System identification for ControlSystems.jl. Examples in the form of jupyter notebooks are provided here.
There exist two methods for identification of statespace models, n4sid and pem. n4sid uses subspace-based identification whereas pem solves the prediction-error problem using an iterative optimization method (from Optim.jl). If unsure which method to use, try n4sid first.
d = iddata(y,u,sampletime)
sys = n4sid(d, :auto; verbose=false)
# or use a robust version of svd if y has outliers or missing values
using TotalLeastSquares
sys = n4sid(d, :auto; verbose=false, svd=x->rpca(x)[3])Estimate a statespace model using the n4sid method. Returns an object of type N4SIDResult where the model is accessed as sys.sys.
d: Identification data object, created usingiddata(y,u,sampletime).y: Measurements N×nyu: Control signal N×nur: Rank of the model (model order)verbose: Print stuff?Wf: A frequency-domain model of measurement disturbances. To focus the attention of the model on a narrow frequency band, try something likeWf = Bandstop(lower, upper, fs=1/Ts)to indicate that there are disturbances outside this band.i: Algorithm parameter, generally no need to tune thisγ: Set this to a value between (0,1) to stabilize unstable models such that the largest eigenvalue has magnitude γ.
The frequency weighting is borrowing ideas from "Frequency Weighted Subspace Based System Identification in the Frequency Domain", Tomas McKelvey 1996. In particular, we apply the output frequency weight matrix (Fy) as it appears in eqs. (16)-(18).
See
sys = era(d::AbstractIdData, r, m = 2r, n = 2r, l = 5r)
H = okid(d::AbstractIdData, r, l = 5r)Models can be simulated using lsim from ControlSystems.jl and using simulate. You may also convert the model to a KalmanFilter from LowLevelParticleFilters.jl by calling KalmanFilter(sys), after which you can perform filtering and smoothing etc. with the utilities provided for a KalmanFilter.
A simple algorithm for identification of discrete-time LTI systems on state-space form:
is provided. The user can choose to minimize either prediction errors or simulation errors, with arbitrary metrics, i.e., not limited to squared errors.
The result of the identification is a custom type StateSpaceNoise <: ControlSystems.LTISystem, with fields A,B,K, representing the dynamics matrix, input matrix and Kalman gain matrix, respectively. The observation matrix C is not stored, as this is always given by [I 0] (you can still access it through sys.C thanks to getproperty).
This package also supports estimating models on the form
through pseudo-linear regression or
through the generalized least squares method (arxar).
Estimation of the more general model form
or any of its other special cases is not supported. Since those models are also LTI systems, estimating a state-space model is in some sense equivalent.
Transfer-function estimation through spectral methods is supported through the functions tfest and coherence.
Below, we generate a system and simulate it forward in time. We then try to estimate a model based on the input and output sequences.
using ControlSystemIdentification, ControlSystems, Random, LinearAlgebra
function ⟂(x)
u,s,v = svd(x)
u*v
end
function generate_system(nx,ny,nu)
U,S = ⟂(randn(nx,nx)), diagm(0=>0.2 .+ 0.5rand(nx))
A = S*U
B = randn(nx,nu)
C = randn(ny,nx)
sys = ss(A,B,C,0,1)
end
Random.seed!(1)
T = 1000 # Number of time steps
nx = 3 # Number of poles in the true system
nu = 1 # Number of control inputs
ny = 1 # Number of outputs
x0 = randn(nx) # Initial state
sim(sys,u,x0=x0) = lsim(sys, u', 1:T, x0=x0)[1]' # Helper function
sys = generate_system(nx,nu,ny)
u = randn(nu,T) # Generate random input
y = sim(sys, u, x0) # Simulate system
d = iddata(y,u,1)
sysh,x0h,opt = pem(d, nx=nx, focus=:prediction) # Estimate model
yh = predict(sysh, d, x0h) # Predict using estimated model
plot([y; yh]', lab=["y" "ŷ"]) # Plot prediction and true outputWe can also simulate the system with colored noise, necessitating estimating also noise models.
