AScall is a web tool providing automatic processing of allele specific real-time PCR experiments: variation conducted real-time whereis each allele is detected by independent reaction separated into individual wells. The positive outcome of the reaction for a particular allele is judged by the increase in the fluorescent signal. This tool is written in R language with graphical user interface based on shiny technology. In addition to general purpose R packages, our program uses a number of specific PCR-oriented packages: chipPCR, qPCR, RDML, shinyMolBio. GUI use standard shiny elements, shinyWidgets for multiply selectors - pickerInput, shinyMolBio for PCR plate - pcrPlateInput and PCR curves plot - renderAmpCurves.
General note: some of operations are time consuming (e.g. curves preprocession). So be patient!!! To avoid recalculations after each option change all setup is applied by clicking Recalculate Results button.
Copy all files (or init.R, generics.R, server.R and ui.R for minimal installation) to any directory on your computer and run as usual shiny application. All necessary packages will be installed during application run.
Process of analysis can be described by several major steps:
- data import from PCR machines via RDML format;
- optional PCR curves preprocessing;
- overall experiment quality control and individual sample control;
- sample genotype calling.
Sample files are available in
examples folder of AScall.
These files are BioRad CFX pcrd format and rdml converted. Also there is
manual genotyping results file - genotyping.xlsx. Click button Use Sample File
to load 01.rdml file.
You can input one or more files in a format supported by the RDML package: rdml, lc96p, xlsx, etc. But AScall need correct sample naming to work as automatic genotype caller. Naming convention and export will be shown for BioRad CFX Manager 3.1 as example. Several files can be imported at once – in such case all subsequent analyses are carried out independently for each plate and one summary table with results is provided (control gene and preprocessing settings will be the same for all plates!).
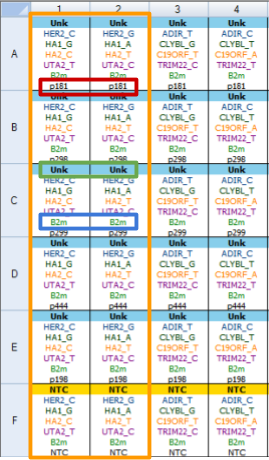
Correct plate setup is shown on the fig.1:
There are several major elements:
- All wells with one sample have to be named equaly. Marked with the red box p181 on the fig.1: despites the different names of targets - sample names are equal. Same rule for replicates - there should not be any indices!
- Target names for not control genes have to contain gene name and allele name after underscore - GENENAME_ALLELENAME pattern. Green box on the figure - HER2_C and HER2_G where HER2 is the gene name and C or G are alleles. For indel targets use + sign (e.g. UGT2b17_+). Using indel targets leads to alternate analysis: no amplification is deletion; amplification is insertion.
- Control gene name have to be equal for all plates. Blue box B2m.
- No template controls must have NTC sample type.
- Target name without allele name called marker.
- All wells with the same markers AScall interpretes as one kit. Orange box on the fig.1.
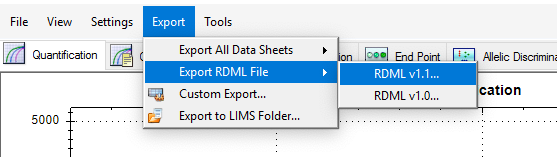
After plate setup is done you can import data by Export>Export RDML File>RDML v1.1 menu (see fig.2).
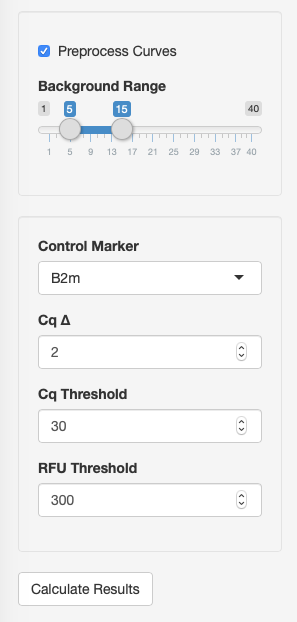
This is optional step and only needed when your use RAW data or want to recalculate Cq values with the independent method (fitting and Cq calculation). Preprocessing is conducted with the chipPCR and qpcR packages usage. You should check Preprocess Curves to enable preprocessing and options (Fig.3).
