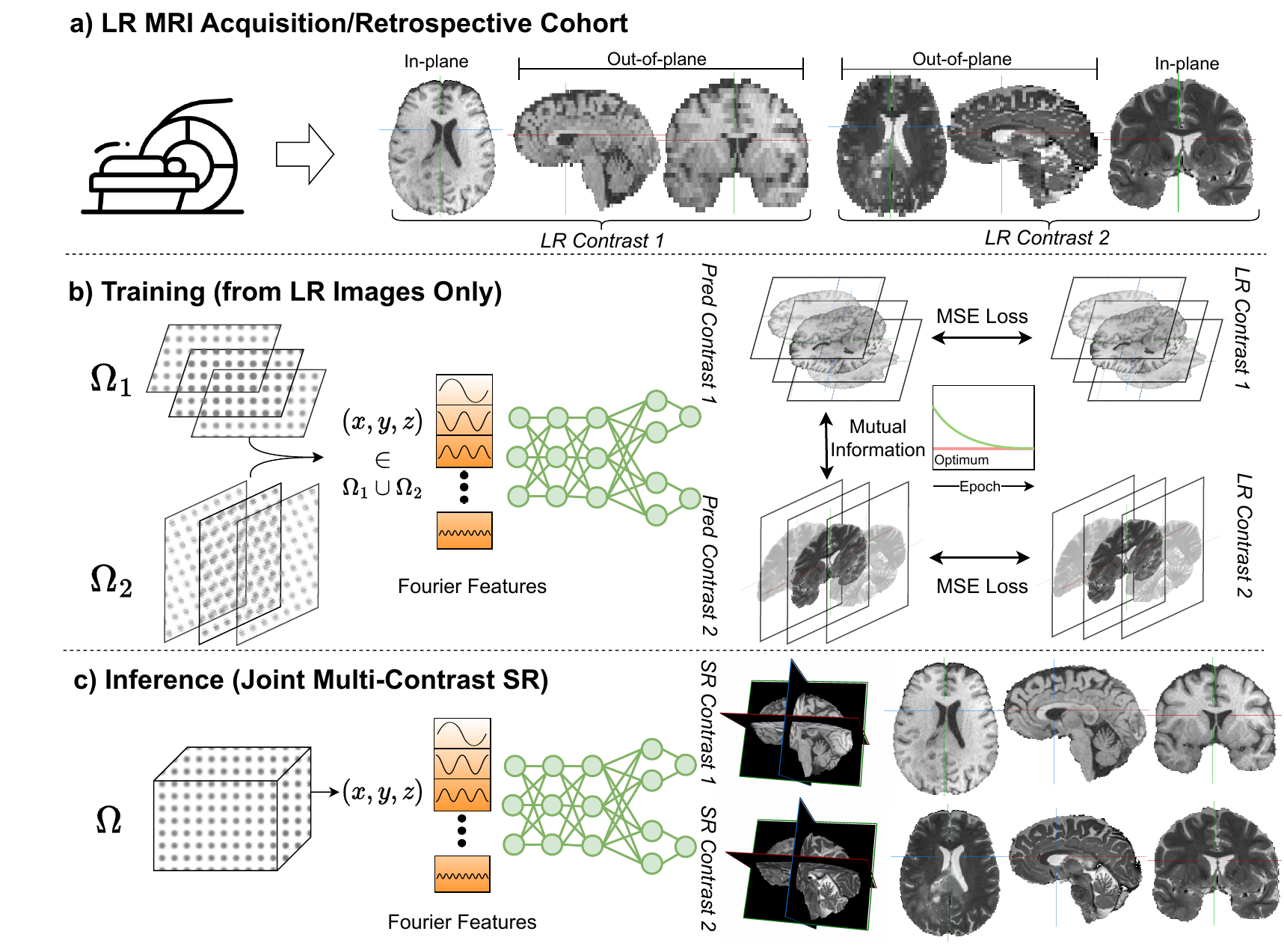
We conduct our experiments on three different Datasets:
- BRATS (25 patients)
- MSSEG-2016 (25 patients)
- in-house MS Dataset (cMS) (25 patients)
├── msseg-test-center07-06
│ └── 01
│ ├── sub-msseg-test-center07-06_ses-01_space-mni_brainmask.nii.gz
│ ├── sub-msseg-test-center07-06_ses-01_space-mni_flair_LR.nii.gz
│ ├── sub-msseg-test-center07-06_ses-01_space-mni_flair_mask_LR.nii.gz
│ ├── sub-msseg-test-center07-06_ses-01_space-mni_flair.nii.gz
│ ├── sub-msseg-test-center07-06_ses-01_space-mni_t1_LR.nii.gz
│ ├── sub-msseg-test-center07-06_ses-01_space-mni_t1_mask_LR.nii.gz
│ └── sub-msseg-test-center07-06_ses-01_space-mni_t1.nii.gz
### BRATS 2019
├── BraTS19_CBICA_AZH_1
│ ├── BraTS19_CBICA_AZH_1_brainmask.nii.gz
│ ├── BraTS19_CBICA_AZH_1_flair_LR.nii.gz
│ ├── BraTS19_CBICA_AZH_1_flair_mask_LR.nii.gz
│ ├── BraTS19_CBICA_AZH_1_flair.nii.gz
│ ├── BraTS19_CBICA_AZH_1_t1_LR.nii.gz
│ ├── BraTS19_CBICA_AZH_1_t1_mask_LR.nii.gz
│ ├── BraTS19_CBICA_AZH_1_t1.nii.gz
│ ├── BraTS19_CBICA_AZH_1_t2_LR.nii.gz
│ ├── BraTS19_CBICA_AZH_1_t2_mask_LR.nii.gz
│ └── BraTS19_CBICA_AZH_1_t2.nii.gz
├── m203013
│ └── 20200227
│ ├── sub-m203013_ses-20200227_space-mni_brainmask.nii.gz
│ ├── sub-m203013_ses-20200227_space-mni_dir_LR.nii.gz
│ ├── sub-m203013_ses-20200227_space-mni_dir_mask_LR.nii.gz
│ ├── sub-m203013_ses-20200227_space-mni_dir.nii.gz
│ ├── sub-m203013_ses-20200227_space-mni_flair_LR.nii.gz
│ ├── sub-m203013_ses-20200227_space-mni_flair_mask_LR.nii.gz
│ ├── sub-m203013_ses-20200227_space-mni_flair.nii.gz
│ ├── sub-m203013_ses-20200227_space-mni_t1_LR.nii.gz
│ ├── sub-m203013_ses-20200227_space-mni_t1_mask_LR.nii.gz
│ └── sub-m203013_ses-20200227_space-mni_t1.nii.gz
