There is a recent explosion of deep learning algorithms that to tackle the computational problem of predicting drug treatment outcome from baseline molecular measurements. To support this,we have built a benchmark dataset that harmonizes diverse datasets to better assess algorithm performance.
This package collects diverse sets of paired molecular datasets with corresponding drug sensitivity data. All data here is reprocessed and standardized so it can be easily used as a benchmark dataset for the This repository leverages existing datasets to collect the data required for deep learning model development. Since each deep learning model requires distinct data capabilities, the goal of this repository is to collect and format all data into a schema that can be leveraged for existing models.
The goal of this repository is two-fold: First, it aims to collate and standardize the data for the broader community. This requires running a series of scripts to build and append to a standardized data model. Second, it has a series of scripts that pull from the data model to create model-specific data files that can be run by the data infrastructure.
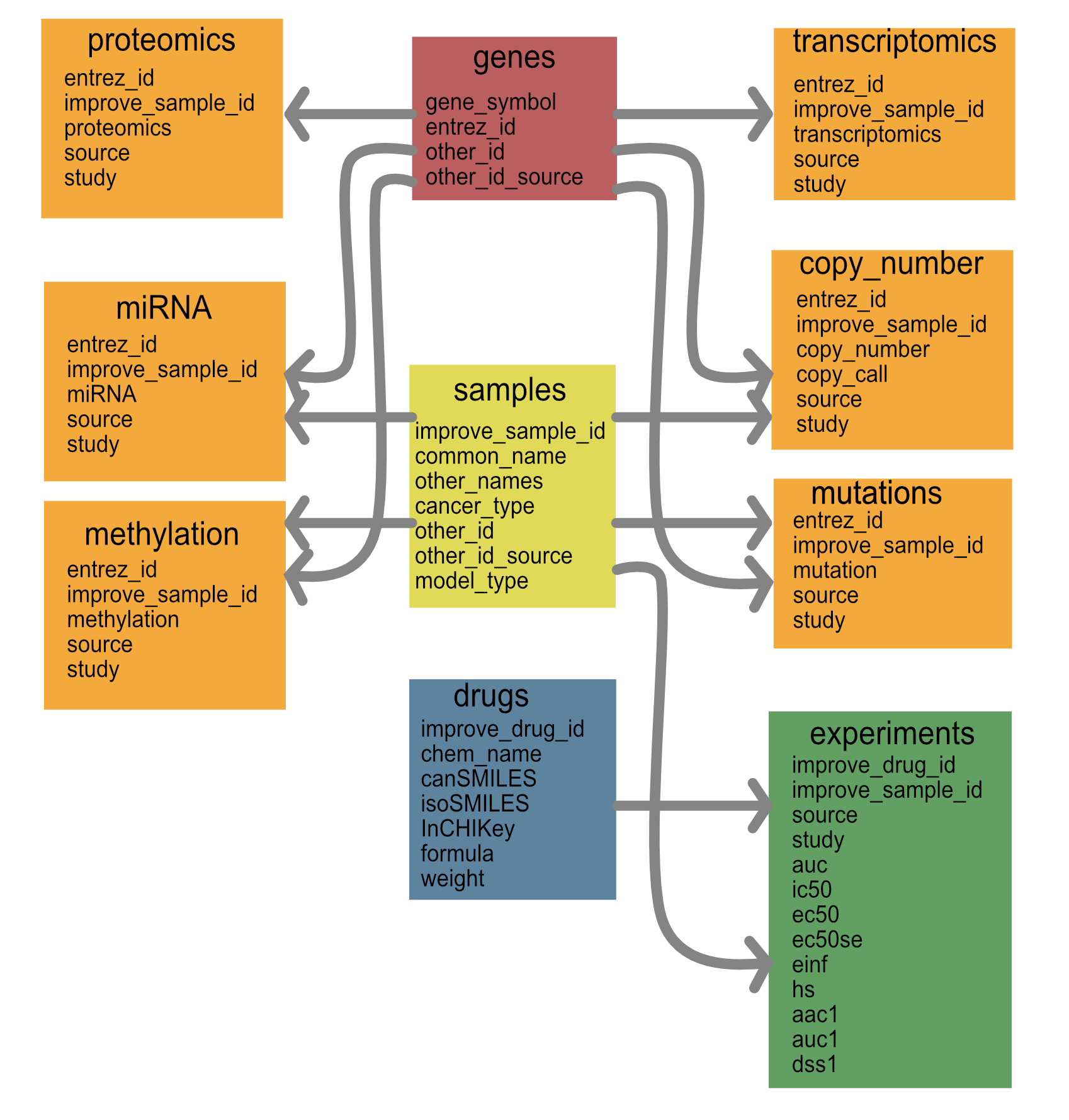
The goal of the data model is to collate drug response data together with molecular data in a way that can be easily ingested by machine learning models. The overall schema is shown below.
We will store the data in tables that are represented by the files below. Each data-specific model can be generated from a smaller set of these tables. The schema for these tables is represented below.
For each dataset added, the files are comma-delimited and named follows:
- genes.csv
- drugs.tsv.gz --> Drug names have commas and quotes in them, therefore require tab delimited
- samples.csv
- experiments.csv.gz --> compressed to fit on github
- transcriptomics.csv.gz
- mutations.csv.gz
- copy_number.csv.gz
- methylation.csv.gz
- mirnas.csv.gz
Below is a description of how the data model is built.
| Data model step | Description/Dependencies | Script | Destination |
|---|---|---|---|
| Build cell line data | Runs through PGX and existing CCLE data to compile all values | cell_line/buildInitialDataset.py | [./cell_line] |
| Build cptac data | This uses the genes files created in the [./cell_line] directory but generates additional samples. | cptac/getCptacData.py | [./cptac] |
| Get HCMI data | This uses a fixed manifest to download the data into the proper schema | hcmi/getHCMIData.py | [./hcmi] |
What data is stored here?
Files are stored on FigShare. We need to build a script that pulls those data as needed.