.7000826
## cor : 0.1634975
## max TPR+TNR at : 0.07265669
##
##
## Environment space model fit (test data): class : ModelEvaluation
## n presences : 52
## n absences : 10000
## AUC : 0.4120712
## cor : -0.01234761
## max TPR+TNR at : 0.4952489
##
##
## Suitability:
## class : RasterLayer
## dimensions : 54, 162, 8748 (nrow, ncol, ncell)
## resolution : 0.1666667, 0.1666667 (x, y)
## extent : -10, 17, 39, 48 (xmin, xmax, ymin, ymax)
## coord. ref. : +proj=longlat +datum=WGS84 +ellps=WGS84 +towgs84=0,0,0
## data source : in memory
## names : layer
## values : 0.01949305, 0.9973357 (min, max)
##
##
##
## Notes:
If you want a more complicated formula, though (e.g., with interactions or polynomial effects), you'll need to supply that manually.
monticola.glm <- enmtools.glm(species = monticola, env = env, f = pres ~ poly(bio1, 2) + poly(bio7, 2) * poly(bio12, 2), test.prop = 0.2)
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
monticola.glm
##
##
## Formula: presence ~ poly(bio1, 2) + poly(bio7, 2) + poly(bio12, 2) + poly(bio7,
## 2):poly(bio12, 2)
## <environment: 0x7fdc20831e60>
##
##
## Data table (top ten lines):
##
## Longitude Latitude bio1 bio12 bio7 presence
## --- ---------- --------- ----- ------ ----- ---------
## 1 -5.171215 43.06957 78 917 249 1
## 2 -6.036635 43.02531 76 1012 246 1
## 3 -7.679727 40.38852 137 1143 247 1
## 4 -7.790437 40.30959 129 1231 242 1
## 5 -7.473340 43.78935 140 931 179 1
## 6 -6.575039 42.91070 84 1012 247 1
## 7 -5.132756 43.49572 133 822 190 1
## 8 -7.787378 40.39362 137 1143 247 1
## 10 -7.621731 40.34170 101 1514 229 1
## 11 -7.645674 40.36543 101 1514 229 1
##
##
## Model:
## Call:
## glm(formula = f, family = "binomial", data = analysis.df[, -c(1,
## 2)], weights = weights)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -1.3269 -0.7112 -0.3866 0.6404 2.6673
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) -0.9591 0.4109 -2.334 0.01958 *
## poly(bio1, 2)1 -30.6734 5.7687 -5.317 1.05e-07 ***
## poly(bio1, 2)2 -32.4723 6.4276 -5.052 4.37e-07 ***
## poly(bio7, 2)1 -16.7741 12.8116 -1.309 0.19043
## poly(bio7, 2)2 5.8198 10.6370 0.547 0.58429
## poly(bio12, 2)1 34.5309 12.6304 2.734 0.00626 **
## poly(bio12, 2)2 -9.2282 9.7090 -0.950 0.34187
## poly(bio7, 2)1:poly(bio12, 2)1 -238.9043 440.0210 -0.543 0.58717
## poly(bio7, 2)2:poly(bio12, 2)1 -76.0475 291.2958 -0.261 0.79404
## poly(bio7, 2)1:poly(bio12, 2)2 624.6585 287.3713 2.174 0.02973 *
## poly(bio7, 2)2:poly(bio12, 2)2 99.7371 177.5479 0.562 0.57429
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 576.70 on 719 degrees of freedom
## Residual deviance: 448.36 on 709 degrees of freedom
## AIC: 238.89
##
## Number of Fisher Scoring iterations: 5
##
##
##
## Model fit (training data): class : ModelEvaluation
## n presences : 208
## n absences : 512
## AUC : 0.7964524
## cor : 0.4205467
## max TPR+TNR at : 0.1333889
##
##
## Environment space model fit (training data): class : ModelEvaluation
## n presences : 208
## n absences : 10000
## AUC : 0.7314553
## cor : 0.1168428
## max TPR+TNR at : 0.3741629
##
##
## Proportion of data wittheld for model fitting: 0.2
##
## Model fit (test data): class : ModelEvaluation
## n presences : 52
## n absences : 512
## AUC : 0.7557843
## cor : 0.2330202
## max TPR+TNR at : -0.08155348
##
##
## Environment space model fit (test data): class : ModelEvaluation
## n presences : 52
## n absences : 10000
## AUC : 0.7097
## cor : 0.0509356
## max TPR+TNR at : 0.2494423
##
##
## Suitability:
## class : RasterLayer
## dimensions : 54, 162, 8748 (nrow, ncol, ncell)
## resolution : 0.1666667, 0.1666667 (x, y)
## extent : -10, 17, 39, 48 (xmin, xmax, ymin, ymax)
## coord. ref. : +proj=longlat +datum=WGS84 +ellps=WGS84 +towgs84=0,0,0
## data source : in memory
## names : layer
## values : 2.220446e-16, 0.992979 (min, max)
##
##
##
## Notes:
To check out the marginal response functions, you only need to type
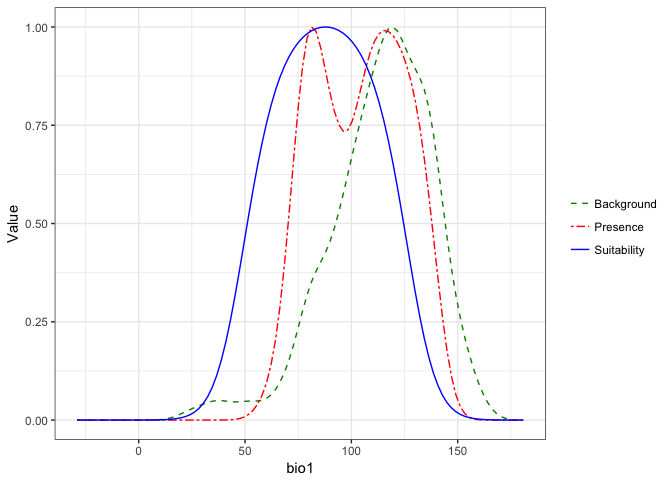
monticola.glm$response.plots
## $bio1
##
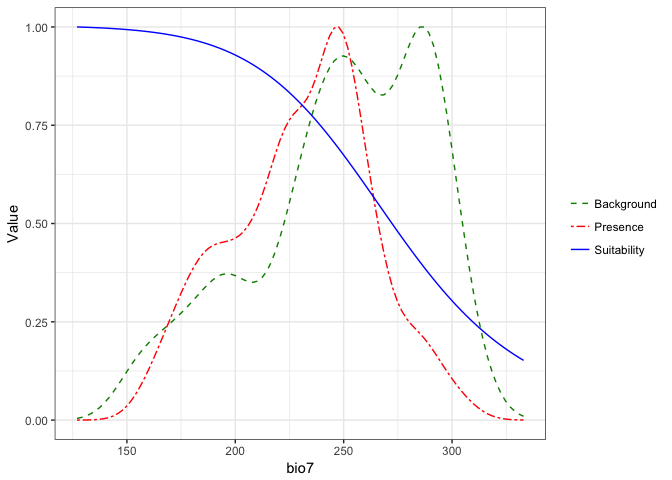
## $bio7
##
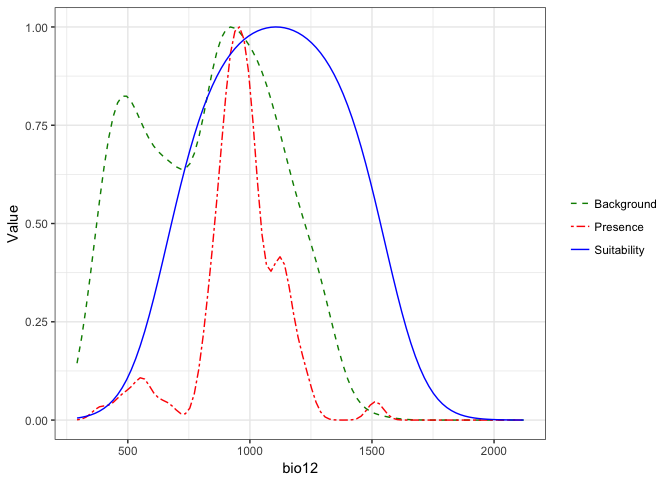
## $bio12
These plots present a smoothed estimate of the frequency of different levels of the environemntal variable in the presence data and the background points, along with the estimated relationship between that environmental predictor and the suitability of habitat from the model.
You can also visualize your models and data in a 2D environment space using any pair of layers from your environment stack. These plots hold all non-plotted variables (bio7 in this case) constant at their mean value across all presence points, then vary the plotted variables between the minimum and maximum values in env.
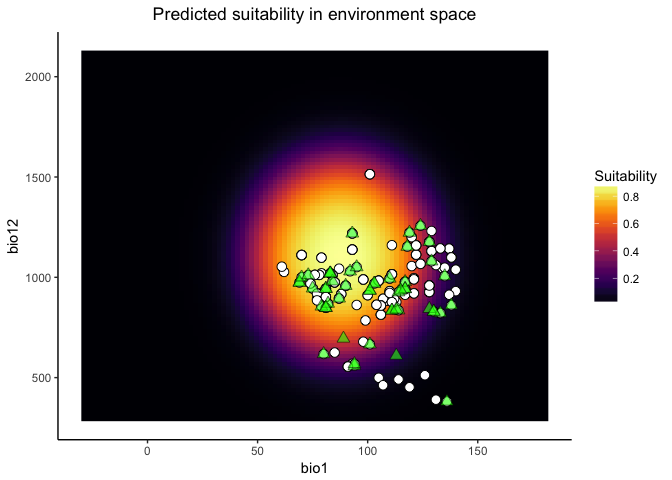
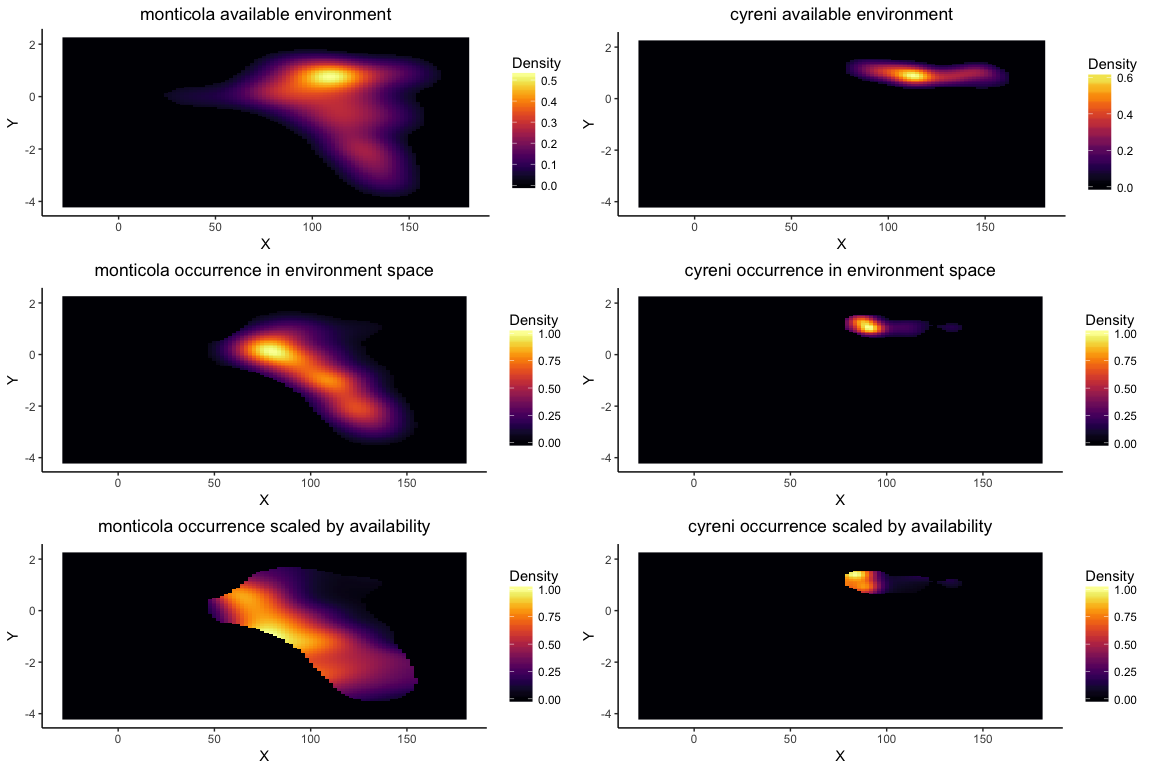
The suit.plot shows you suitability in environment space as a function of your two variables, with brighter colors representing variable combinations predicted to be more suitable. The points represent the occurrence points for your species in that environment space.
