ktweedie: Kernel-based Tweedie compound Poisson gamma model using
high-dimensional covariates for the analyses of zero-inflated response
variables.
ktweedie is a package that fits nonparametric Tweedie compound Poisson
gamma models in the reproducing kernel Hilbert space. The package is
based on two algorithms, the ktweedie for kernel-based Tweedie model
and the sktweedie for sparse kernel-based Tweedie model. The
ktweedie supports a wide range of kernel functions implemented in the
R package kernlab and the optimization algorithm is a
Broyden–Fletcher–Goldfarb–Shanno (BFGS) algorithm with bisection line
search. The package includes cross-validation functions for
one-dimensional tuning of the kernel regularization parameter
and for two-dimensional joint tuning of one kernel parameter and
.
The
sktweedie uses variable weights to achieve variable selection. It
is a meta-algorithm that alternatively updates the kernel parameters and
a set of variable weights.
The ktweedie solves the problem
where
is the index parameter,
is the dispersion parameter,
is an
kernel matrix computed according to the user-specified kernel function
,
whose entries are
are calculated based on the
-dimensional
predictors from subjects
.
In the kernel-based Tweedie model, the mean parameter
for the
-th
observation is modelled by
The sktweedie solves
where
involves variable weights
.
- From the CRAN.
install.packages("ktweedie")- From the Github.
devtools::install_github("ly129/ktweedie")First we load the ktweedie package:
library(ktweedie)The package includes a toy data for demonstration purpose. The
predictor matrix
x is generated from standard normal distribution and
y is generated according to
where
.
That said, only the first two predictors are associated with the
response.
data(dat)
x <- dat$x
y <- dat$yAn input matrix x and an output vector y are now loaded. The
ktd_estimate() function can be used to fit a basic ktweedie model.
The regularization parameter lam1 can be a vector, which will be
solved in a decreasing order with warm start.
fit.ktd <- ktd_estimate(x = x,
y = y,
kern = rbfdot(sigma = 0.1),
lam1 = c(0.01, 0.1, 1))
str(fit.ktd$estimates)
#> List of 3
#> $ lambda 1 :List of 3
#> ..$ fn : num 110
#> ..$ coefficient: num [1:30, 1] 0.5558 -0.062 -0.0381 0.0523 -0.0251 ...
#> ..$ convergence: int 0
#> $ lambda 0.1 :List of 3
#> ..$ fn : num 51
#> ..$ coefficient: num [1:30, 1] 1.662 -0.235 -0.177 0.867 -0.143 ...
#> ..$ convergence: int 0
#> $ lambda 0.01:List of 3
#> ..$ fn : num 39.2
#> ..$ coefficient: num [1:30, 1] 7.692 -0.49 -0.841 4.624 -0.696 ...
#> ..$ convergence: int 0fit.ktd$estimates stores the estimated coefficients and the final
objective function value. The estimated kernel-based model coefficients
for the
-th
lam1 can be accessed by the index l:
fit.ktd$estimates[[l]]$coefficient.
The function can also be used to fit the sktweedie model by setting
the argument sparsity to TRUE, and specifying the regularization
coefficient
in the argument
lam2.
fit.sktd <- ktd_estimate(x = x,
y = y,
kern = rbfdot(sigma = 0.1),
lam1 = 5,
sparsity = TRUE,
lam2 = 1)And we can access the fitted coefficients in a similar manner to the
fit.ktd. Additionally, the fitted variable weights
can be accessed by
fit.sktd$estimates[[1]]$weight
#> [,1]
#> [1,] 1.0000000
#> [2,] 0.4462078
#> [3,] 0.0000000
#> [4,] 0.0000000
#> [5,] 0.0000000Variables with weights close to 0 can be viewed as noise variables.
The ktweedie and sktweedie algorithms require careful tuning of one
to multiple hyperparameters, depending on the choice of kernel
functions. For the ktweedie, we recommend either a one-dimensional
tuning for lam1
()
or a two-dimensional random search for
lam1 and the kernel parameter
using cross-validation. Tuning is achieved by optimizing a
user-specified criterion, including log likelihood loss = "LL", mean
absolute difference loss = "MAD" and root mean squared error
loss = "RMSE". Using the Laplacian kernel as an example.
laplacedot(sigma = 1)
#> Laplace kernel function.