σu = 0.1 # Noise variances
σy = 0.1
sysn = generate_system(nx,nu,ny) # Noise system
un = u + sim(sysn, σu*randn(size(u)),0*x0) # Input + load disturbance
y = sim(sys, un, x0)
yn = y + sim(sysn, σy*randn(size(u)),0*x0) # Output + measurement noise
dn = iddata(yn,un,1)The system now has 3nx poles, nx for the system dynamics, and nx for each noise model, we indicated this to the main estimation function pem:
sysh,x0h,opt = pem(dn,nx=3nx, focus=:prediction)
yh = predict(sysh, dn, x0h) # Form prediction
plot([y; yh]', lab=["y" "ŷ"]) # Compare true output (without noise) to predictionWe can have a look at the singular values of a balanced system Gramian:
s = ss(sysh) # Convert to standard state-space type
s2,G = balreal(s) # Form balanced representation (obs. and ctrb. Gramians are the same
diag(G) # Singular values of Gramians
# 9-element Array{Float64,1}:
# 3.5972307807882844
# 0.19777167699663994
# 0.0622528285731599
# 0.004322765397504325
# 0.004270259700592557
# 0.003243449461350837
# 0.003150873301312319
# 0.0005827927965893053
# 0.00029732262107216666Note that there are 3 big singular values, corresponding to the system poles, there are also 2×3 smaller singular values, corresponding to the noise dynamics.
The estimated noise model can be extracted by noise_model(sys), we can visualize it with a bodeplot.
bodeplot(noise_model(sysh), exp10.(range(-3, stop=0, length=200)), title="Estimated noise dynamics")See the example notebooks for these plots.
sys, x0, opt = pem(d; nx, kwargs...)
d: Identification data object, created usingiddata(y, u sampletime)y: Measurements, either a matrix with time along dim 2, or a vector of vectorsu: Control signals, same structure asy
nx: Number of poles in the estimated system. This number should be chosen as number of system poles plus number of poles in noise models for measurement noise and load disturbances.focus: Either:predictionor:simulation. If:simulationis chosen, a two stage problem is solved with prediction focus first, followed by a refinement for simulation focus.metric: A Function determining how the size of the residuals is measured, defaultsse(e'e), but any Function such asnorm,e->sum(abs,e)ore -> e'Q*ecould be used.regularizer(p) = 0: function for regularization of the parameter vectorp. The structure ofpis detailed below. L₂ regularization, for instance, can be achieved byregularizer = p->sum(abs2, p)solverDefaults toOptim.BFGS()kwargs: additional keyword arguments are sent toOptim.Options.
The parameter vector is of type ComponentVector and the fields A,B,K,x0 can be accessed as p.A etc. The internal storage is according to
A = size(nx,nx)
B = size(nx,nu)
K = size(nx,ny)
x0 = size(nx)
p = [A[:];B[:];K[:];x0]sys::StateSpaceNoise: identified system. Can be converted toStateSpacebyconvert(StateSpace, sys)orss(sys), but this will discard the Kalman gain matrix, seeinnovation_form.x0: Estimated initial stateopt: Optimization problem structure. Contains info of the result of the optimization problem
pem: Main estimation function, see above.predict(sys, d, x0=zeros): Form predictions using estimatedsys, this essentially runs a stationary Kalman filter.simulate(sys, u, x0=zeros): Simulate the system using inputu. The noise model and Kalman gain does not have any influence on the simulated output.innovation_form: Extract the noise model from the estimated system (ss(A,K,C,0)).
