- Input: a OBO Graph JSON object
- Optional: a JSON ontology stylesheet
- Output: a Dot-format / Graphviz file
Quickstart (command line):
./bin/og2dot.js tests/simple-og.json > test.dot
dot test.dot -Tpng -Grankdir=BT > test.png
Command line; from python obographs package
ogr -p subClassOf BFO:0000050 -r obo:go -t png g nucleus
API:
var compoundRelations = ['BFO:0000050']
var styleMap = {}
var gv = new ogv.OboGraphViz(result.data)
var dot = gv.renderDot(compoundRelations, styleMap)
One or more predicates can be designated as 'compound', i.e. used for nesting.
On the command line, use -c. In the API, use compoundRelations
Example:
./bin/og2dot.js -c is_a tests/simple-og.json > test.dot
Generates:
Note only works for subgraphs that exhibit disjointness over this property, acyclicity
In the API can be passed using styleMap. On the command line, by using either -s (to pass a JSON file) or -S (to pass stringified JSON object directly on command line)
E.g.
./bin/og2dot.js -s examples/example-style.json -c is_a tests/simple-og.json > test.dot
These go in the root of the stylemap object
{
"style": "filled",
"fillcolor": "green"
}
this sets all nodes to be filled green
Each relationship type can have its own individual style, by passing relationProperties. This is keyed by the CURIE for the relation (or "is_a" for subClassOf):
{
"relationProperties": {
"is_a": {
"color": "black",
"penwith": 3,
"arrowhead": "open",
"label": ""
},
"BFO:0000050": {
"arrowhead": "tee",
"color": "blue"
}
}
}
Pass in prefixProperties to be able to assign individual properties for ontology prefixes. This can be useful when visualization graphs that combine multiple ontologies
{
"prefixProperties": {
"SO": {
"fillcolor": "yellow"
},
"RO": {
"fillcolor": "pink"
},
"BFO": {
"fillcolor": "cyan"
}
}
}
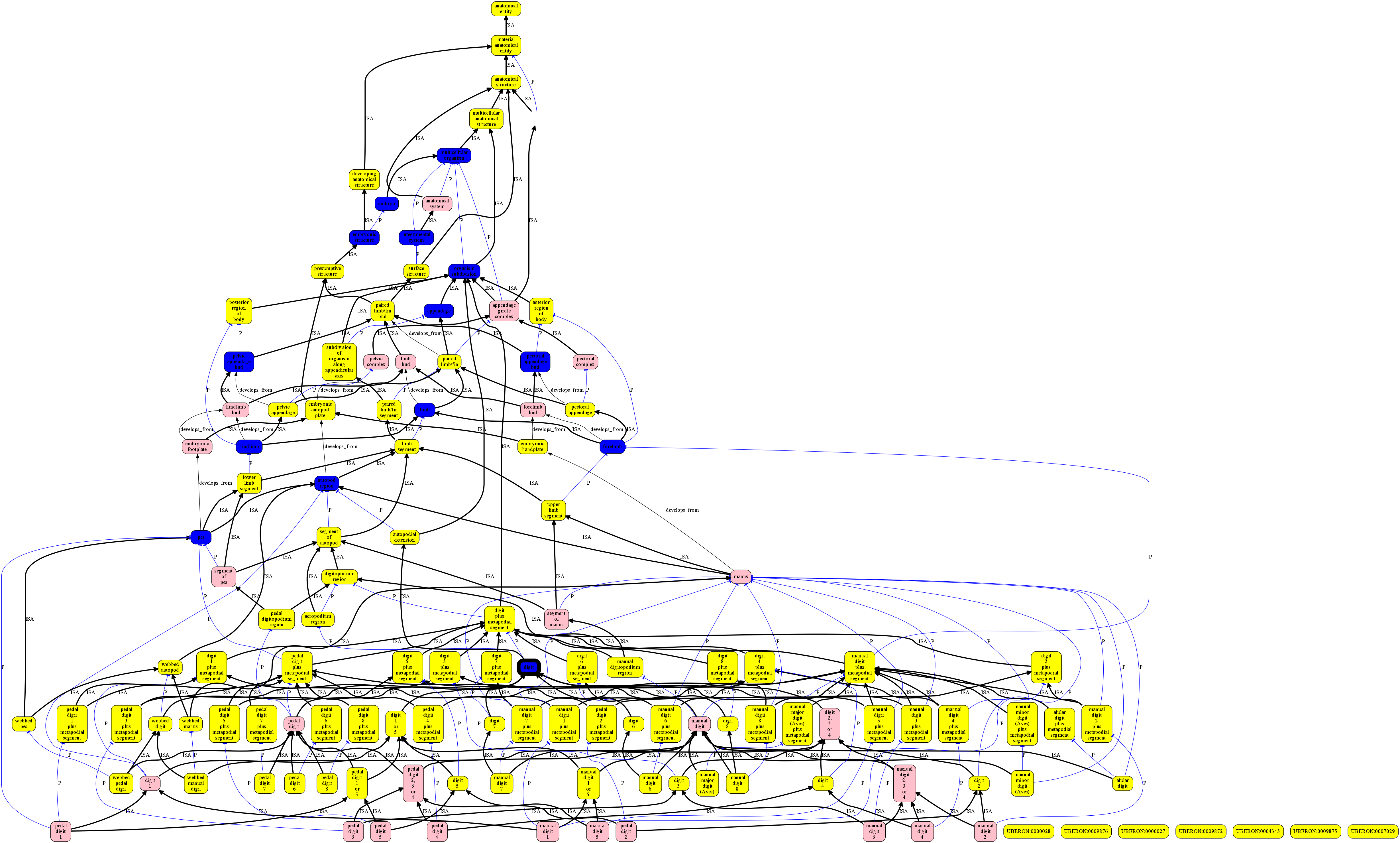
The following example uses all subclasses of digit in Uberon, plus their ancestors, which forms a complex lattic structure.
See [digit.json](examples/digit.json] for the underlying ontology. See examples/uberon-style.json for the stylesheet.
og2dot.js -s examples/uberon-style.json examples/digit.json -t png -o examples/digit.png
Renders:
E.g. GO-CAM models
{
nodeFilter : {
"type" : "INDIVIDUAL"
},
labelFrom : "type"
}
See:
https://github.com/biolink/biolink-api/tree/master/obographs
(note: this python API may move to its own repo in future)
obographs-python command line:
ogr -p subClassOf BFO:0000050 -r go -t png a nucleus
This proceeds by:
- Using the python obographs library to extract a networkx subgraph around the specified node
- Write as obographs-json
- Calls og2dot.js
Go to http://api.monarchinitiative.org/api/
See the /ontol/subgraph/ route
This exports obographs which can be fed in to this js lib
TODO - link to demo site
AmiGO uses bbop-graphs; these are similar enough that they can be passed in instead of obographs.
Why not D3, cytoscape js etc?
These are all very nice and pretty, but GraphViz has some powerful features that I have not found in any other framework (or have been too lazy to find out how to do). In particular:
- Easy to run on command line
- The ability to nest relationships (update: compound graphs in cytoscape.js)
- simple control over box and edge visual attributes
- embedding arbitrary HTML
This is intended to replace blipkit graphviz generation. For some examples, see mondo report