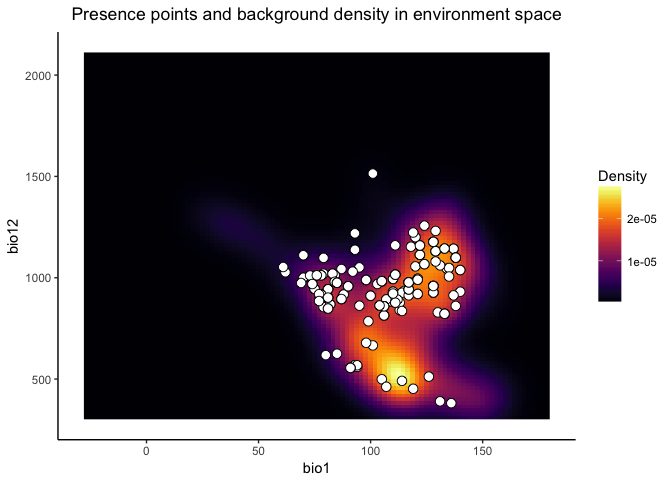
The colored raster of the background.plot shows you the density of background points in environment space, while the white points again represent your occurrence points in environment space.
visualize.enm(monticola.glm, env, layers = c("bio1", "bio12"), plot.test.data = TRUE)
## $background.plot
##
## $suit.plot
The procedure for building Bioclim, Domain, and Maxent models is similar to the procedure for GLMs, with the exception that you do not need to pass a formula to the model function for Maxent, Domain, and Bioclim models. Note that running Maxent models requires a bit of extra setup; see dismo documentation for details.
monticola.gam <- enmtools.gam(monticola, env, f = pres ~ poly(bio1, 2) + poly(bio7, 2) * poly(bio12, 2), test.prop = 0.2)
monticola.dm <- enmtools.dm(monticola, env, test.prop = 0.2)
monticola.bc <- enmtools.bc(monticola, env, test.prop = 0.2)
monticola.mx <- enmtools.maxent(monticola, env, test.prop = 0.2)
ENMTools provides a number of metrics for ENMs and for similarities between ENMs. These include measures of niche breadth, based on Levins(1968). An important caveat when interpreting these metrics is that they are driven to some (variable) extent by the availability of different combinations of environmental predictors. As such they are more accurately interpreted as a measurment of the smoothness of the geographic distribution of suitability scores than as an estimate of the breadth of the fundamental niche; an organism with narrow fundamental niche breadth that nonetheless encompasses a set of environmental conditions that is quite common will have a high breadth when measured using ENMs, while having a low breadth in environment space.
raster.breadth(monticola.glm)
## $B1
## [1] 0.9440004
##
## $B2
## [1] 0.5296247
ENMTools also provides metrics for measuring similarity between ENMs. These include Schoener's D (Schoener 1968), I (Warren et al. 2008), and the Spearman rank correlation coefficient between two rasters. While D and I are commonly used in the ENM literature, they may tend to overestimate similarity between ENMs when many grid cells are of similar values (e.g., when two species prefer different habitat but the region contains a great deal of habitat that is unsuitable for both).
monticola.glm <- enmtools.glm(species = monticola, env = env, f = pres ~ poly(bio1, 2) + poly(bio7, 2) + poly(bio12, 2), test.prop = 0.2)
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
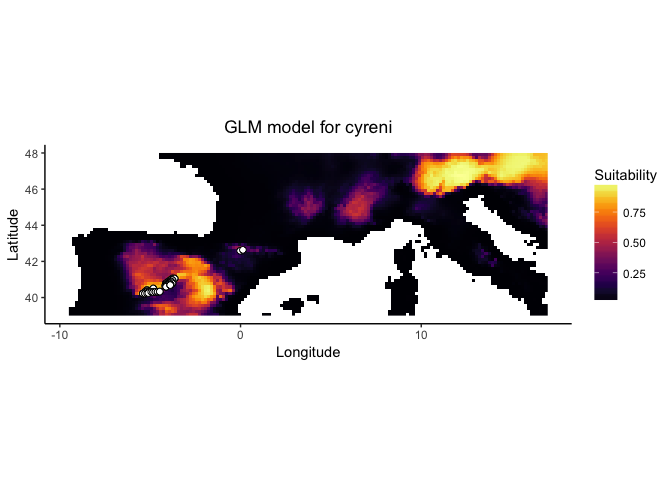
cyreni.glm <- enmtools.glm(species = cyreni, env = env, f = pres ~ poly(bio1, 2) + poly(bio7, 2) + poly(bio12, 2), test.prop = 0.2)
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
raster.overlap(monticola.glm, cyreni.glm)
## $D
## [1] 0.6964057
##
## $I
## [1] 0.9026603
##
## $rank.cor
## [1] 0.3249225
A new feature of the R version of ENMTools is that you can now use these same metrics in the n-dimensional space of all combinations of environmental variables, instead of restricting your measures of model similarity to those sets of conditions that appear in the training region. This is done by repeatedly drawing Latin hypercube samples from the space of all possible combinations of environmental variables given the min and max of each variable within the training region. ENMTools continues to draw samples until subsequent iterations differ by less than a specified tolerance value. Lower tolerance values result in more precise estimates of overlap, but can take much longer to calculate.
monticola.glm <- enmtools.glm(species = monticola, env = env, f = pres ~ poly(bio1, 2) + poly(bio7, 2) + poly(bio12, 2), test.prop = 0.2)
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
cyreni.glm <- enmtools.glm(species = monticola, env = env, f = pres ~ poly(bio1, 2) + poly(bio7, 2) + poly(bio12, 2), test.prop = 0.2)
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
env.overlap(monticola.glm, cyreni.glm, env, tolerance = .001)
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## $env.D
## [1] 0.9747509
##
## $env.I
## [1] 0.9992639
##
## $env.cor
## [1] 0.999296
##
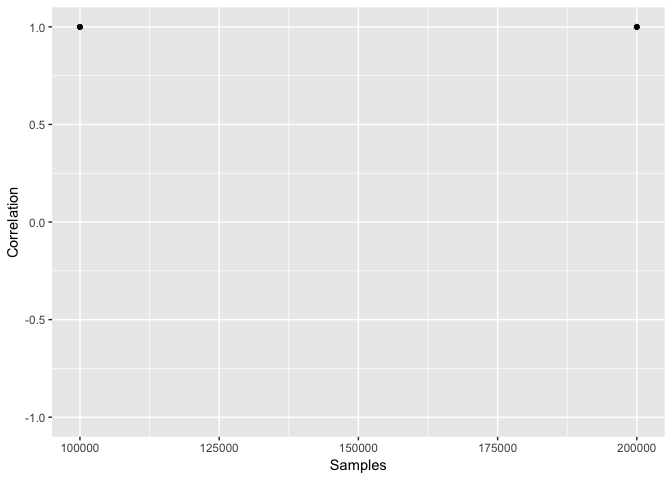
## $env.D.plot
##
## $env.I.plot
##
## $env.cor.plot
The plots that come out of these environment space functions are used for diagnosing convergence of the overlap/breadth metric. Ideally what you want is a relationship between the metric and the number of samples that shows no clear directional trend.
In this example, we will run a niche identity (also called equivalency) test, as in Warren et al. 2008. This test takes the presence points for a pair of species and randomly reassigns them to each species, then builds ENMs for these randomized occurrences. By doing this many times, we can estimate the probability distribution for ENM overlap between species under the null hypothesis that the two species' occurrences in the environment are effectively a random draw from the same underlying distribution. Note that niche evolution is only one of many reasons why two species' realized environmental distributions might cause departures from this null hypothesis. See Warren et al. 2014 for details.
To run an identity test, we need to decide what type of models we will build, how many replicates we will run, and (in the case of GLM and GAM) a model formula to use for empirical models and the Monte Carlo replicates. The resulting object contains the replicate models, p values, and plots of the results. Typically idenity tests are run with at least 99 replicates, but we are using a smaller number here for the sake of execution time.
NOTE: In order for models to be comparable, both empirical and pseudoreplicate models for the identity test are conducted with pseudoabsence points pooled for the two species being compared.