We provide an enviornment file that can be used to setup a conda environment. As our implementation is purely based on PyTorch, Scikit-Learn, Numpy, Nibabel, Pyyaml and the lpips repo, it should be easily possible to use other (or older) versions of libaries and CUDA, and tailor the environment to your needs.
As we train on single subjects, we decided to integrate training and inference into one python file, main.py.
Essentially, we run inference after every run - to log the performance of the isotropically upsampled image.
Please feel free to modify the codebase according to your needs or application.
To run the code, please execute:
python3 main.py --logging --config configs/"your_custom_config.yaml
To run our code for your datasets, please create a config file that you pass as an argument --config configs/"your_custom_config.yaml.
We provide all of our experiment configurations, with the following convention:
├── config_brats_ctr1.yaml -> Single Contrast for Contrast 1 (i.e. one output channel)
├── config_brats_ctr2.yaml -> Single Contrast for Contrast 2 (i.e. one output channel)
├── config_brats_mlpv2.yaml -> Multi Contrast Model with Split_Head Architecture (best performing model)
├── config_brats.yaml -> Multi Contrast Model without Split_Head Architecture (vanilla MLP, ablation)
To log to wandb, please use the --logging option.
Please cite this work if any of our code or ideas are helpful for your research.
@inproceedings{mcginnisandshit2023single,
title={Single-subject Multi-contrast MRI Super-resolution via Implicit Neural Representations},
author={McGinnis, Julian and Shit, Suprosanna and Li, Hongwei Bran and Sideri-Lampretsa, Vasiliki and Graf, Robert and Dannecker, Maik and Pan, Jiazhen and Stolt-Ans{\'o}, Nil and M{\"u}hlau, Mark and Kirschke, Jan S and others},
booktitle={International Conference on Medical Image Computing and Computer-Assisted Intervention},
pages={173--183},
year={2023},
organization={Springer}
}