#> Hyperparameter : sigma = 1The one-dimensional search for the optimal lam1, can be achieved with
the ktd_cv() function from a user-specified vector of candidate
values:
ktd.cv1d <- ktd_cv(x = x,
y = y,
kern = laplacedot(sigma = 0.1),
lambda = c(0.0001, 0.001, 0.01, 0.1, 1),
nfolds = 5,
loss = "LL")
ktd.cv1d
#> $LL
#> 1 0.1 0.01 0.001 1e-04
#> -82.30040 -60.33054 -55.68177 -55.68835 -65.38823
#>
#> $Best_lambda
#> [1] 0.01The two-dimensional joint search for the optimal lam1 and sigma
requires ktd_cv2d(). In the example below, a total of ncoefs = 10
pairs of candidate lam1 and sigma values are randomly sampled
(uniformly on the log scale) within the ranges lambda = c(1e-5, 1e0)
and sigma = c(1e-5, 1e0), respectively. Then the nfolds = 5-fold
cross-validation is performed to select the pair that generates the
optimal cross-validation loss = "MAD".
ktd.cv2d <- ktd_cv2d(x = x,
y = y,
kernfunc = laplacedot,
lambda = c(1e-5, 1e0),
sigma = c(1e-5, 1e0),
nfolds = 5,
ncoefs = 10,
loss = "MAD")
ktd.cv2d
#> $MAD
#> Lambda=0.000435692, Sigma=0.174196 Lambda=0.00855899, Sigma=0.00201436
#> 354.1993 431.4734
#> Lambda=0.00518177, Sigma=0.000749782 Lambda=7.25693e-05, Sigma=0.0620986
#> 469.7289 327.0395
#> Lambda=0.0513091, Sigma=0.000344321 Lambda=0.0108477, Sigma=0.000277883
#> 626.3884 589.4097
#> Lambda=9.72691e-05, Sigma=2.19179e-05 Lambda=0.0682224, Sigma=0.000455657
#> 433.5755 624.1514
#> Lambda=0.000228745, Sigma=0.0247239 Lambda=0.166265, Sigma=0.00695988
#> 332.0113 544.0900
#>
#> $Best_lambda
#> [1] 7.25693e-05
#>
#> $Best_sigma
#> [1] 0.0620986Then the model is fitted using the hyperparameter(s) selected by the
ktd_cv() or ktd_cv2d(). In the example below, the selected lam1
and sigma values are accessed by ktd.cv2d$Best_lambda and
ktd.cv2d$Best_sigma, which are then be fed into the ktd_estimate()
to perform final model fitting.
ktd.fit <- ktd_estimate(x = x,
y = y,
kern = laplacedot(sigma = ktd.cv2d$Best_sigma),
lam1 = ktd.cv2d$Best_lambda)
str(ktd.fit$estimates)
#> List of 1
#> $ lambda 7.25693e-05:List of 3
#> ..$ fn : num 36.6
#> ..$ coefficient: num [1:30, 1] 24.82 -9.63 -17.4 44.79 3.7 ...
#> ..$ convergence: int 0For the sktweedie, only the Gaussian radial basis function (RBF)
kernel rbfdot() is supported. We recommend using the same set of tuned
parameters as if a ktweedie model is fitted and tuning lam2
manually:
sktd.cv2d <- ktd_cv2d(x = x,
y = y,
kernfunc = rbfdot,
lambda = c(1e-3, 1e0),
sigma = c(1e-3, 1e0),
nfolds = 5,
ncoefs = 10,
loss = "LL")
sktd.fit <- ktd_estimate(x = x,
y = y,
kern = rbfdot(sigma = sktd.cv2d$Best_sigma),
lam1 = sktd.cv2d$Best_lambda,
sparsity = TRUE,
lam2 = 1,
ftol = 1e-3,
partol = 1e-3,
innerpartol = 1e-5)The function ktd_predict() can identify necessary information stored
in ktd.fit$data and sktd.fit$data to make predictions at the
user-specified newdata. If the argument newdata is unspecified, the
prediction will be made at the original x used in model training and
fitting.
ktd.pred <- ktd_predict(ktd.fit, type = "response")
head(ktd.pred$prediction)
#> [,1]
#> [1,] 6.448220e+02
#> [2,] 1.750695e-03
#> [3,] 9.215399e-02
#> [4,] 4.713962e+00
#> [5,] 1.678452e-01
#> [6,] 1.650646e+00If newdata with the same dimension as x is provided, the prediction
will be made at the new data points.