id.glm <- identity.test(species.1 = monticola, species.2 = cyreni, env = env, type = "glm", nreps = 4)
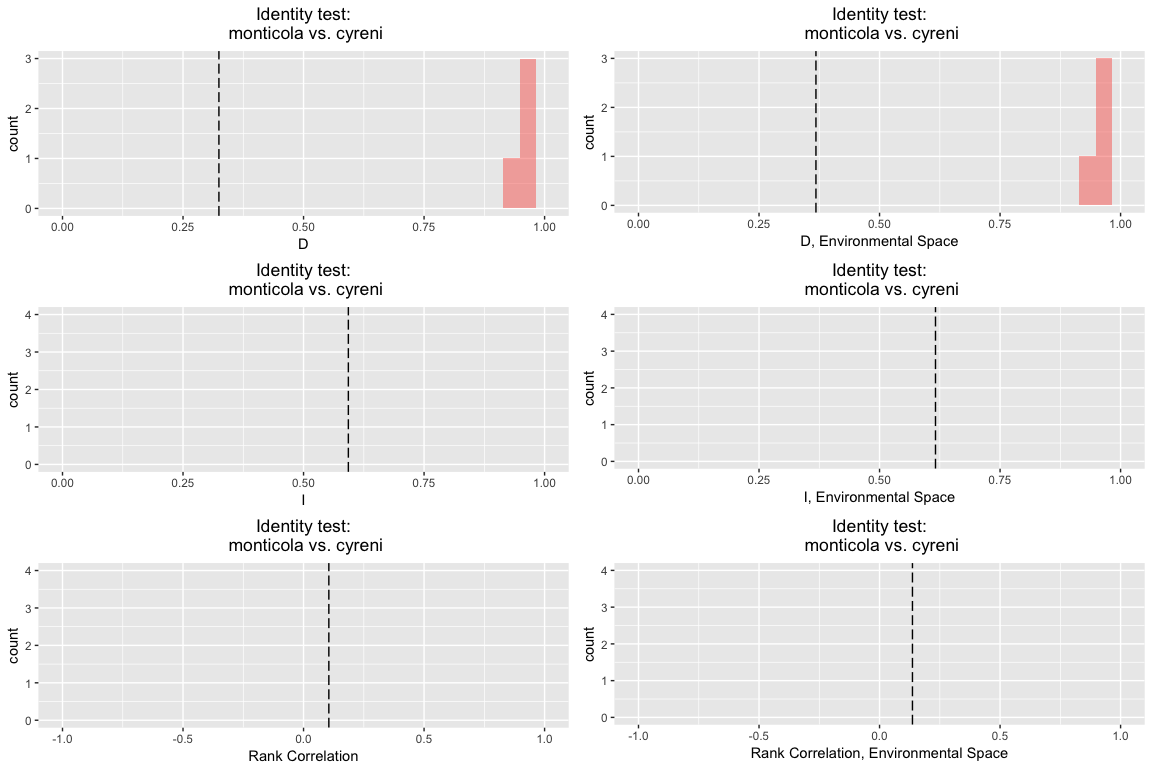
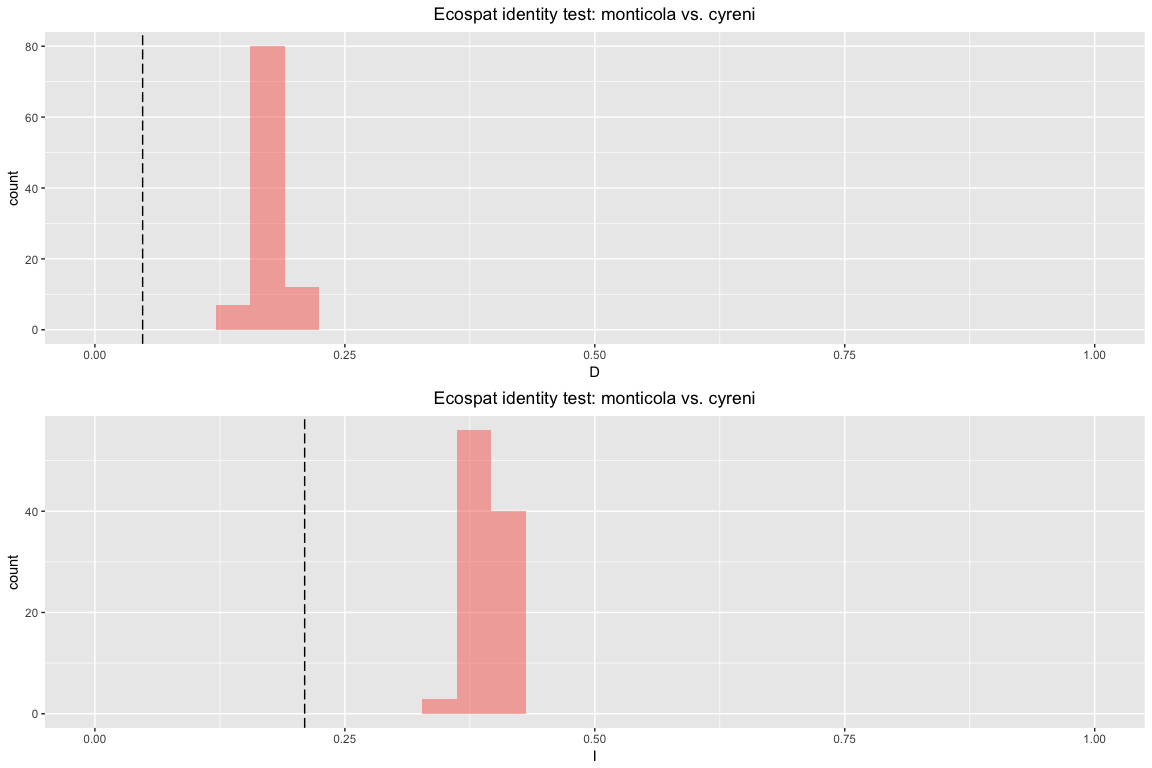
id.glm
##
##
##
##
## Identity test monticola vs. cyreni
##
## objectentity test p-values:
## D I rank.cor env.D env.I env.cor
## 0.2 0.2 0.2 0.2 0.2 0.2
##
##
## Replicates:
##
##
## D I rank.cor env.D env.I env.cor
## ---------- ---------- ---------- ---------- ---------- ---------- ----------
## empirical 0.3244487 0.5929443 0.1049357 0.3680675 0.6160525 0.1365335
## rep 1 0.9677634 0.9990155 0.9920496 0.9653146 0.9979226 0.9947297
## rep 2 0.9489356 0.9972812 0.9647560 0.9451813 0.9947636 0.9718006
## rep 3 0.9765731 0.9993665 0.9920094 0.9739029 0.9991720 0.9953646
## rep 4 0.9422789 0.9965344 0.9877409 0.9628450 0.9978574 0.9886635
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
The background or similarity test compares the overlap seen between two species' ENMs to the overlap expected by chance if one or both species was effectively choosing habitat at random from within their broad geographic range. The purpose of this test is to correct for the availability of habitat and ask whether the observed similarity between species or populations is significantly more (or less) than expected given the available set of environments in the regions in which they occur.
NOTE: In order for models to be comparable, both empirical and pseudoreplicate models for the background test are conducted with pseudoabsence points pooled for the two species being compared.
In Warren et al. 2008, we developed this test in the context of comparing one species' actual occurrence to the random background occurrences of the other species. This is what we call an "asymmetric" test, and in our case we did the test in both directions with the idea that we might compare the results of A vs. B background to the results of B vs. A background. This may be informative in some cases, but many people have also found this asymmetry confusing (and indeed it is often difficult to interpret). For that reason, the background test here can be conducted against a null hypothesis that is generated from "asymmetric" (species.1 vs species.2 background) or "symmetric" (species.1 background vs. species.2 background) comparisons.
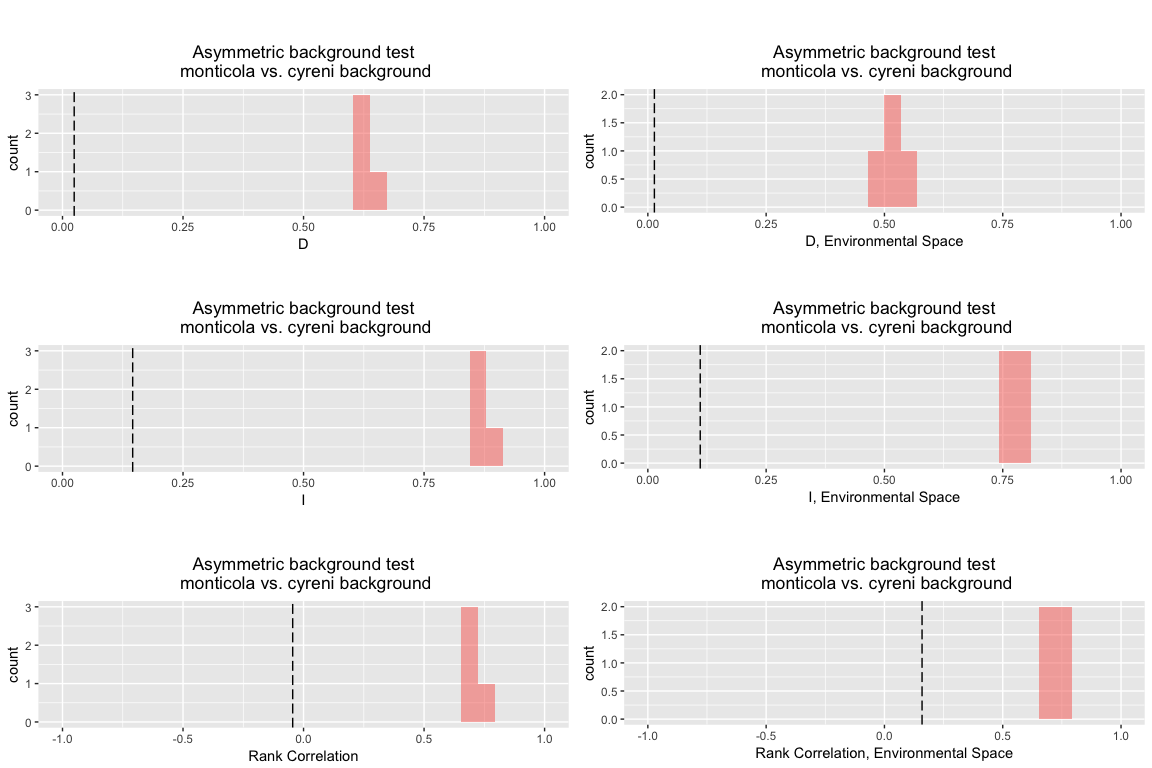
Here, for instance, is a Bioclim background test using the classical asymmetric approach:
bg.bc.asym <- background.test(species.1 = monticola, species.2 = cyreni, env = env, type = "bc", nreps = 4, test.type = "asymmetric" )
bg.bc.asym
##
##
##
##
## Asymmetric background test
## monticola vs. cyreni background
##
## background test p-values:
## D I rank.cor env.D env.I env.cor
## 0.4 0.4 0.4 0.4 0.4 0.4
##
##
## Replicates:
##
##
## D I rank.cor env.D env.I env.cor
## ---------- ---------- ---------- ----------- ---------- ---------- ----------
## empirical 0.0242365 0.1455829 -0.0450249 0.0138222 0.1107173 0.1589788
## rep 1 0.6214600 0.8700845 0.6715790 0.5110334 0.7742323 0.6747580
## rep 2 0.6140519 0.8692591 0.6881657 0.4890054 0.7671731 0.6784506
## rep 3 0.6477756 0.8921239 0.7362763 0.5359914 0.8078705 0.7271245
## rep 4 0.6349293 0.8741857 0.6796873 0.5338093 0.7872621 0.7335395
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