After that several additional option Background Region appears. You can set signal background region (linear part of the curves before exponentional growth) for all curves using this slider (Click Recalculate Results to apply changes).
Processing consists of the following three steps:
- Background subtraction
chipPCR::CPP(fpoints$cyc, fpoints$fluor,
trans = TRUE,
bg.range = bgRange)$y.norm- Model fitting
qpcR::pcrfit(fpoints[, c("cyc", "fluor")],
cyc = 1, fluo = 2,
model = get(modelType))- Cq calculation
qpcR::efficiency(fitted, plot = FALSE,
type = cqMethod)After loading all data files Recalculate Results button appears and subsequent analisys can be done. First of all you can fine tune several options (Fig.3):
- Control Marker - select any detected marker as control marker - reaction that have to be positive in all samples.
- Cq ∆ - maximum difference between Cq values of reactions replicates.
- Cq Threshold - max Cq values to be reaction treated as positive.
- RFU Threshold - minimum fluorescence signal to be reaction treated as positive.
- Low RFU - all reactions with RFU lower than RFU Threshold are marked with
RFU_QC = "Low"
RFU_QC = ifelse(endPt < input$rfuThr,
"Low", "Ok"))- Mark amplification status - negative for reactions with
RFU_QC != "Ok"and Cq higher than Cq Threshold
ampStatus_QC = ifelse(RFU_QC != "Ok" | cq > input$cqThr,
"NoAmp", "Ok")- Replicate match check - all replicates have to be
ampStatus_QC = "NoAmp"or (ampStatus_QC = "Ok"and difference between Cq values lower than Cq ∆ option)
meanCq = mean(cq),
deltaCq = max(cq) - min(cq)
replicateMatch_QC = {
if ((all(ampStatus_QC == "Ok") && deltaCq[1] < input$cqDelta) ||
all(ampStatus_QC != "Ok")) "Ok" else "Fail"- ∆ between alleles - alleles Cq difference (and minus delta between ctrlMarkerCqs to normalize samples) have to be lower than Cq ∆ option.
dTbl <- dTbl %>%
group_by(position) %>%
mutate(
ctrlMarkerCq = cq[marker == input$ctrlMarker]
)
dTbl <- dTbl %>%
group_by(kit, marker, sample) %>%
mutate(
allelesDeltaCqUnnorm = meanCq - min(meanCq),
allelesDeltaCq = allelesDeltaCqUnnorm -
(ctrlMarkerCq - min(ctrlMarkerCq)),
allelesDeltaCq_QC =
ifelse(abs(allelesDeltaCq) < input$cqDelta,
"Ok", "Fail"))- NTC no amplification - all NTC reactions in kit have to be
ampStatus_QC = "NoAmp"
noAmpNTC_QC =
{
if (any(ampStatus_QC[sample.type == "ntc"] == "Ok"))
"Fail" else "Ok"
}- Control Marker QC -
ampStatus_QC = "Ok"andreplicateMatch_QC = "Ok"for Control Marker reactions
ctrlMarker_QC =
{
if (any(replicateMatch_QC[marker == input$ctrlMarker] != "Ok") ||
any(ampStatus_QC[marker == input$ctrlMarker] == "NoAmp"))
"Fail" else "Ok"
}- Kit total QC -
noAmpNTC_QC != "Ok"for all NTC
kit_QC =
{
if (any(noAmpNTC_QC != "Ok"))
"Fail" else "Ok"
}- Results Calc - calculates for all sample which are
kit_QC == "Ok" & replicateMatch_QC == "Ok" & ctrlMarker_QC == "Ok". Then result is a combination of alleles withampStatus_QC == "Ok"or insertion/deletion ifampStatus_QC == "Ok"/ampStatus_QC != "Ok"for indel markers.