Internally, Optim.jl is used to optimize the system parameters, using automatic differentiation to calculate gradients (and Hessians where applicable). Optim solver options can be controlled by passing keyword arguments to pem, and by passing a manually constructed solver object. The default solver is BFGS()
Basic support for ARX/ARMAX model estimation, i.e. a model on any of the forms
is provided. The ARX estimation problem is convex and the solution is available on closed-form. Usage example:
N = 2000 # Number of time steps
t = 1:N
Δt = 1 # Sample time
u = randn(N) # A random control input
G = tf(0.8, [1,-0.9], 1)
y = lsim(G,u,t)[1][:]
yn = y
d = iddata(y,u,Δt)
na,nb = 1,1 # Number of polynomial coefficients
Gls = arx(d,na,nb,stochastic=false) # set stochastic to true to get a transfer function of MonteCarloMeasurements.Particles
@show Gls
# TransferFunction{ControlSystems.SisoRational{Float64}}
# 0.8000000000000005
# --------------------------
# 1.0*z - 0.8999999999999997As we can see, the model is perfectly recovered. In reality, the measurement signal is often affected by noise, in which case the estimation will suffer. To combat this, a few different options exist:
e = randn(N)
yn = y + e # Measurement signal with noise
d = iddata(yn,u,Δt)
na,nb,nc = 1,1,1
Gls = arx(d,na,nb, stochastic=true) # Regular least-squares estimation
Gtls = arx(d,na,nb, estimator=tls) # Total least-squares estimation
Gwtls = arx(d,na,nb, estimator=wtls_estimator(y,na,nb)) # Weighted Total least-squares estimation
Gplr, Gn = plr(d,na,nb,nc, initial_order=20) # Pseudo-linear regression
@show Gls; @show Gtls; @show Gwtls; @show Gplr; @show Gn;
# TransferFunction{ControlSystems.SisoRational{MonteCarloMeasurements.Particles{Float64,500}}}
# 0.824 ± 0.029
# ---------------------
# 1.0*z - 0.713 ± 0.013
# Gtls = TransferFunction{ControlSystems.SisoRational{Float64}}
# 1.848908051191616
# -------------------------
# 1.0*z - 0.774385918070221
# Gwtls = TransferFunction{ControlSystems.SisoRational{Float64}}
# 0.8180228878106678
# -------------------------
# 1.0*z - 0.891939152690534
# Gplr = TransferFunction{ControlSystems.SisoRational{Float64}}
# 0.8221837077656046
# --------------------------
# 1.0*z - 0.8896345125395438
# Gn = TransferFunction{ControlSystems.SisoRational{Float64}}
# 0.9347035105826179
# --------------------------
# 1.0*z - 0.8896345125395438We now see that the estimate using standard least-squares is heavily biased and it is wrongly certain about the estimate (notice the ± in the transfer function coefficients). Regular Total least-squares does not work well in this example, since not all variables in the regressor contain equally much noise. Weighted total least-squares does a reasonable job at recovering the true model. Pseudo-linear regression also fares okay, while simultaneously estimating a noise model. The helper function wtls_estimator(y,na,nb) returns a function that performs wtls using appropriately sized covariance matrices, based on the length of y and the model orders. Weighted total least-squares estimation is provided by TotalLeastSquares.jl. See the example notebooks for more details.
Uncertain transfer function with Particles coefficients can be used like any other model. Try, e.g., nyquistplot(Gls) to get a Nyquist plot with confidence bounds.
See also function arma for estimation of signal models without inputs.
arx: Transfer-function estimation using closed-form solution.ar: Estimate an AR model.arma: Estimate an ARMA model.plr: Transfer-function estimation using pseudo-linear regressionarxar: Transfer-function estimation using generalized least squares methodgetARXregressor/getARregressor: For low-level control over the estimation See docstrings for further help.
The model estimation procedures can be used to estimate spectrograms. This package extends some methods from DSP.jl to accept a estimation function as the second argument. To create a suitable such function, we provide the function model_spectrum. Usage is illustrated below.
