Environment variables found in principles.yml (conda env)
FeatureMatching.py
Below Code is found in main.py
[F,mask]=cv2.findFundamentalMat(kp1,kp2, method=3,ransacReprojThreshold=3.0,confidence=0.99)
[E1,mask]=cv2.findEssentialMat(kp1,kp2,cameraMatrix=K, method=cv2.RANSAC, prob=0.999, threshold=3.0 )
[E2,mask]=cv2.findEssentialMat(kp2,kp1,cameraMatrix=K, method=cv2.RANSAC, prob=0.999, threshold=3.0 )[_, R1, t1, mask] = cv2.recoverPose(E1, kp1,kp2, cameraMatrix=K,mask=mask)
[_, R2, t2, mask] = cv2.recoverPose(E2, kp2,kp1, cameraMatrix=K,mask=mask)
projMatr1=np.column_stack((R1,t1))
projMatr2=np.column_stack((R2,t2))R Recovered relative rotation, 3x3 matrix. t Recovered relative translation, 3x1 vector.
mask Output mask for inliers in points1 and points2. In the output mask only inliers which pass the cheirality check.
mat_4D=cv2.triangulatePoints(projMatr1=projMatr1,projMatr2=projMatr2,projPoints1=kp1,projPoints2=kp2).astype(np.float64)
mat_3D = mat_4D[:,:3]/mat_4D[:,3:4]changing to 3D according to since we require projection in 3D space. they're just 3D points in a 4D projective space, analogous to 2D points in a 3D projective space. all points (x,y,z,1) * w, for arbitrary nonzero w, in the projective space represent the same 3D point (x,y,z), and (x,y,z,1) is the canonical representative. https://stackoverflow.com/questions/69429075/what-could-be-the-reason-for-triangulation-3d-points-to-result-in-a-warped-para
used solvePnPRefineLM() for better control of outliers
r, t=solvePnPRefineLM()(points_3D,points_2D,K,dist_coeffs,r1,t1,criteria=criteria)
R,jacobian=cv2.Rodrigues(r)