# Use a subset of the original x as newdata.
newdata <- x[1:6, ]
ktd.pred.new <- ktd_predict(ktd.fit,
newdata = newdata,
type = "response")
sktd.pred.new <- ktd_predict(sktd.fit,
newdata = newdata,
type = "response")
data.frame(ktweedie = ktd.pred.new$prediction,
sktweedie = sktd.pred.new$prediction)
#> ktweedie sktweedie
#> 1 6.448220e+02 421.931421
#> 2 1.750695e-03 22.543092
#> 3 9.215399e-02 23.415272
#> 4 4.713962e+00 1.642355
#> 5 1.678452e-01 12.034229
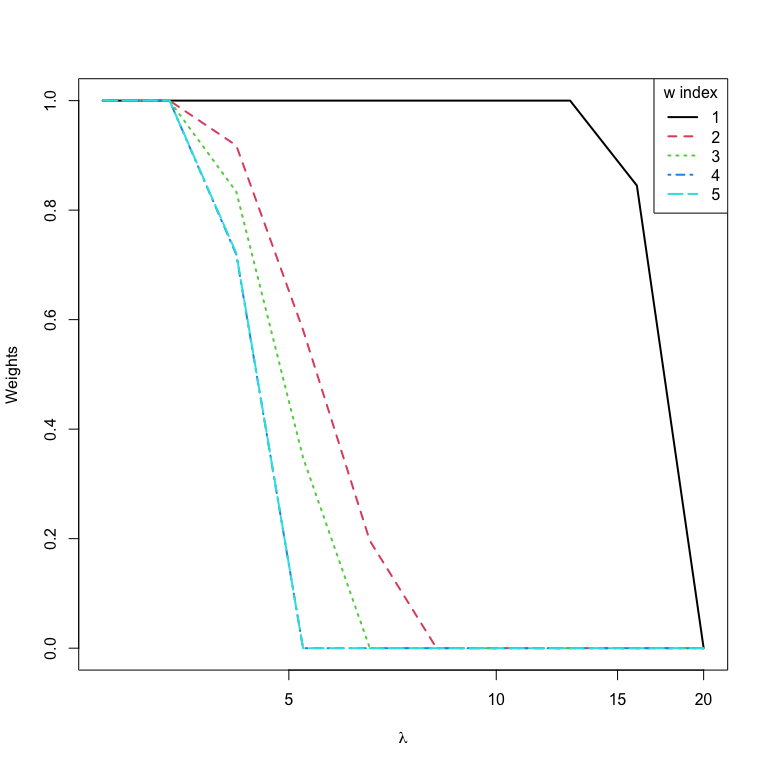
#> 6 1.650646e+00 122.187222In practice, the variable selection results of the sktweedie is more
meaningful. An effective way to fit the sktweedie is to start with an
arbitrarily big lam2 that sets all weights to zero and gradually
decrease its value. Note that a warning message is generated for the
first lam2, suggesting that all weights are set to zero.
nlam2 <- 10
lam2.seq <- 20 * 0.8^(1:nlam2 - 1)
wts <- matrix(NA, nrow = nlam2, ncol = ncol(x))
for (i in 1:nlam2) {
sktd.tmp <- ktd_estimate(x = x,
y = y,
kern = rbfdot(sigma = sktd.cv2d$Best_sigma),
lam1 = sktd.cv2d$Best_lambda,
sparsity = TRUE,
lam2 = lam2.seq[i],
ftol = 1e-3,
partol = 1e-3,
innerpartol = 1e-5)
wt.tmp <- sktd.tmp$estimates[[1]]$weight
if (is.null(wt.tmp)) wts[i, ] <- 0 else wts[i, ] <- wt.tmp
}
#> WARNING: All weights are zero in weight update iteration:
#> [1] 2
# plot the solution path with graphics::matplot()
matplot(y = wts,
x = lam2.seq,
type = "l",
log = "x",
ylab = "Weights",
xlab = expression(paste(lambda)),
lwd = 2)
legend("topright",
title = "w index",
legend = 1:5,
lty = 1:5,
col = 1:6,
lwd = 2)