using ControlSystemIdentification, DSP
T = 1000
fs = 1
s = sin.((1:1/fs:T) .* 2pi/10) + 0.5randn(T)
S1 = spectrogram(s,window=hanning, fs=fs) # Standard spectrogram
estimator = model_spectrum(ar,1/fs,6)
S2 = spectrogram(s,estimator,window=rect, fs=fs) # Model-based spectrogram
plot(plot(S1,title="Standard Spectrogram"),plot(S2,title="AR Spectrogram")) # Requires the package LPVSpectral.jlNon-parametric estimation is provided through spectral estimation. To illustrate, we once again simulate some data:
T = 100000
h = 1
sim(sys,u) = lsim(sys, u, 1:T)[1][:]
σy = 0.5
sys = tf(1,[1,2*0.1,0.1])
ωn = sqrt(0.3)
sysn = tf(σy*ωn,[1,2*0.1*ωn,ωn^2])
u = randn(T)
y = sim(sys, u)
yn = y + sim(sysn, randn(size(u)))
d = iddata(y,u,h)
dn = iddata(yn,u,h)We can now estimate the coherence function to get a feel for whether or nor our data seems to be generated by a linear system:
k = coherence(d) # Should be close to 1 if the system is linear and noise free
k = coherence(dn) # Slightly lower values are obtained if the system is subject to measurement noiseWe can also estimate a transfer function using spectral techniques, the main entry point to this is the function tfest, which returns a transfer-function estimate and an estimate of the power-spectral density of the noise (note, the unit of the PSD is squared compared to a transfer function, hence the √N when plotting it in the code below):
G,N = tfest(dn)
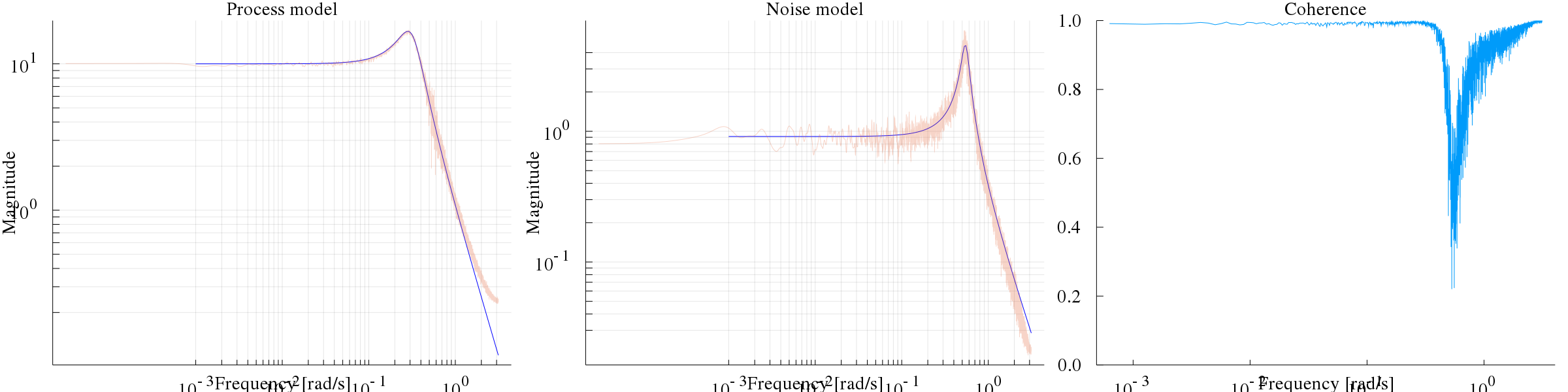
bodeplot([sys,sysn], exp10.(range(-3, stop=log10(pi), length=200)), layout=(1,3), plotphase=false, subplot=[1,2,2], size=(3*800, 600), ylims=(0.1,300), linecolor=:blue)
coherenceplot!(dn, subplot=3)
plot!(G, subplot=1, lab="G Est", alpha=0.3, title="Process model")
plot!(√N, subplot=2, lab="N Est", alpha=0.3, title="Noise model")The left figure displays the Bode magnitude of the true system, together with the estimate (noisy), and the middle figure illustrates the estimated noise model. The right figure displays the coherence function, which is close to 1 everywhere except for at the resonance peak of the noise log10(sqrt(0.3)) = -0.26.
See the example notebooks for more details.
To estimate a parametric, rational transfer function from frequency-domain data, call tfest with an FRD object and an initial guess for the system model. This initial guess determines the number of coefficients in the numerator and denominator of the estimated model.
G0 = tf(1.0, [1,1,1]) # Initial guess
G = tfest(d::FRD, G0)Internally, Optim is using a gradient-based optimizer to find the optimal fit of the bode curve of the system. The defaut optimizer BFGS can be changed, see the docstring ?tfest.