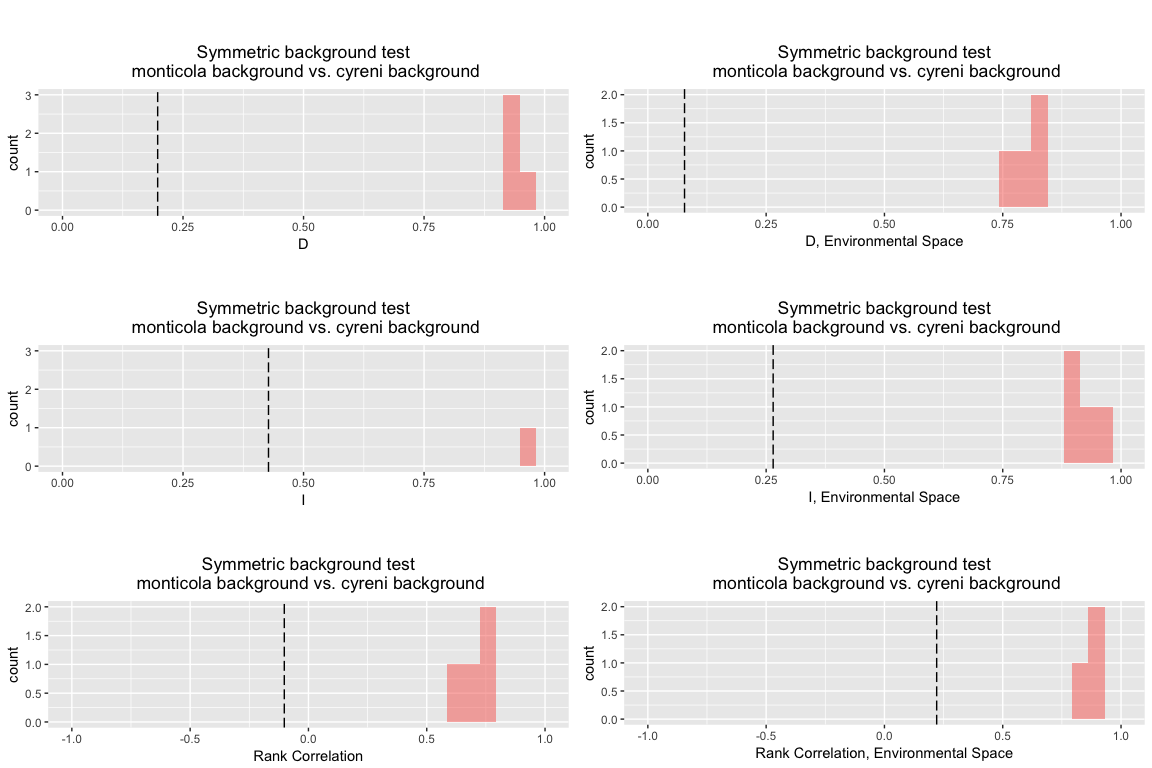
And here is a Domain background test using the symmetric approach:
bg.dm.sym <- background.test(species.1 = monticola, species.2 = cyreni, env = env, type = "dm", nreps = 4, test.type = "symmetric" )
bg.dm.sym
##
##
##
##
## Symmetric background test
## monticola background vs. cyreni background
##
## background test p-values:
## D I rank.cor env.D env.I env.cor
## 0.4 0.4 0.4 0.4 0.4 0.4
##
##
## Replicates:
##
##
## D I rank.cor env.D env.I env.cor
## ---------- ---------- ---------- ----------- ---------- ---------- ----------
## empirical 0.1974048 0.4272913 -0.1020590 0.0775272 0.2648275 0.2207919
## rep 1 0.9462649 0.9964841 0.6346617 0.8440406 0.9521950 0.9254358
## rep 2 0.9546793 0.9976078 0.7412483 0.8426959 0.9476840 0.8970172
## rep 3 0.9213012 0.9824419 0.7453075 0.7434447 0.8853417 0.8458307
## rep 4 0.9407627 0.9926325 0.6575177 0.7768920 0.9123329 0.9387315
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
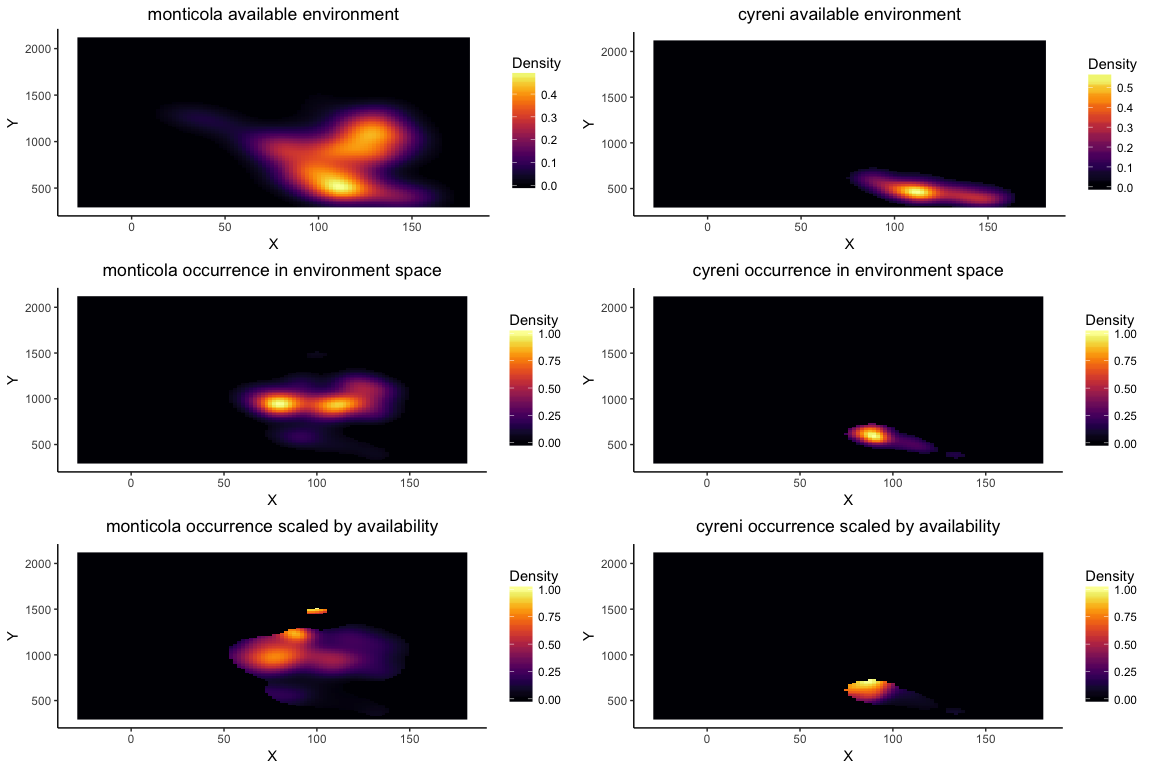
Using enmtools.species objects also provides a simplified interface to the niche equivalency and similarity tests (or identity and background tests, respectively) that were developed by Broennimann et al. (2012). These tests do not rely on ENMs, instead using kernel density smoothing to estimate density of the species in environment space. Ecospat also uses the density of the available environment to correct for availability when measuring overlaps, so that overlaps are not strictly driven by availability of combinations of environmental variables.
These tests only work with two environmental axes, so they are often done with the top two PC axes of a set of environments. In our case we'll just pick a couple of environmental layers, though (bio1 and bio2). Here's an equivalency/identity test:
esp.id <- enmtools.ecospat.id(monticola, cyreni, env[[c("bio1", "bio12")]])
##
##
## No background points provided, drawing background from range raster.
##
##
##
## No background points provided, drawing background from range raster.
esp.id
##
##
##
##
## Ecospat identity test monticola vs. cyreni
##
## ecospat.id test empirical overlaps:
## $D
## [1] 0.0477392
##
## $I
## [1] 0.2097866
##
##
##
## ecospat.id test p-values:
## D I
## 0 0
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## NULL
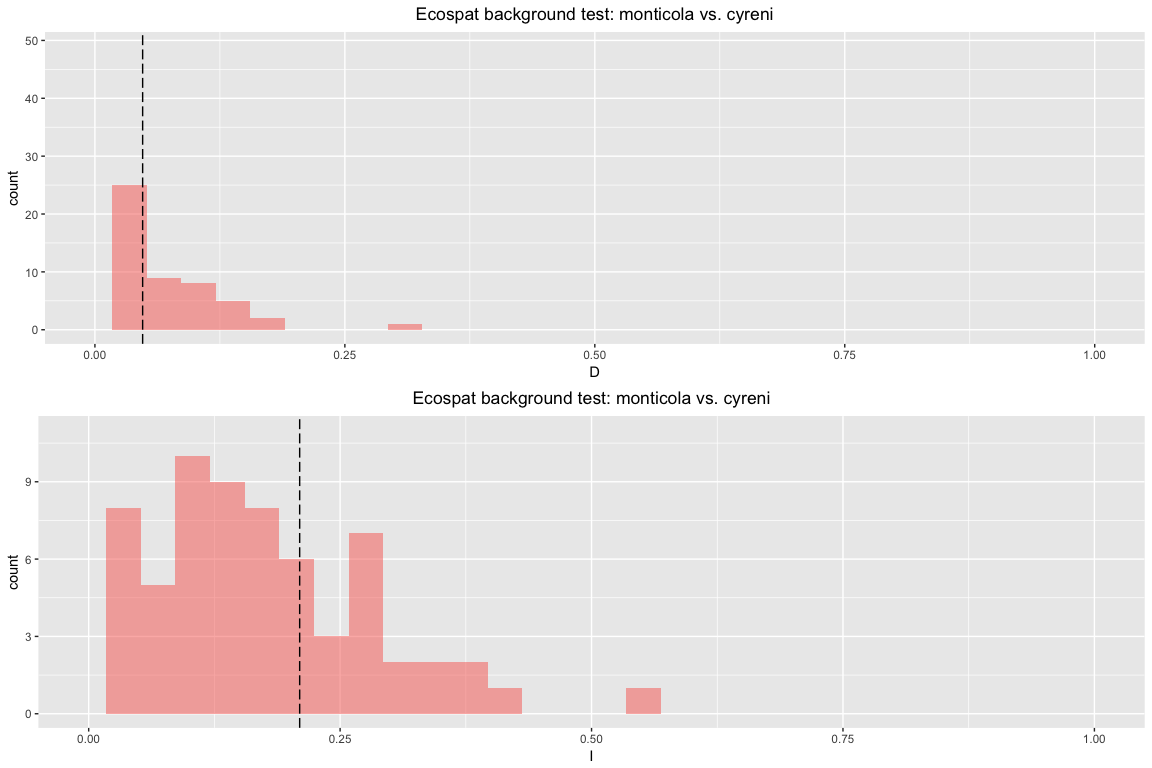
And here's a symmetric background test. The difference between symmetric and asymmetric for these tests is the same as for the background tests presented above.
esp.bg.sym <- enmtools.ecospat.bg(monticola, cyreni, env[[c("bio1", "bio12")]], test.type = "symmetric")
##
##
## No background points provided, drawing background from range raster.
##
##
##
## No background points provided, drawing background from range raster.
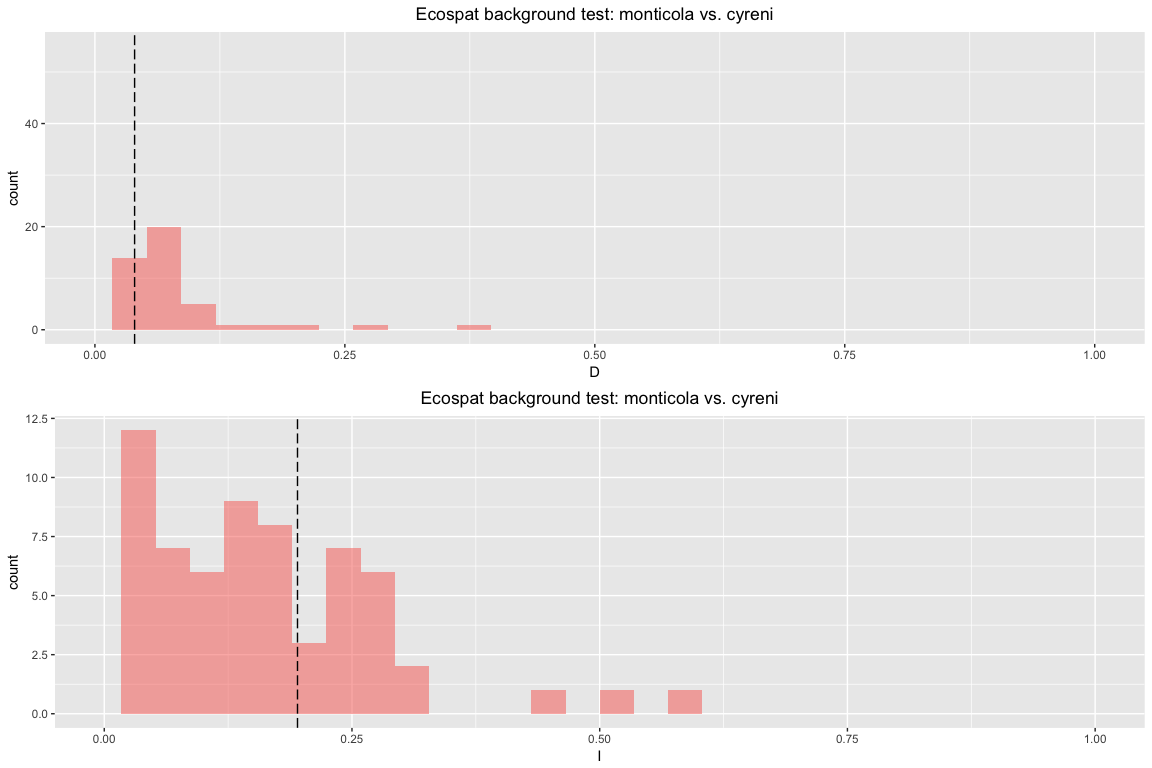
esp.bg.sym
##
##
##
##
## Ecospat background test symmetric monticola vs. cyreni
##
## ecospat.bg test empirical overlaps:
## $D
## [1] 0.0477392
##
## $I
## [1] 0.2097866
##
##
##
## ecospat.bg test p-values:
## D I
## 0.56 0.48
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 24 rows containing non-finite values (stat_bin).