pnp_mat=np.hstack([R,t])
CurrentPose=np.dot(K,pnp_mat)Least squares with soft l1 loss: rho(z) = 2 * ((1 + z)**0.5 - 1)
res1=least_squares(RMSE, r.flatten(),jac='2-point', method='dogbox',loss='soft_l1',max_nfev=2000) #implemented least square optimization
Breadth search based on the number of matched features. Sorted by the most number of features and expanded.
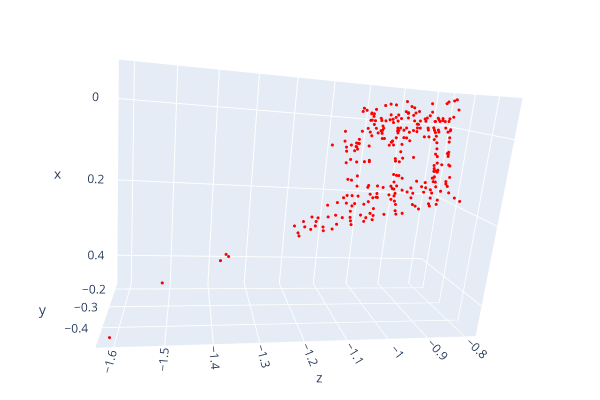
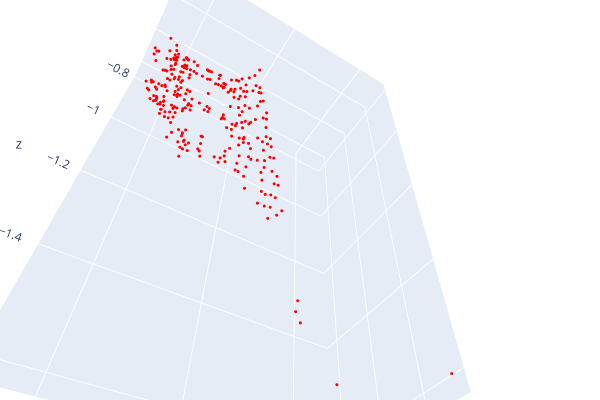
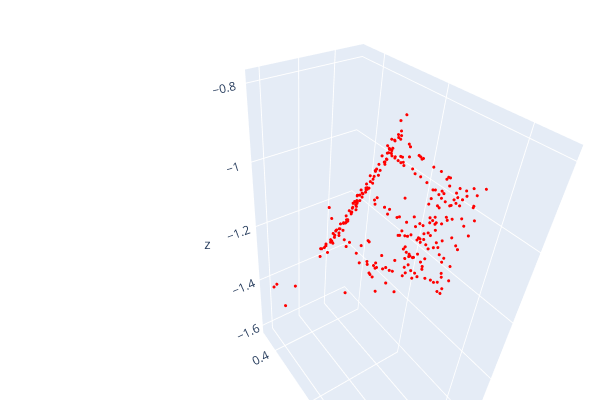
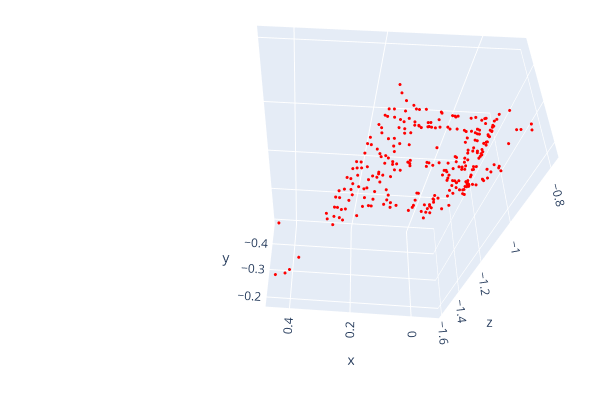
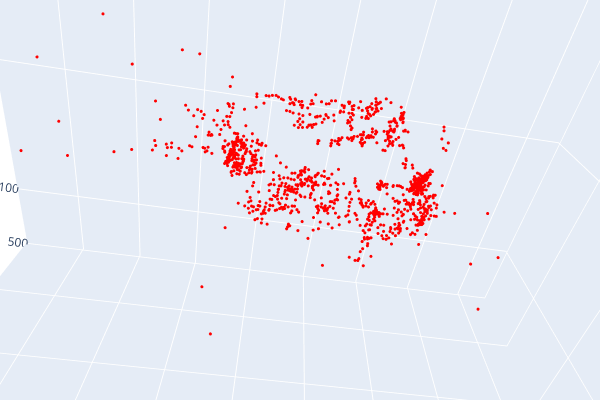
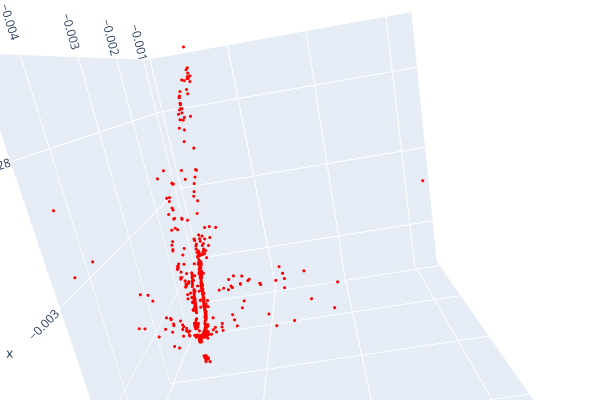
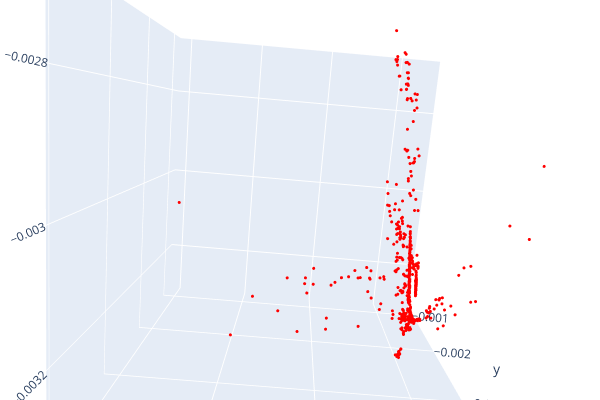
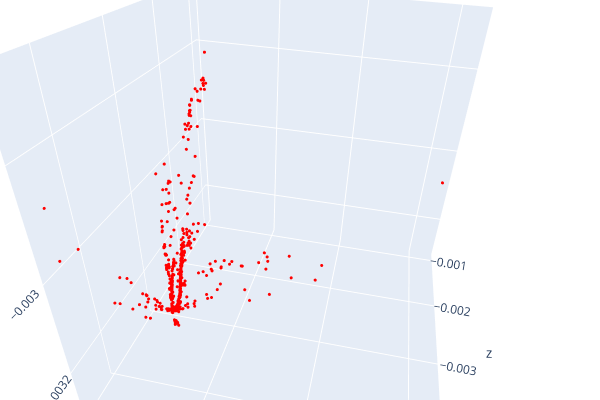
Open3d point cloud print("Load a ply point cloud, print it, and render it") pcd = o3d.geometry.PointCloud()
Datasets processed in the file Dataload.py
Intrinsic matrix constructed from pose information:
K=np.array([[fx,0,cx], [0 ,fy, cy],[0, 0, 1]]).astype(np.float64) ##intrinsic matrixRMSE_r+=sum(np.power(r.flatten()-r0.flatten(),2)) #calculate RMSE r and update
TOTAL_RMSE_r = np.sqrt( (1/len(images)**2)* RMSE_r): RMSE for TRex rotation matrix: 0.0015021174744879215 RMSE for TRex translation vector 14627410.962035134 Average loss 1.752949594479218e-12
--------------------------------------------------------------------------------
Loaded 54 poses from: kitti_trex_gt.txt
Loaded 54 poses from: kitti_trex_00.txt
--------------------------------------------------------------------------------
Aligning using Umeyama's method...
Rotation of alignment:
[[ 0.44750528 0.89419974 -0.01207684]
