A simple process automation tool created by the Gladfelter lab at UNC Chapel Hill to streamline fungal cell biology data analysis.
When working with microscopy data, the pain points for the lab are:
- organizing the data due to its large size
- adapting the data to 3rd party tools given its proprietary
*.nd2format - cropping ROIs given the complex, inter-weaving cells
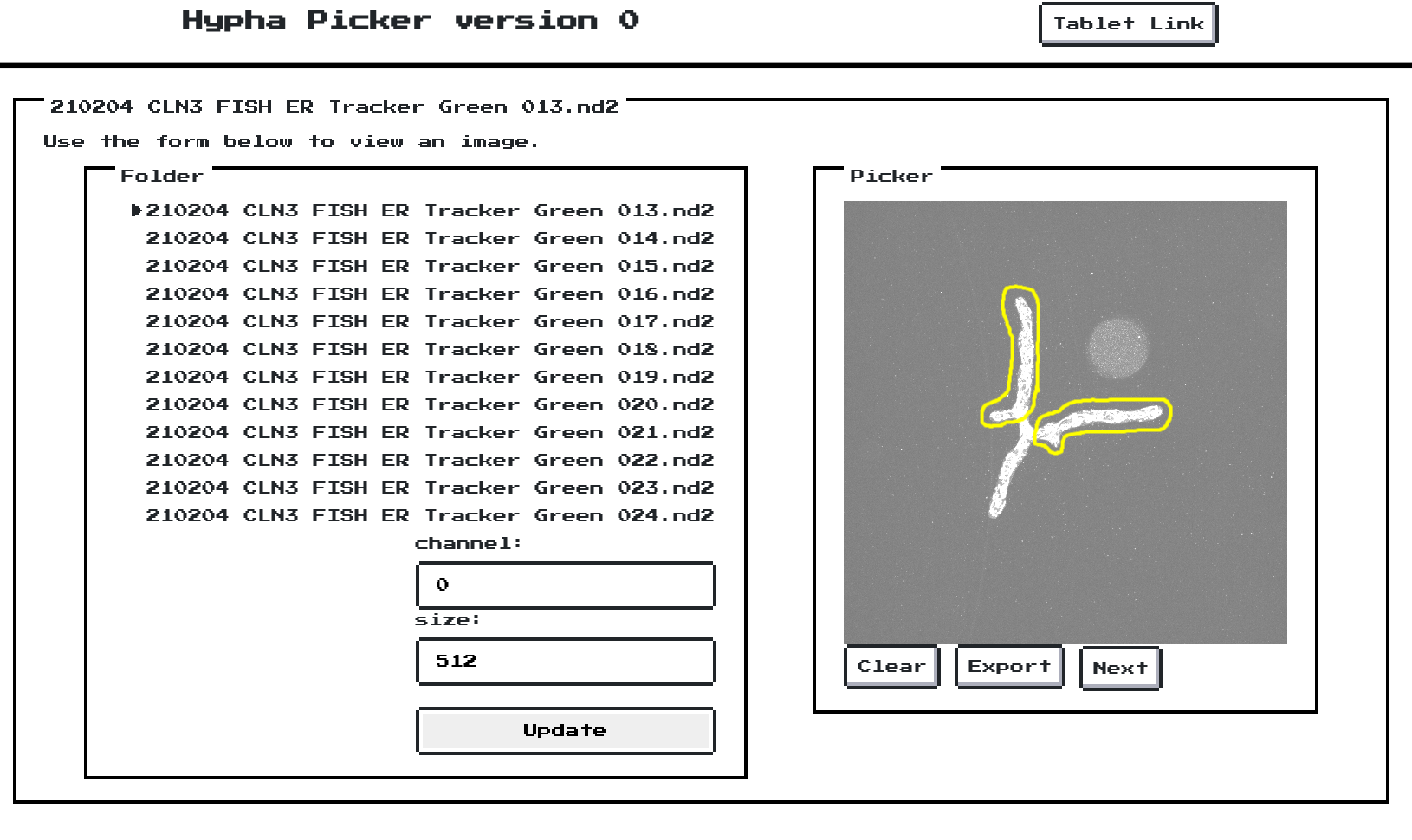
The tool crawls through folders of *.nd2 files and caches max projections of
each z-stack ahead-of-time. When ready, the user can launch a local web
interface to manually lasso-select ROIs that are cropped, padded, and dumped
into more-accessible *.TIFF files.