## NULL
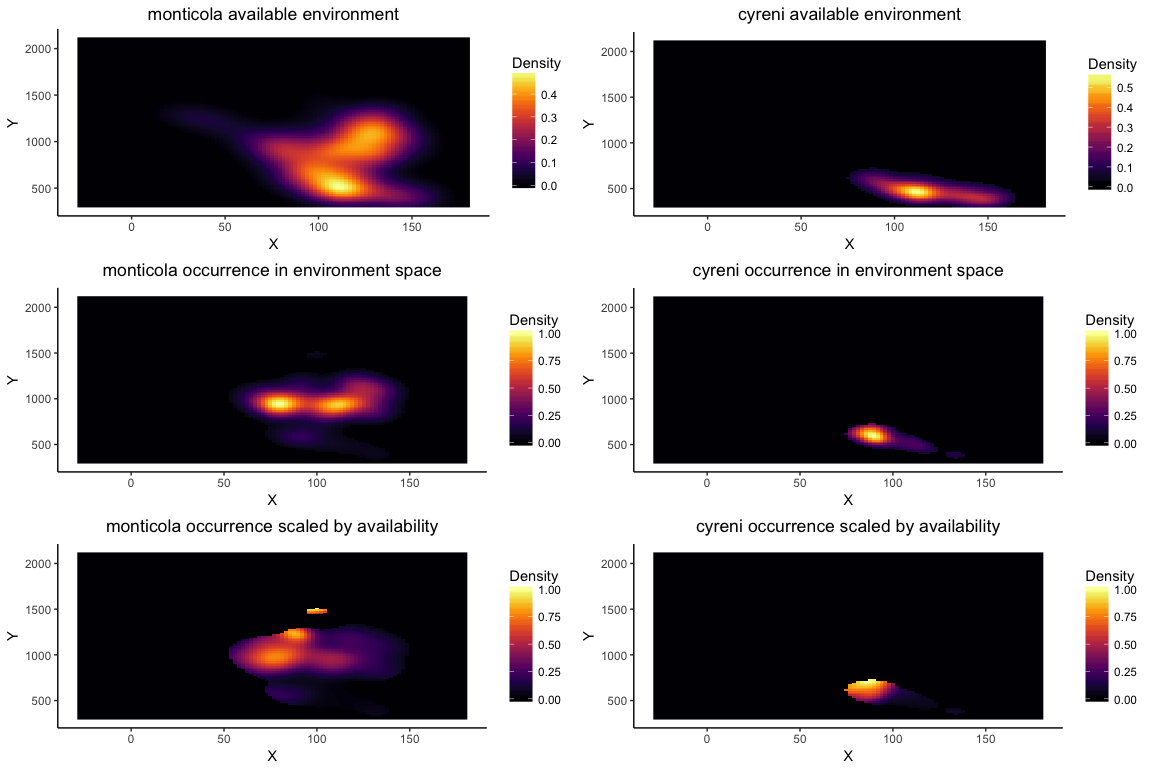
Note that if you provide more than two layers to the enmtools.ecospat function, it will performa a PCA analysis on the provided layers and measure overlaps on the first two axes of that PCA space.
esp.bg.sym <- enmtools.ecospat.bg(monticola, cyreni, env, test.type = "symmetric")
##
##
## No background points provided, drawing background from range raster.
##
##
##
## No background points provided, drawing background from range raster.
##
## [1] "More than two layers in environment stack and no layers argument passed, performing PCA..."
esp.bg.sym
##
##
##
##
## Ecospat background test symmetric monticola vs. cyreni
##
## ecospat.bg test empirical overlaps:
## $D
## [1] 0.039695
##
## $I
## [1] 0.195018
##
##
##
## ecospat.bg test p-values:
## D I
## 0.64 0.44
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 32 rows containing non-finite values (stat_bin).
## NULL
ENMTools also allows you to perform linear, blob, and ribbon rangebreak tests as developed in Glor and Warren 2011. The linear and blob tests are two versions of a test that permit one to ask whether the geographic regions occupied by two species are more environmentally different than expected by chance. The ribbon test, meanwhile, is designed to test whether the ranges of two species are divided by a region that is relatively unsuitable to one or both forms.
For the linear and blob tests, you call them very much like you would the identity and background tests. Here's a linear one using GLM models:
rbl.glm <- rangebreak.linear(monticola, cyreni, env, type = "glm", nreps = 4)
##
##
## No background points provided, drawing background from range raster.
##
##
##
## No background points provided, drawing background from range raster.
##
##
## Building empirical models...
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Building replicate models...
##
## Replicate 1 ...
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Replicate 2 ...
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Replicate 3 ...
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Replicate 4 ...
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
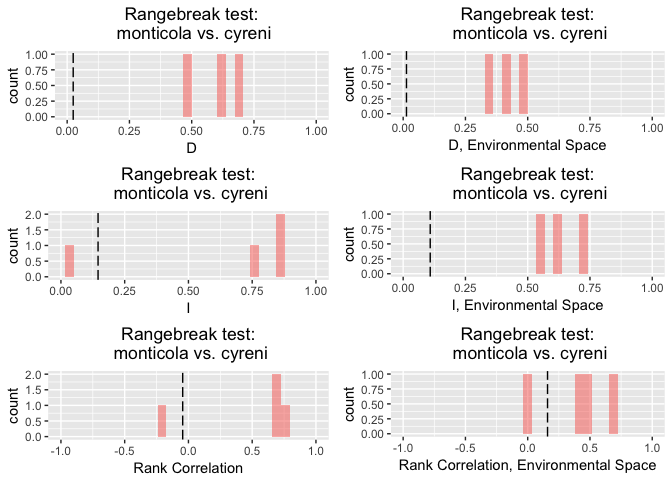
rbl.glm
##
##
##
##
## Linear rangebreak test monticola vs. cyreni
##
## rangebreak test p-values:
## D I rank.cor env.D env.I env.cor
## 0.2 0.2 0.4 0.2 0.2 0.4
##
##
## Replicates:
##
##
## D I rank.cor env.D env.I env.cor
## ---------- ---------- ---------- ---------- ---------- ---------- -----------
## empirical 0.3244487 0.5929443 0.1049357 0.3682391 0.6158986 0.1370378
## rep 1 0.4164302 0.6969905 0.2230349 0.4452615 0.6985640 0.3871524
## rep 2 0.4221705 0.7136580 0.2200321 0.4554071 0.7113299 0.4364791
## rep 3 0.4723282 0.7568531 0.0458225 0.5744372 0.7832484 0.2085522
## rep 4 0.5971889 0.8541483 0.1981989 0.5651623 0.7668461 -0.2813777
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
And here's a blob test using Bioclim:
rbb.bc <- rangebreak.blob(monticola, cyreni, env, type = "bc", nreps = 4)
##
##
## No background points provided, drawing background from range raster.
##
##
##
## No background points provided, drawing background from range raster.
##
##
## Building empirical models...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Building replicate models...
##
## Replicate 1 ...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Replicate 2 ...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Replicate 3 ...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Replicate 4 ...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
rbb.bc
##
##
##
##
## blob rangebreak test monticola vs. cyreni
##
## rangebreak test p-values:
## D I rank.cor env.D env.I env.cor
## 0.4 0.4 0.4 0.4 0.4 0.4
##
##
## Replicates:
##
##
## D I rank.cor env.D env.I env.cor
## ---------- ---------- ---------- ----------- ---------- ---------- ----------
## empirical 0.0242365 0.1455829 -0.0450249 0.0133100 0.1086260 0.1592556
## rep 1 0.6896760 0.8578952 0.7220233 0.4986668 0.7336902 0.7237154
## rep 2 0.0046566 0.0245867 -0.2164500 0.0019719 0.0132680 0.0017443
## rep 3 0.4882798 0.7556071 0.7118018 0.3291984 0.5505698 0.4024447
## rep 4 0.6220568 0.8484927 0.7545716 0.4055820 0.6054990 0.4720669
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
If you want to access the individual replicates (for instance to see how your ranges are being split up), you can find them in the list named "replicate.models" inside your rangebreak test object.
rbl.glm$replicate.models$monticola.rep.1
##
##
## Formula: presence ~ bio1 + bio12 + bio7
## <environment: 0x7fdc11984fb8>
##
##
## Data table (top ten lines):
##
## Longitude Latitude bio1 bio12 bio7 presence
## ---------- --------- ----- ------ ----- ---------
## -8.07 43.75 134 1045 166 1
## -7.94 43.75 131 1061 171 1
## -8.07 43.66 135 1048 168 1
## -7.82 43.75 135 1006 174 1
## -7.95 43.66 129 1080 176 1
## -7.70 43.75 135 1006 174 1
## -7.82 43.66 122 1113 184 1
## -7.95 43.57 129 1080 176 1
## -8.07 43.48 129 1131 178 1
## -8.20 43.40 140 1038 164 1
##
##
## Model:
## Call:
## glm(formula = f, family = "binomial", data = analysis.df[, -c(1,
## 2)], weights = weights)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -1.8790 -0.7547 -0.3902 0.7147 1.8858
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) 8.7745104 2.1895024 4.008 6.14e-05 ***
## bio1 -0.0409593 0.0065667 -6.237 4.45e-10 ***
## bio12 0.0011127 0.0007125 1.562 0.118
## bio7 -0.0217979 0.0048563 -4.489 7.17e-06 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 720.87 on 901 degrees of freedom