[ 0.72676047 -0.37151271 -0.57775213]
[-0.5211125 0.24977015 -0.81612293]]
Translation of alignment:
[ 45.31216322 -10.67234959 6.78472044]
Scale correction: 1.0
--------------------------------------------------------------------------------
Compared 54 absolute pose pairs.
Calculating APE for translation part pose relation...
--------------------------------------------------------------------------------
APE w.r.t. translation part (m)
(with SE(3) Umeyama alignment)
max 537.908827
mean 172.434304
median 53.996346
min 32.582834
rmse 243.921671
sse 3212880.198135
std 172.523020RMSE for Fern rotation matrix:14187.827955083636 RMSE for Fern translation vector: 6697.837169699395
--------------------------------------------------------------------------------
Loaded 19 poses from: kitti_trex_gt.txt
Loaded 19 poses from: kitti_trex_00.txt
--------------------------------------------------------------------------------
Aligning using Umeyama's method...
Rotation of alignment:
[[-0.70920034 -0.70429341 0.03171228]
[-0.67597208 0.66652846 -0.31432716]
[ 0.20024141 -0.24435755 -0.94878489]]
Translation of alignment:
[17.08704688 24.74655038 -7.53542607]
Scale correction: 1.0
--------------------------------------------------------------------------------
Compared 19 absolute pose pairs.
Calculating APE for translation part pose relation...
--------------------------------------------------------------------------------
APE w.r.t. translation part (m)
(with SE(3) Umeyama alignment)
max 472.822399
mean 177.974571
median 33.815540
min 20.278510
rmse 258.609392
sse 1270697.532936
std 187.626943