genResult <- function(okAlleles) {
okAlleles <- unique(okAlleles)
if (length(okAlleles) == 1) {
okAlleles <- c(okAlleles, okAlleles)
}
paste(okAlleles, collapse = "/")
}
genIndelResult <- function(ampStatus_QC) {
if (all(ampStatus_QC == "Ok")) {
"Ins"
} else if (all(ampStatus_QC != "Ok")) {
"Del"
} else {
"Error"
}
}
result = {
if (marker[1] != input$ctrlMarker) {
if (allele[1] == "+") {
genIndelResult(ampStatus_QC)
} else {
genResult(allele[ampStatus_QC == "Ok"])
}
} else {
""
}
}
resultZygosity =
sapply(result,
function(x)
{
if (x[1] == "")
""
else
switch(
as.character(str_split(x,
"/")[[1]] %>%
unique() %>%
length()),
"1" = "Homo",
"2" = "Hetero",
"Error")
})There are three main elements:
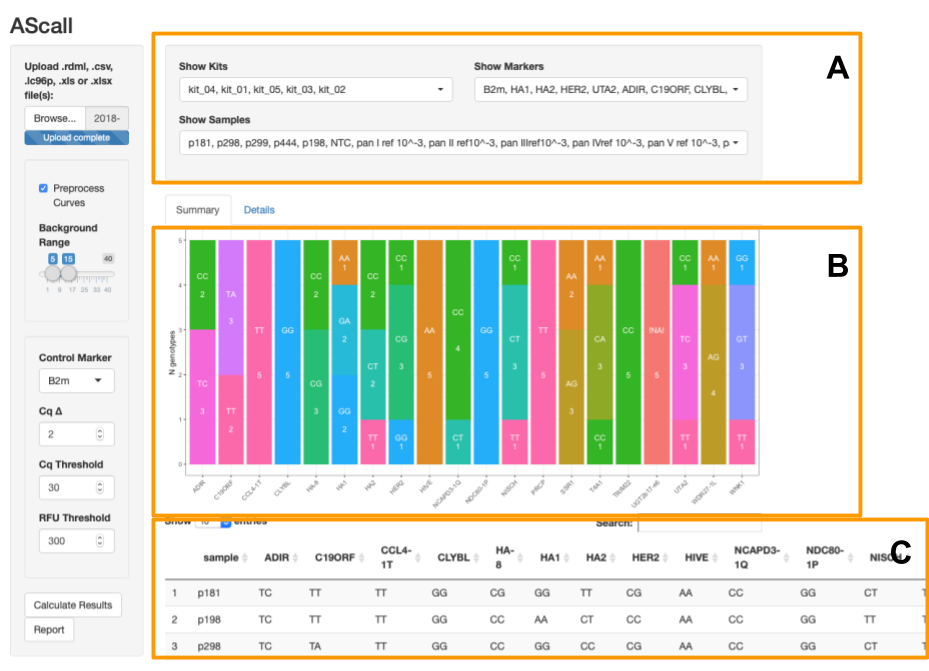
- Global filters - serve to filter kits, samples and markers for summary and details views (fig.4A).
- Summary view - global genotyping results for all PCR files.
- Details view - per file view of PCR curves and plate with QC results.
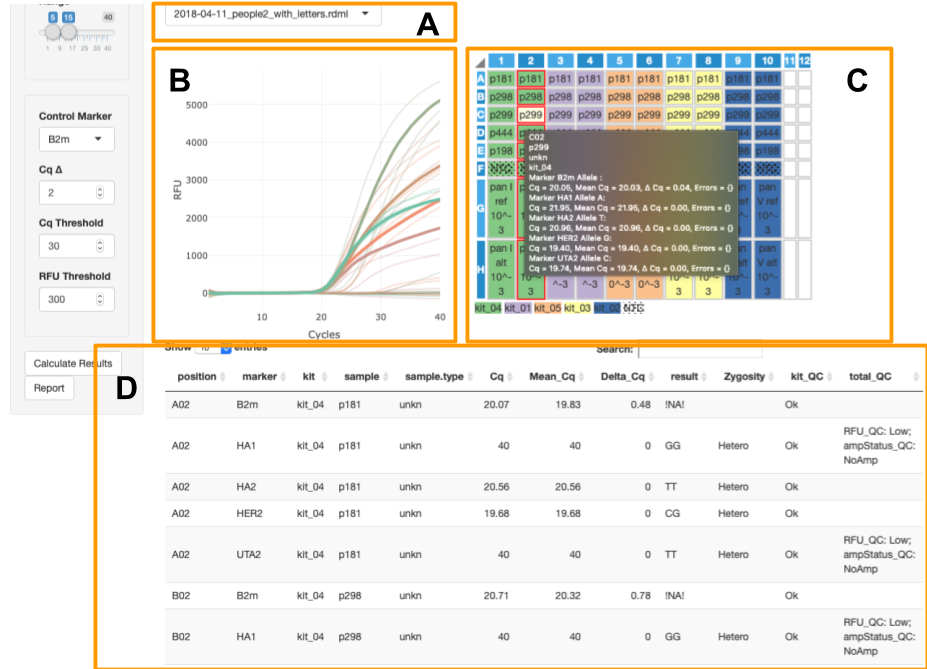
Global filters allow to select individual samples, markers or kits for viewing at details and summary (fig.4A).