## Residual deviance: 584.14 on 898 degrees of freedom
## AIC: 273.49
##
## Number of Fisher Scoring iterations: 5
##
##
##
## Model fit (training data): class : ModelEvaluation
## n presences : 260
## n absences : 642
## AUC : 0.7830068
## cor : 0.3935289
## max TPR+TNR at : -0.4716333
##
##
## Environment space model fit (training data): class : ModelEvaluation
## n presences : 260
## n absences : 10000
## AUC : 0.4248523
## cor : -0.008857617
## max TPR+TNR at : 0.3841534
##
##
## Proportion of data wittheld for model fitting: 0
##
## Model fit (test data): [1] NA
##
##
## Environment space model fit (test data): [1] NA
##
##
## Suitability:
## class : RasterLayer
## dimensions : 54, 162, 8748 (nrow, ncol, ncell)
## resolution : 0.1666667, 0.1666667 (x, y)
## extent : -10, 17, 39, 48 (xmin, xmax, ymin, ymax)
## coord. ref. : +proj=longlat +datum=WGS84 +ellps=WGS84 +towgs84=0,0,0
## data source : in memory
## names : layer
## values : 0.01243438, 0.9996217 (min, max)
##
##
##
## Notes:
rbl.glm$replicate.models$cyreni.rep.1
##
##
## Formula: presence ~ bio1 + bio12 + bio7
## <environment: 0x7fdc3c89aee0>
##
##
## Data table (top ten lines):
##
## Longitude Latitude bio1 bio12 bio7 presence
## ---------- --------- ----- ------ ----- ---------
## -5.18 40.41 85 625 294 1
## -5.18 40.41 85 625 294 1
## -5.29 40.31 89 646 296 1
## -5.29 40.31 89 646 296 1
## -5.17 40.36 85 625 294 1
## -5.41 40.22 93 645 295 1
## -5.41 40.22 93 645 295 1
## -5.18 40.32 89 646 296 1
## -4.83 40.50 107 462 298 1
## -4.83 40.50 107 462 298 1
##
##
## Model:
## Call:
## glm(formula = f, family = "binomial", data = analysis.df[, -c(1,
## 2)], weights = weights)
##
## Deviance Residuals:
## Min 1Q Median 3Q Max
## -0.7418 -0.3423 -0.1757 -0.0051 1.9055
##
## Coefficients:
## Estimate Std. Error z value Pr(>|z|)
## (Intercept) -5.386065 10.904553 -0.494 0.6214
## bio1 -0.105238 0.019167 -5.491 4e-08 ***
## bio12 -0.006370 0.003437 -1.854 0.0638 .
## bio7 0.070905 0.033311 2.129 0.0333 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
##
## (Dispersion parameter for binomial family taken to be 1)
##
## Null deviance: 210.72 on 847 degrees of freedom
## Residual deviance: 129.35 on 844 degrees of freedom
## AIC: 65.765
##
## Number of Fisher Scoring iterations: 7
##
##
##
## Model fit (training data): class : ModelEvaluation
## n presences : 76
## n absences : 772
## AUC : 0.8721366
## cor : 0.2682345
## max TPR+TNR at : -0.1611919
##
##
## Environment space model fit (training data): class : ModelEvaluation
## n presences : 76
## n absences : 10000
## AUC : 0.8260658
## cor : 0.1177308
## max TPR+TNR at : 0.31541
##
##
## Proportion of data wittheld for model fitting: 0
##
## Model fit (test data): [1] NA
##
##
## Environment space model fit (test data): [1] NA
##
##
## Suitability:
## class : RasterLayer
## dimensions : 54, 162, 8748 (nrow, ncol, ncell)
## resolution : 0.1666667, 0.1666667 (x, y)
## extent : -10, 17, 39, 48 (xmin, xmax, ymin, ymax)
## coord. ref. : +proj=longlat +datum=WGS84 +ellps=WGS84 +towgs84=0,0,0
## data source : in memory
## names : layer
## values : 1.15296e-08, 0.9931977 (min, max)
##
##
##
## Notes:
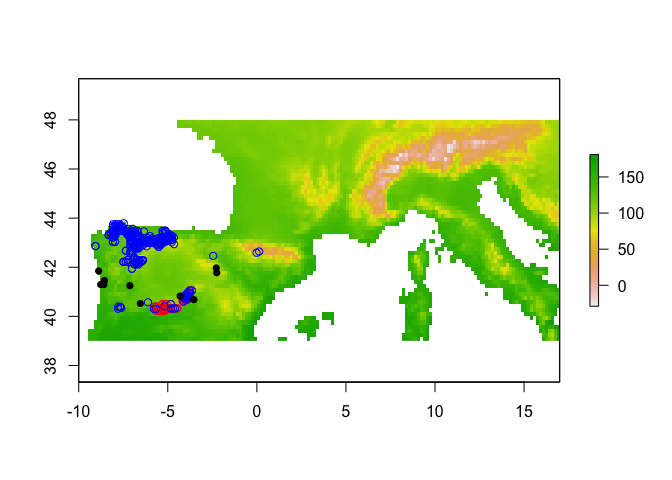
For the ribbon rangebreak test, you will need one extra thing; a third enmtools.species object representing the occurrence points (for one or both species) that fall within the ribbon of putatively unsuitable habitat. In the case of these two lizards we don't have such a ribbon, so we'll just simulate one based on some random points.
plot(env[[1]])
points(cyreni$presence.points, col = "red")
points(monticola$presence.points, col = "blue")
ribbon <- enmtools.species(species.name = "ribbon")
ribbon$presence.points <- data.frame(Longitude = runif(n = 10, min = -9, max = 0),
Latitude = runif(n = 10, min = 40.5, max = 42))
points(ribbon$presence.points, pch = 16)
ribbon$range <- background.raster.buffer(ribbon$presence.points, 20000, mask = env)
ribbon
##
##
## Range raster:
## class : RasterLayer
## dimensions : 54, 162, 8748 (nrow, ncol, ncell)
## resolution : 0.1666667, 0.1666667 (x, y)
## extent : -10, 17, 39, 48 (xmin, xmax, ymin, ymax)
## coord. ref. : +proj=longlat +datum=WGS84 +ellps=WGS84 +towgs84=0,0,0
## data source : in memory
## names : bio1
## values : 1, 1 (min, max)
##
##
##
## Presence points (first ten only):
##
## Longitude Latitude
## ---------- ---------
## -8.767481 41.29724
## -7.129208 41.25078
## -8.560360 41.45911
## -2.252357 41.77763
## -8.881490 41.84858
## -6.539390 40.52293
## -8.585099 41.29406
## -4.308883 40.82632
## -3.536658 40.67851
## -2.280341 41.96987
##
##
## Background points not defined.
##
## Species name: ribbon
Now we'll run a ribbon rangebreak test using GLM models with quadratic effects. We also need to tell it the width of the ribbons to generate for the replicates. The units for the width argument are the same units that the presence points are in; e.g., if the points are in decimal degrees you should supply the width of the barrier in decimal degrees.
rbr.glm <- rangebreak.ribbon(monticola, cyreni, ribbon, env, type = "glm", f = pres ~ poly(bio1, 2) + poly(bio12, 2) + poly(bio7, 2), width = 0.5, nreps = 4)
##
##
## No background points provided, drawing background from range raster.
##
##
##
## No background points provided, drawing background from range raster.
##
##
##
## No background points provided, drawing background from range raster.
##
##
## Building empirical models...
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
## Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
## Adding environmental data to species ribbon
## Processing presence points...
## Processing background points...
## Adding environmental data to species outside
## Processing presence points...
## Processing background points...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Building replicate models...
##
## Replicate 1 ...
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
## Adding environmental data to species ribbon
## Processing presence points...
## Processing background points...
## Adding environmental data to species outside
## Processing presence points...
## Processing background points...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Replicate 2 ...
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
## Adding environmental data to species ribbon
## Processing presence points...
## Processing background points...
## Adding environmental data to species outside
## Processing presence points...
## Processing background points...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Replicate 3 ...
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
## Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
## Adding environmental data to species ribbon
## Processing presence points...
## Processing background points...
## Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
## Adding environmental data to species outside
## Processing presence points...
## Processing background points...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
##
## Replicate 4 ...
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
## Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
## Adding environmental data to species ribbon
## Processing presence points...
## Processing background points...
## Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
## Adding environmental data to species outside
## Processing presence points...
## Processing background points...
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
## [1] "Trying to find starting conditions, attempt 1"
## [1] "Building replicates..."
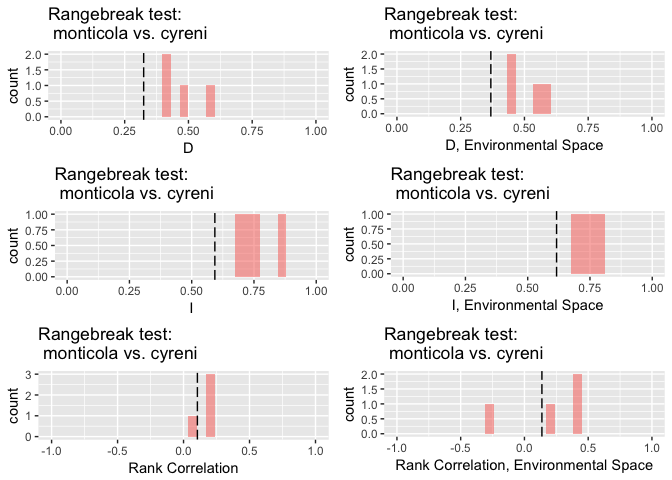
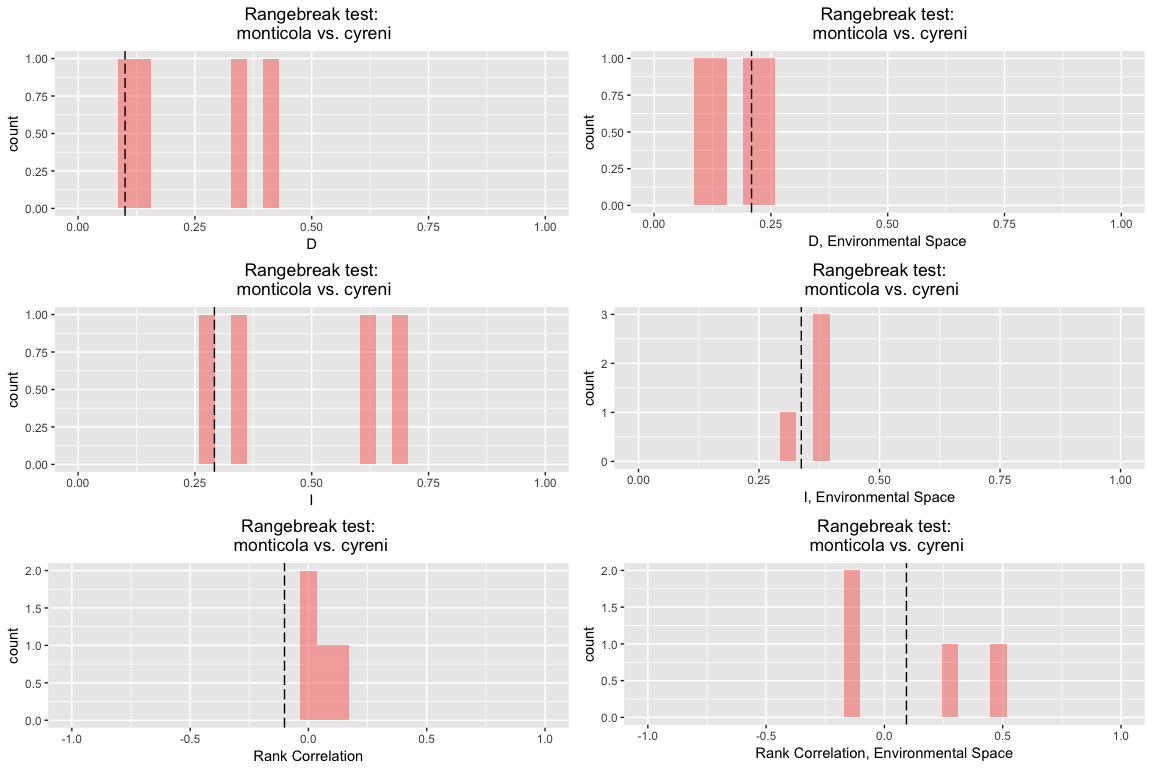
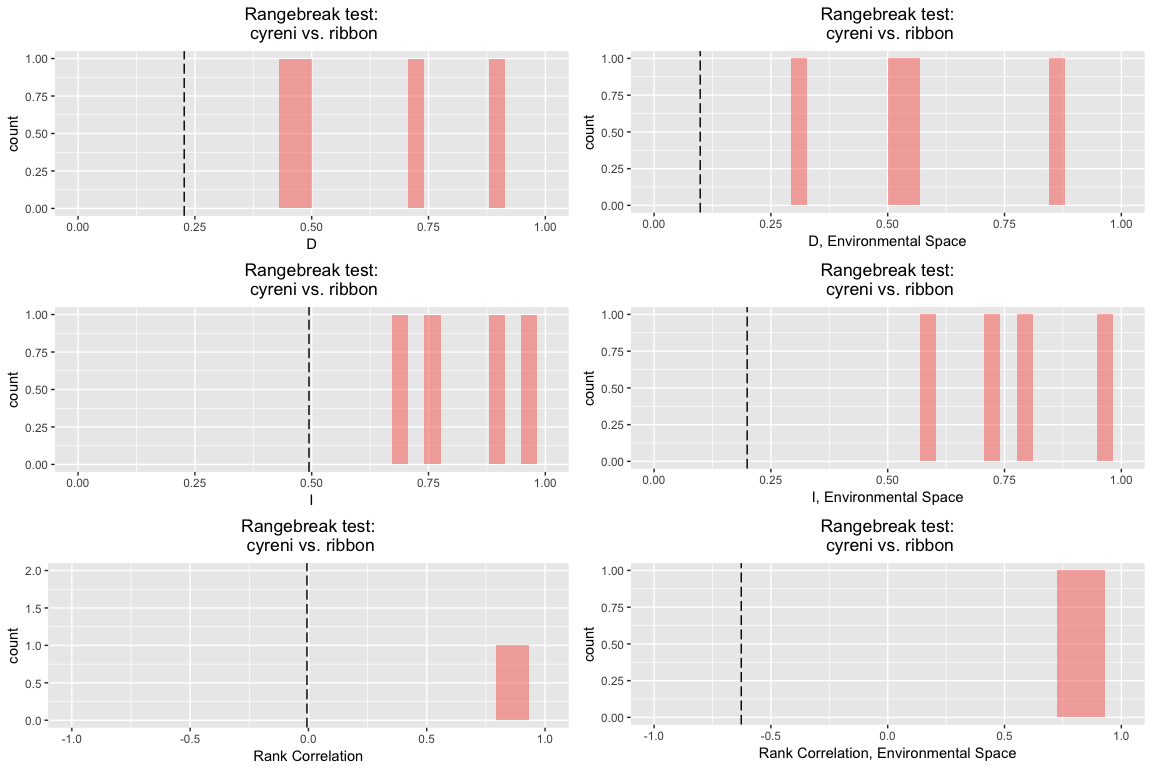
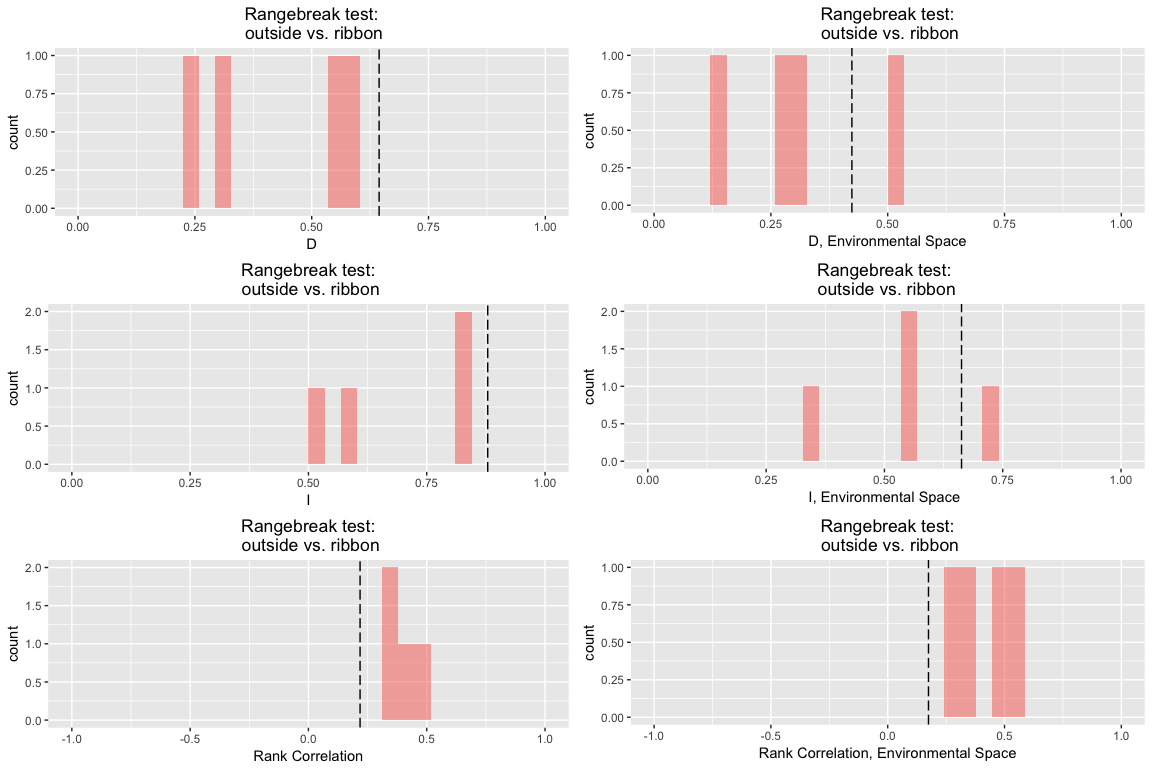
rbr.glm
##
##
##
##
## ribbon rangebreak test monticola vs. cyreni
##
## rangebreak test p-values...
##
## Species 1 vs. Species 2:
## D I rank.cor env.D env.I env.cor
## 0.8 0.8 0.4 1.2 0.8 1.2
##
## Species 1 vs. Ribbon:
## D I rank.cor env.D env.I env.cor
## 0.4 0.4 0.4 0.4 0.4 0.4
##
## Species 2 vs. Ribbon:
## D I rank.cor env.D env.I env.cor
## 0.4 0.4 0.4 0.4 0.4 0.4
##
## Outside vs. Ribbon:
## D I rank.cor env.D env.I env.cor
## 0.4 0.4 0.4 0.8 0.8 0.4
##
##
## Replicates:
##
## Species 1 vs. Species 2:
## D I rank.cor env.D env.I env.cor
## empirical 0.1005914 0.2917623 -0.10114941 0.2085903 0.3376812 0.09302784
## rep 1 0.3565518 0.6355876 -0.02596101 0.2230470 0.3737421 -0.12110338
## rep 2 0.4104463 0.6761612 0.08390495 0.2489801 0.3822046 -0.14229292
## rep 3 0.1213811 0.3424171 0.13038532 0.1501157 0.3719485 0.45153350
## rep 4 0.0902216 0.2843522 0.03205496 0.1009398 0.2933058 0.25625175
##
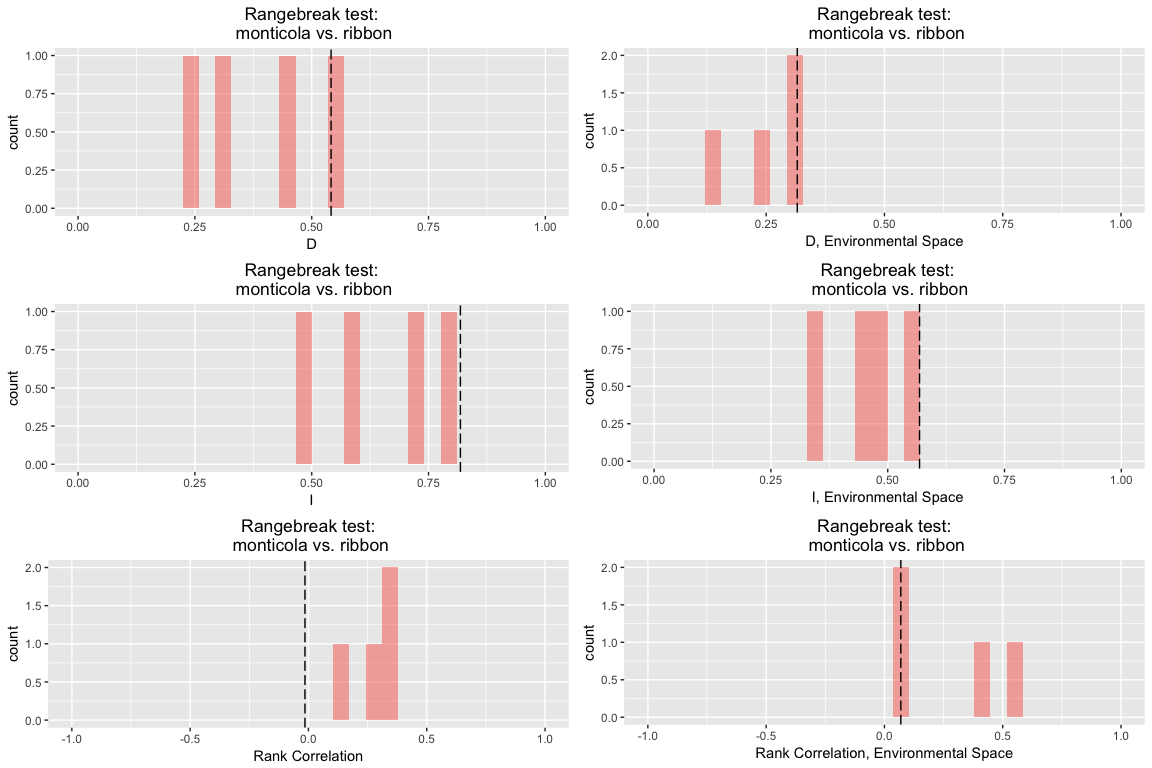
## Species 1 vs. Ribbon:
## D I rank.cor env.D env.I env.cor
## empirical 0.5414841 0.8182575 -0.01472933 0.3157483 0.5681497 0.06923811
## rep 1 0.4468853 0.7307242 0.12674612 0.3126485 0.4898966 0.09526138
## rep 2 0.5364706 0.7818077 0.35937225 0.2548644 0.4567155 0.07342587
## rep 3 0.2277833 0.4805389 0.24684606 0.1355698 0.3490069 0.38817853
## rep 4 0.3071671 0.5701145 0.37629252 0.2941895 0.5521452 0.53269544
##
## Species 2 vs. Ribbon:
## D I rank.cor env.D env.I env.cor
## empirical 0.2271375 0.4942254 -0.006247587 0.09873118 0.1992178 -0.6268091
## rep 1 0.8877557 0.9808929 0.972142787 0.85482266 0.9521763 0.9597145
## rep 2 0.7236891 0.9066791 0.823449461 0.54225084 0.7802574 0.8955788
## rep 3 0.4828211 0.7527276 0.932253465 0.50798800 0.7113498 0.8374995
## rep 4 0.4361225 0.6995519 0.870359442 0.32311605 0.5853179 0.7575722
##
## Outside vs. Ribbon:
## D I rank.cor env.D env.I env.cor
## empirical 0.6442788 0.8788862 0.2182055 0.4233755 0.6629133 0.1745643
## rep 1 0.5592673 0.8266039 0.3906555 0.5056916 0.7089894 0.4978725
## rep 2 0.5820008 0.8144199 0.4767257 0.3168624 0.5440349 0.2482848
## rep 3 0.2510722 0.5079192 0.3177119 0.1276517 0.3374701 0.3715698
## rep 4 0.3027025 0.5692949 0.3240719 0.2907300 0.5496800 0.5307634
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## Warning: Removed 1 rows containing missing values (geom_bar).
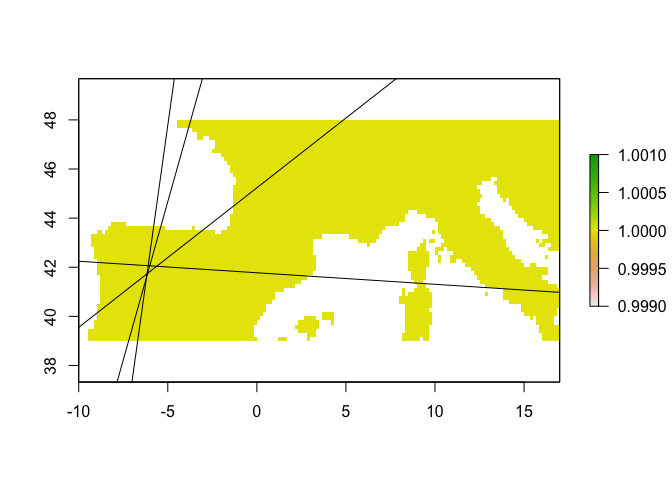
Note that the output table here has slope, intercept, and intercept offset.
rbr.glm$lines.df
## slope intercept offset
## 1 -1.0966462 34.70667 0.3710317
## 2 -0.6354215 39.14627 0.2962010
## 3 0.9441800 49.63428 0.3438273
## 4 0.1388375 44.14309 0.2523980
The intercept denotes the intercept corresponding to the CENTER of each ribbon. To get the lines denoting the edges of the ribbons (for example if you want to plot the ribbons on a map), you add and substract the offset. In other words, the top edge of the ribbon is given by y = (slope * x) + intercept + offset, while the bottom edge is given by y = (slope * x) + intercept - offset.
