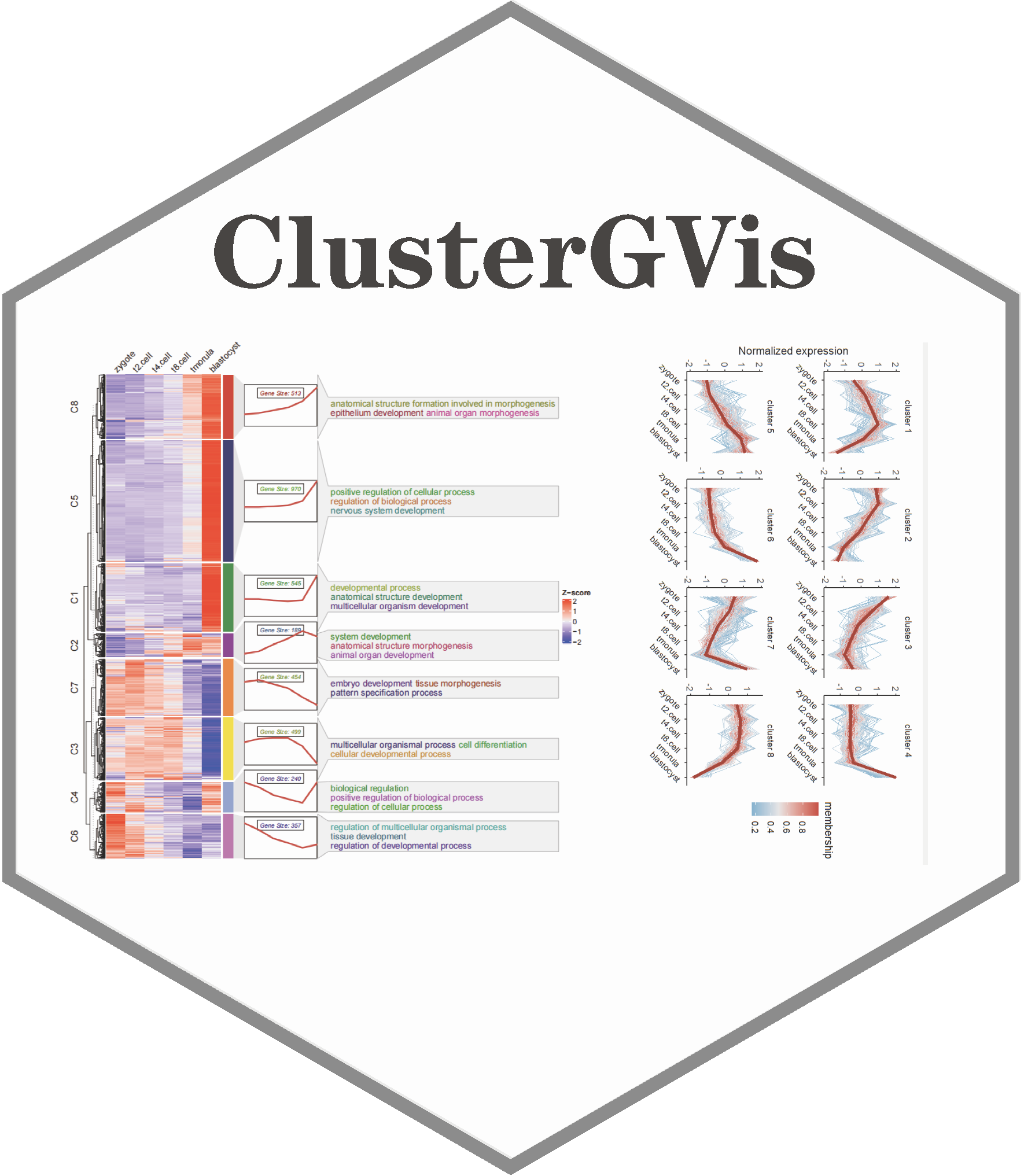
To enhance clustering and visualization of time-series gene expression data from RNA-Seq experiments, we present the ClusterGVis package. This tool enables concise and elegant analysis of time-series gene expression data in a simple, one-step operation. Additionally, you can perform enrichment analysis for each cluster using the enrichCluster function, which integrates seamlessly with clusterProfiler. ClusterGVis empowers you to create publication-quality visualizations with ease.
Thanks for the contributions for clusterProfiler and ComplexHeatmap!
There are some R package to make sure have been installed for better installing ClusterGVis:
BiocManager::install("ComplexHeatmap")
BiocManager::install("clusterProfiler")
BiocManager::install("TCseq")
BiocManager::install("Mfuzz")
BiocManager::install("monocle")
devtools::install_github('cole-trapnell-lab/monocle3')
install.packages("circlize")
install.packages("Seurat")You can install the development version of ClusterGVis like so:
# install from cran
install.packages("ClusterGVis")
# Note: please update your ComplexHeatmap to the latest version!
# install.packages("devtools")
devtools::install_github("junjunlab/ClusterGVis")Jun Zhang (2022). ClusterGVis: One-step to Cluster and Visualize Gene Expression Matrix. https://github.com/junjunlab/ClusterGVis
The comprehensive documentation: https://junjunlab.github.io/ClusterGvis-manual/
https://github.com/junjunlab/ClusterGvis-app
- ClusterGVis 对基因表达时间序列聚类和可视化
- ClusterGVis 的问题解答及优化
- ClusterGVis 对指定 cluster 添加注释
- ClusterGVis 继续查缺补漏
- ClusterGVis 添加多个样本注释
- bugs 报告和修复
- ClusterGVis 对接单细胞啦
- 听说你还想添加 KEGG 注释?
- ClusterGVis 对接 monocle2 拟时序热图
- 听说你想画个 monocle3 的拟时序热图?
- ClusterGVis 添加自定义图形注释
- 听说你想插入 GO 和 KEGG 图形注释?
- enrichCluster 关于非模式物种富集分析的使用
- 听说你想给 visCluster 添加行注释?
- ClusterGVis shiny 交互式 web 来了?