This view shows all genotyping results for all loaded files as bar plot and table. Barplot allows to overview results by markers: x-axis is marker and y-axis is number of samples grouped by genotypes of this marker (fig.4B). Table represents genotyping results by samples (fig.4C).
Provides access to additional per plate details about PCR curves and analysis. It consists of:
- File selector - you can switch uploaded experiments by this elements (fig.5A).
- PCR curves - created by
shinyMolBio::renderAmpCurves()function (fig.5B). Curves are colored by marker. - PCR plate - created by
shinyMolBio::pcrPlateInput()function (fig.5C). Wells are color by kit; dotted wells contain NTC (see legend under plate). Selected wells have red border and light yellow background is for on hover well (selection filters curves on PCR curves plot and the on hover curves are solid while the others are - transparent). On hover well provides additionalinfo inside black box. - Details table - show information about every PCR curve including genotyping results and QC (fig.5D).
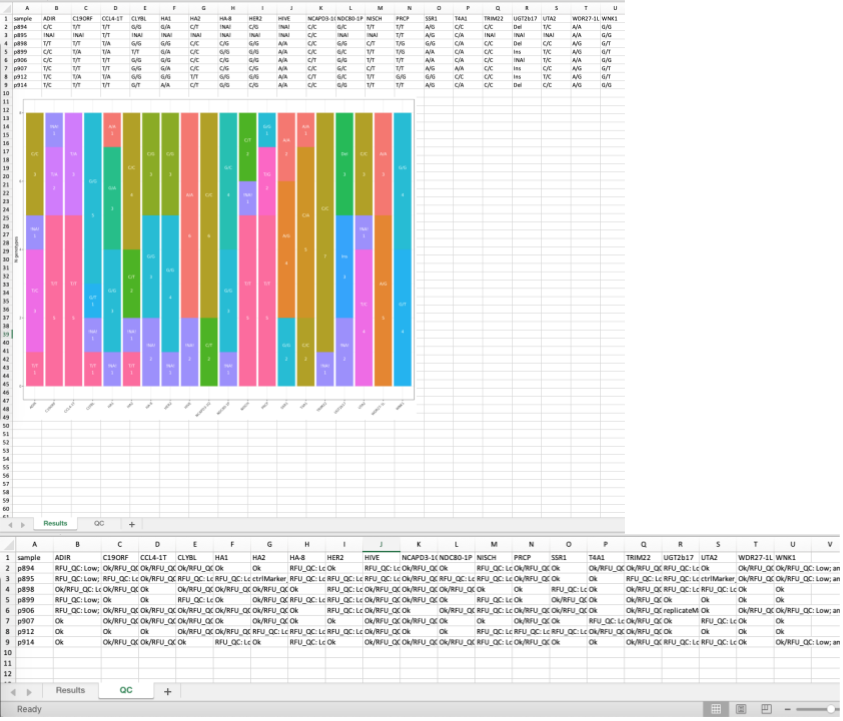
After genotype analysis summary report can be generated by clicking Report button. Xlsx report consists of (fig.6):
- Genotyping results table
- Genotypes count plot
- QC table
- Analysis settings
Report generation is carried out by openxlsx package.
- RFU_QC
- Low - curve endpoint fluorescence signal is lower than RFU Threshold
- Ok
- ampStatus_QC
- NoAmp - anmlification for this curve is not detected
- Ok
- replicateMatch_QC
- Fail - Cq difference between replicates is bigger than Cq ∆ option
- Ok
- allelesDeltaCq_QC
- Fail - Cq difference between alleles of one marker is bigger than Cq ∆ option
- Ok
- noAmpNTC_QC
- Fail - any NTC sample has positive amplification
- Ok
- ctrlMarker_QC
- Fail - control marker does not have positive amplification in any well
- Ok