Some of the tests in ENMTools, including some really neat ones that are still in development, require you to build an enmtools.clade object. These objects are simply lists that contain a phylogeny and a set of enmtools.species objects. It's important that the names of the species objects and their species.name attributes match the names in the phylogeny's tip.labels. For demonstration, we're going to build an object for a clade of five anoles from Hispaniola. We have the tree, so we're just going to grab occurrence data from GBIF using the rgbif package.
library(rgbif)
library(ape)
##
## Attaching package: 'ape'
## The following objects are masked from 'package:raster':
##
## rotate, zoom
hisp.anoles <- read.nexus(file = "test/testdata/StarBEAST_MCC.species.txt")
keepers <- c("brevirostris", "marron", "caudalis", "websteri", "distichus")
hisp.anoles <- drop.tip(phy = hisp.anoles, tip = hisp.anoles$tip.label[!hisp.anoles$tip.label %in% keepers])
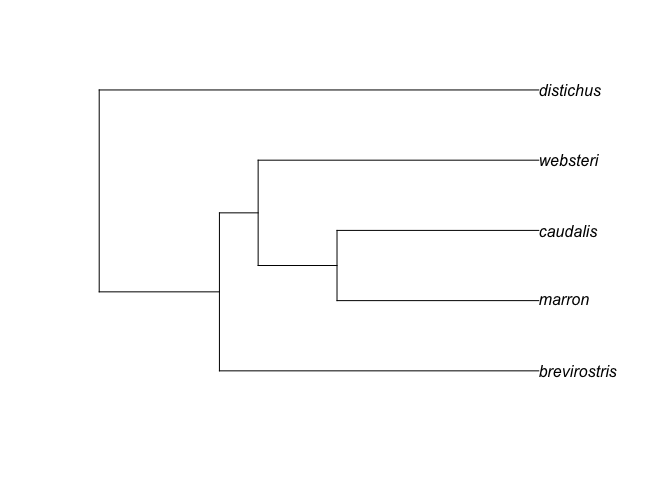
plot(hisp.anoles)
So there's our tree. Now we're going to grab some environmental data.
hisp.env <- stack(list.files("test/testdata/Hispaniola_Worldclim", full.names = TRUE))
hisp.env <- setMinMax(hisp.env)
And then we'll create a function to build species from GBIF.
# Automate the process of downloading data and removing duds and dupes
species.from.gbif <- function(genus, species, name = NA, env){
# Name it after the species epithet unless told otherwise
if(is.na(name)){
name <- species
}
# Get GBIF data
this.sp <- enmtools.species(presence.points = gbif(genus = genus, species = species)[,c("lon", "lat")],
species.name = name)
# Rename columns, get rid of duds
colnames(this.sp$presence.points) <- c("Longitude", "Latitude")
this.sp$presence.points <- this.sp$presence.points[complete.cases(extract(env, this.sp$presence.points)),]
this.sp$presence.points <- this.sp$presence.points[!duplicated(this.sp$presence.points),]
this.sp$range <- background.raster.buffer(this.sp$presence.points, 50000, mask = hisp.env)
return(this.sp)
}
Now we'll create five species and add them to a species.clade object that is called brev.clade.
brevirostris <- species.from.gbif(genus = "Anolis", species = "brevirostris", env = hisp.env)
marron <- species.from.gbif(genus = "Anolis", species = "marron", env = hisp.env)
caudalis <- species.from.gbif(genus = "Anolis", species = "caudalis", env = hisp.env)
websteri <- species.from.gbif(genus = "Anolis", species = "websteri", env = hisp.env)
distichus <- species.from.gbif(genus = "Anolis", species = "distichus", env = hisp.env)
brev.clade <- enmtools.clade(species = list(brevirostris, marron, caudalis, websteri, distichus), tree = hisp.anoles)
check.clade(brev.clade)
##
##
## An enmtools.clade object with 5 species
##
## Species names:
## brevirostris caudalis distichus marron websteri
##
## Tree:
##
## Phylogenetic tree with 5 tips and 4 internal nodes.
##
## Tip labels:
## [1] "brevirostris" "caudalis" "distichus" "marron"
## [5] "websteri"
##
## Rooted; includes branch lengths.
##
##
## Data Summary:
##
##
## species.names in.tree presence background range
## ------------- -------------- -------- --------- ----------- --------
## brevirostris brevirostris TRUE 173 0 present
## caudalis caudalis TRUE 21 0 present
## distichus distichus TRUE 678 0 present
## marron marron TRUE 12 0 present
## websteri websteri TRUE 17 0 present
That's one way to build a clade object by hand, but there's already one built into ENMTools to experiment with so we'll just use that.
data(iberolacerta.clade)
The AOC tests allow you to examine patterns of range, point, and ENM overlap in the context of a phylogeny. This is effectively a generalized version of several analyses: age-range correlation (e.g., Fitzpatrick and Turelli 2006), ENM overlap in the context of a phylogeny (e.g., Knouft et al. 2006, Warren et al. 2008), and point overlaps (e.g., Cardillo and Warren 2016).
These tests require the creation of an enmtools.clade object, as above. AOC tests consist of two steps: first, the average overlap at each node in the phylogeny is calcualted using a method that takes tree topology into account (see Fitzpatrick and Turelli 2006), then we perform a linear regression to measure the relationship between node age and average overlap. Due to the fact that these overlaps violate many of the assumptions of a regular linear regression, however (e.g., errors are not iid), we can't calculate significance in the typical way. Instead we performa Monte Carlo test, permuting the identity of the tips of the tree and repeating the node averaging and modeling steps. Finally we measure statistical significance by comparing the empirical slope and intercept to the distribution of slopes and intercepts from the Monte Carlo replicates.
First, let's do one using geog.range.overlaps, as in Fitzpatrick and Turelli 2006. Note that this analysis requires that each of your species have a range raster stored in their species object (we did that as part of the function used above).
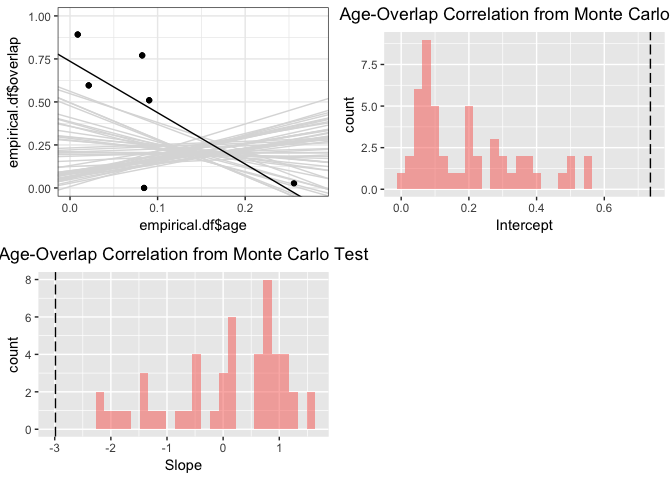
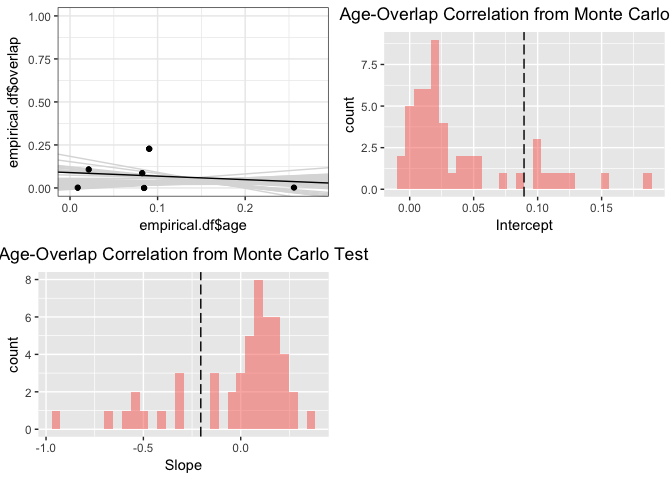
range.aoc <- enmtools.aoc(clade = iberolacerta.clade, nreps = 50, overlap.source = "range")
summary(range.aoc)
##
##
## Age-Overlap Correlation test
##
## 50 replicates
##
## p values:
## (Intercept) empirical.df$age
## 0.03921569 0.03921569
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## NULL
Now we can do one using point overlaps just by changing the overlap.source argument:
point.aoc <- enmtools.aoc(clade = iberolacerta.clade, nreps = 50, overlap.source = "points")
summary(point.aoc)
##
##
## Age-Overlap Correlation test
##
## 50 replicates
##
## p values:
## (Intercept) empirical.df$age
## 0.3921569 0.4313725
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## NULL
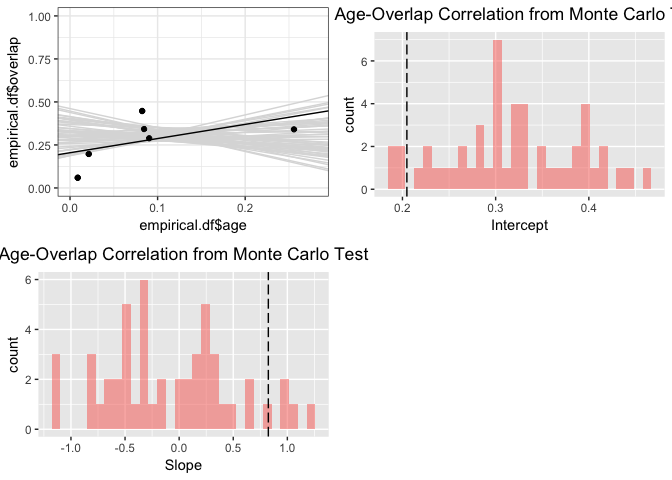
Or we can use similarity between ENMs built for each species. Here we'll use GLM models:
glm.aoc <- enmtools.aoc(clade = iberolacerta.clade, env = env, nreps = 50, overlap.source = "glm", f = pres ~ poly(bio1, 2) + poly(bio12, 2))
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species monticola
## Processing presence points...
## Processing background points...
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species martinezricai
## Processing presence points...
## Processing background points...
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species cyreni
## Processing presence points...
## Processing background points...
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species horvathi
## Processing presence points...
## Processing background points...
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species aurelioi
## Processing presence points...
## Processing background points...
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species aranica
## Processing presence points...
## Processing background points...
##
##
## No background points provided, drawing background from range raster.
##
## Adding environmental data to species bonnali
## Processing presence points...
## Processing background points...
summary(glm.aoc)
##
##
## Age-Overlap Correlation test
##
## 50 replicates
##
## p values:
## (Intercept) empirical.df$age
## 0.1960784 0.2352941
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## `stat_bin()` using `bins = 30`. Pick better value with `binwidth`.
## NULL
*Broennimann, O., Fitzpatrick, M. C., Pearman, P. B., Petitpierre, B., Pellissier, L., Yoccoz, N. G., Thuiller, W., Fortin, M.-J., Randin, C., Zimmermann, N. E., Graham, C. H. and Guisan, A. (2012), Measuring ecological niche overlap from occurrence and spatial environmental data. Global Ecology and Biogeography, 21: 481–497. doi:10.1111/j.1466-8238.2011.00698.x*
Fitzpatrick, B. M., & Turelli, M. (2006). The geography of mammalian speciation: mixed signals from phylogenies and range maps. Evolution, 60(3), 601-615.
Knouft, J. H., Losos, J. B., Glor, R. E., & Kolbe, J. J. (2006). Phylogenetic analysis of the evolution of the niche in lizards of the Anolis sagrei group. Ecology, 87(sp7).
Levins, R. 1968. Evolution In Changing Environments. Monographs in Population Biology, volume 2. Princeton University Press, Princeton, New Jersey, USA.
Schoener, T. W. 1968. Anolis lizards of Bimini: resource partitioning in a complex fauna. Ecology 49:704- 726.
Warren, D.L., R.E. Glor, and M. Turelli. 2008. Environmental niche identity versus conservatism: quantitative approaches to niche evolution. Evolution 62:2868-2883. doi: 10.1111/j.1558-5646.2008.00482.x
Warren, D.L., M. Cardillo, D.F. Rosauer, and D.I. Bolnick. 2014. Mistaking geography for biology: inferring processes from species distributions. Trends in Ecology and Evolution 29 (10), 572-580. doi: 10.1016/j.tree.2014.08.003