v0.9.3 (12 Aug 2008)
Author: John Quinn
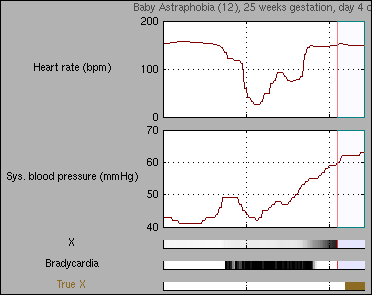
This demo implements a factorial dynamical Bayesian network and applies it to physiological monitoring data from babies receiving intensive care. A set of known artifactual and physiological patterns can be detected, as well as periods of clinically significant novelty.
Data in this demo comes from 15 babies who were treated in the Neonatal Intensive Care Unit at the Royal Infirmary of Edinburgh, UK. For each baby, there is a 24 hour sequence of vital signs data taken at 1-second intervals, including heart rate, blood pressures, saturation of gases in the blood and temperatures.
Details about the model and data can be found in:
- J.A. Quinn, C.K.I. Williams and N. McIntosh, Factorial Switching Linear Dynamical Systems applied to Condition Monitoring in Neonatal Intensive Care, IEEE Transactions on Pattern Analysis and Machine Intelligence 31(9), 2008. [pdf]
- J.A. Quinn, Bayesian Condition Monitoring in Neonatal Intensive Care, PhD thesis, University of Edinburgh, 2007. [pdf]
The data used in the experiments is contained in the file data/15days.mat. The
struct array data has two fields, raw and preprocessed, each of which is a
cell array with elements representing each of the the 15 babies. These elements
are also struct arrays, with fields containing the raw physiological data for each baby, and other information such as gestation and anonymised identifiers.
The struct array intervals contains annotations provided by the clinical
experts. For example, intervals.BloodSample{3} contains an array of times
for which a blood sample was thought to have occurred for baby 3. This is an
n × 2 matrix in which each row represents [start_index, stop_index] for a
particular episode of blood sampling. Indices are relative to the start of the 24 hour monitoring period.
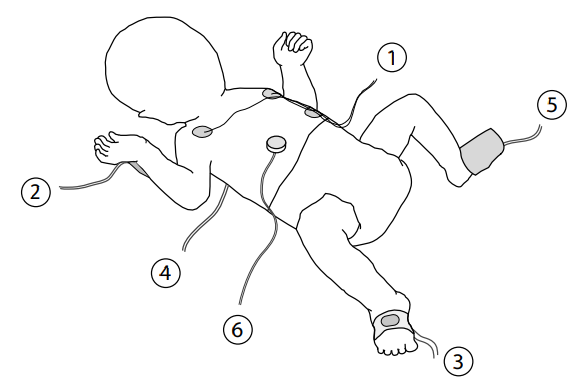
Probes used to collect vital signs data from an infant in intensive care: 1) three-lead ECG, 2) arterial line (connected to blood pressure transducer), 3) pulse oximeter, 4) core temperature probe (underneath shoulder blades), 5) peripheral temperature probe, 6) transcutaneous probe.
Start Matlab and run the command chooseexperiment.
Experiment with the system by editing the files in the
settings directory.
See doc/overview.pdf for high-level details of the code.
Bayes Net Toolbox: http://bnt.sourceforge.net/ (for the 'learn_kalman' function)
Matlab Signal Processing Toolbox: http://www.mathworks.com/products/signal/ (for the 'spectrum' function).
Rao-Blackwellised Particle Filtering Code: http://www.cs.ubc.ca/~nando/software/demorbpfdbn.tar.gz (for the 'deterministicR' function).