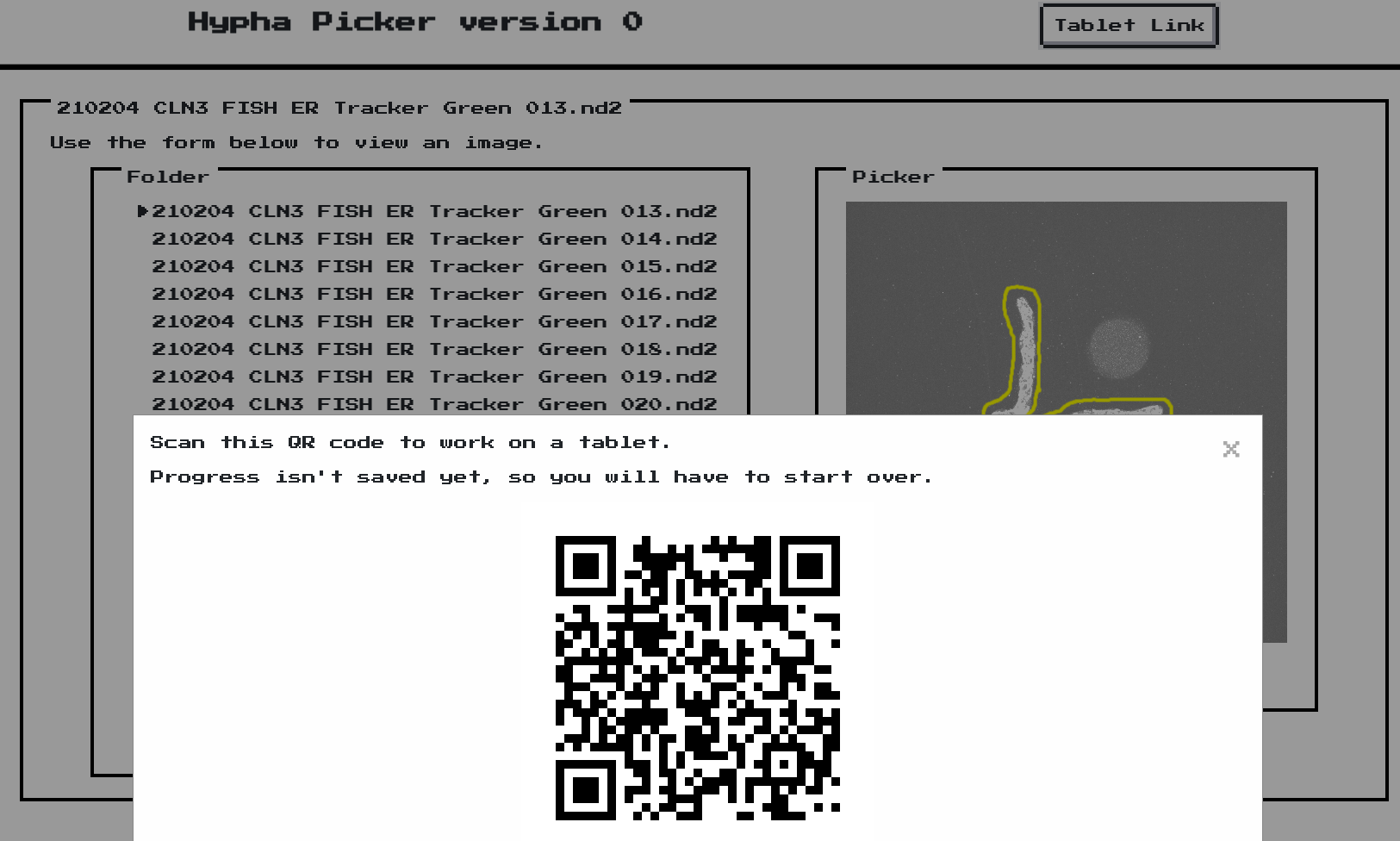
The web interface shows a QR code so that the user can draw these ROIs on a tablet if they so choose.
Refer to the gladlab-ip wiki.