For a comparison between estimation in the time and frequency domains, see this notebook.
If the above problem is hard to solve, you may parametrize the model using, e.g., a Laguerre basis expansion, example:
basis = laguerre_oo(1, 50) # Use 50 basis functions, the final model order may be reduced with baltrunc
Gest,p = tfest(d::FRD, basis)The functions impulseest(h,y,u,order) and impulseestplot performs impulse-response estimation by fitting a high-order FIR model.
Example
T = 200
h = 1
t = h:h:T
sim(sys,u) = lsim(sys, u, t)[1][:]
sys = c2d(tf(1,[1,2*0.1,0.1]),h)
u = randn(length(t))
y = sim(sys, u)
d = iddata(y,u,h)
impulseestplot(d,50, lab="Estimate")
impulseplot!(sys,50, lab="True system")See the example notebooks for more details.
A number of functions are made available to assist in validation of the estimated models. We illustrate by an example
Generate some test data:
Random.seed!(1)
T = 200
nx = 2
nu = 1
ny = 1
x0 = randn(nx)
σy = 0.5
sim(sys,u) = lsim(sys, u', 1:T)[1]'
sys = tf(1,[1,2*0.1,0.1])
sysn = tf(σy,[1,2*0.1,0.3])
# Training data
u = randn(nu,T)
y = sim(sys, u)
yn = y + sim(sysn, randn(size(u)))
dn = iddata(yn,u,1)
# Validation data
uv = randn(nu,T)
yv = sim(sys, uv)
ynv = yv + sim(sysn, randn(size(uv)))
dv = iddata(yv,uv,1)
dnv = iddata(ynv,uv,1)We then fit a couple of models, the flag difficult=true causes pem to solve an initial global optimization problem with constraints on the stability of A-KC to provide a good guess for the gradient-based solver
res = [pem(dn,nx=nx, iterations=1000, difficult=true, focus=:prediction) for nx = [1,3,4]]After fitting the models, we validate the results using the validation data and the functions simplot and predplot (cf. Matlab sys.id's compare):
ω = exp10.(range(-2, stop=log10(pi), length=150))
fig = plot(layout=4, size=(1000,600))
for i in eachindex(res)
(sysh,x0h,opt) = res[i]
simplot!( sysh,dnv,x0h; subplot=1, ploty=i==1)
predplot!(sysh,dnv,x0h; subplot=2, ploty=i==1)
end
bodeplot!(ss.(getindex.(res,1)), ω, plotphase=false, subplot=3, title="Process", linewidth=2*[4 3 2 1])
bodeplot!(innovation_form.(getindex.(res,1)), ω, plotphase=false, subplot=4, linewidth=2*[4 3 2 1])
bodeplot!(sys, ω, plotphase=false, subplot=3, lab="True", linecolor=:blue, l=:dash, legend = :bottomleft, title="System model")
bodeplot!(innovation_form(ss(sys),syse=ss(sysn)), ω, plotphase=false, subplot=4, lab="True", linecolor=:blue, l=:dash, ylims=(0.1, 100), legend = :bottomleft, title="Noise model")
display(fig)In the figure, simulation output is compared to the true model on the top left and prediction on top right. The system models and noise models are visualized in the bottom plots. Both high-order models capture the system dynamics well, but struggle slightly with capturing the gain of the noise dynamics. The figure also indicates that a model with 4 poles performs best on both prediction and simulation data. The true system has 4 poles (two in the process and two in the noise process) so this is expected. However, the third order model performs almost equally well and may be a better choice.
- For estimation of linear time-varying models (LTV), see LTVModels.jl.
- For estimation of linear and nonlinear grey-box models in continuous time, see DifferentialEquations.jl (parameter estimation)
- Estimation of nonlinear black-box models in continuous time DiffEqFlux.jl and in discrete time Flux.jl
- For more advanced spectral estimation, cross coherence, etc., see LPVSpectral.jl
- This package interacts well with MonteCarloMeasurements.jl. See example file.