This short package provides fast estimation of the estimators of agreement for multiple observers. The data are assumed to include a matrix or data-frame with subjects as the rows and observers as the columns, with nominal or ordinal categories as the data. The methods are described by O’Connell and Dobson (Biometrics 1984; 40: 973-983) and Schouten (Statistica Neerlandica 1982: 36: 45-61).
There are several estimators of particular interest here:
- O’Connell-Dobson-Schouten estimator of average agreement across the observers.
- Average agreement by subject (O’Connell and Dobson 1984)
- Average agreement by observer (Schouten 1982)
P-values compared with the null hypothesis (that is, H0: average agreement is zero) are available for these estimators. For the average agreement by observer, p-values are also available compared with the average agreement across observers.
The summary methods describe a range of estimators, including confidence intervals. Several plots are available.
The Fortran code from O’Connell and Dobson was provided by Professor O’Connell. The algorithms described by Schouten (1982) were implemented in Fortran 77.
Both O’Connell and Dobson (1984) and Schouten (1982) use the data from Landis and Koch (Biometrics 1977; 33: 363-374). The data are included as the landis dataset in the R package.
A simple description of the average agreement across the observers is given by printing the magree object:
require(magree)
magree(landis)#+RESULTS[3f3328a44897fea629d021aa38d85d7da20e960f]:
O'Connell-Dobson-Schouten estimator (unweighted) Sav(hetero): 0.361290 (se: 0.028881; 95% CI: 0.306811, 0.419587) Sav(homoge): 0.354335 (se: 0.030018; 95% CI: 0.297924, 0.415112) Pr(Overall agreement due to chance | hetero): < 2.22e-16 Pr(Overall agreement due to chance | homoge): < 2.22e-16
We can obtain considerably more information from the summary of the magree object:
summary(magree(landis))#+RESULTS[fa757baf5e4f9c8efde3a95f84b1890b0ca98e41]:
O'Connell-Dobson-Schouten estimator (unweighted)
Sav(hetero): 0.361290 (se: 0.028881; 95% CI: 0.306811, 0.419587)
Sav(homoge): 0.354335 (se: 0.030018; 95% CI: 0.297924, 0.415112)
Pr(Overall agreement due to chance | hetero): < 2.22e-16
Pr(Overall agreement due to chance | homoge): < 2.22e-16
Observed marginal distributions for categories:
1 2 3 4 5
0.28087167 0.25423729 0.36440678 0.07384988 0.02663438
Observed marginal distributions for categories by observer:
1 2 3 4 5
A 0.2203390 0.2203390 0.3220339 0.186440678 0.050847458
B 0.2288136 0.1016949 0.5847458 0.059322034 0.025423729
C 0.2627119 0.3559322 0.3135593 0.050847458 0.016949153
D 0.3220339 0.4067797 0.1949153 0.067796610 0.008474576
E 0.1355932 0.2627119 0.4491525 0.118644068 0.033898305
F 0.5254237 0.2627119 0.1694915 0.008474576 0.033898305
G 0.2711864 0.1694915 0.5169492 0.025423729 0.016949153
Agreement statistics S_i for each subject:
1 2 3 4 5 6
0.08088076 1.00000000 1.00000000 0.34348626 1.00000000 0.34348626
7 8 9 10 11 12
0.60609175 0.21218351 0.27783488 0.60609175 0.60609175 0.60609175
13 15 16 17 18 19
0.60609175 0.21218351 0.08088076 0.60609175 0.21218351 0.21218351
22 23 24 25 26 27
0.08088076 0.60609175 0.34348626 0.34348626 1.00000000 0.34348626
28 29 30 31 32 33
0.08088076 0.34348626 0.60609175 1.00000000 0.27783488 1.00000000
34 35 36 37 38 39
1.00000000 0.27783488 0.27783488 0.08088076 0.01522939 0.34348626
40 41 42 43 44 45
0.08088076 0.60609175 1.00000000 0.08088076 0.27783488 0.60609175
46 47 48 49 51 52
-0.05042199 0.21218351 0.34348626 0.08088076 0.60609175 0.27783488
53 54 55 56 57 58
0.27783488 0.01522939 0.60609175 0.60609175 0.01522939 1.00000000
59 60 61 62 63 64
1.00000000 0.60609175 0.08088076 0.27783488 0.08088076 0.21218351
65 66 67 68 69 70
0.60609175 0.08088076 1.00000000 0.34348626 0.27783488 1.00000000
71 72 73 74 76 77
0.60609175 0.27783488 0.60609175 0.01522939 0.60609175 0.34348626
78 79 80 81 82 83
0.08088076 0.34348626 -0.11607336 1.00000000 0.21218351 0.08088076
84 85 86 87 88 89
0.27783488 -0.18172474 0.60609175 0.60609175 0.01522939 -0.11607336
90 91 92 93 94 95
0.08088076 0.01522939 -0.11607336 0.21218351 0.34348626 0.27783488
96 98 99 100 101 102
-0.11607336 0.21218351 0.21218351 0.08088076 0.34348626 0.34348626
103 104 105 106 107 108
1.00000000 0.01522939 0.60609175 0.08088076 0.21218351 0.08088076
110 111 112 113 114 115
0.21218351 0.60609175 0.21218351 0.08088076 0.08088076 0.21218351
116 117 118 119 120 121
0.60609175 0.34348626 0.08088076 0.60609175 1.00000000 0.21218351
122 123 124 126
-0.11607336 -0.11607336 0.60609175 0.01522939
Agreement statistics for each observer:
Kappa [Lower, Upper] Pr(kappa_av=kappa_observer)
A 0.37274 0.30394 0.4471 0.60718
B 0.40591 0.33691 0.4788 0.03444 *
C 0.38173 0.31292 0.4556 0.40306
D 0.33866 0.27026 0.4145 0.33175
E 0.32894 0.25801 0.4086 0.23888
F 0.24269 0.17536 0.3257 < 1e-05 ***
G 0.46538 0.40385 0.5280 < 1e-05 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
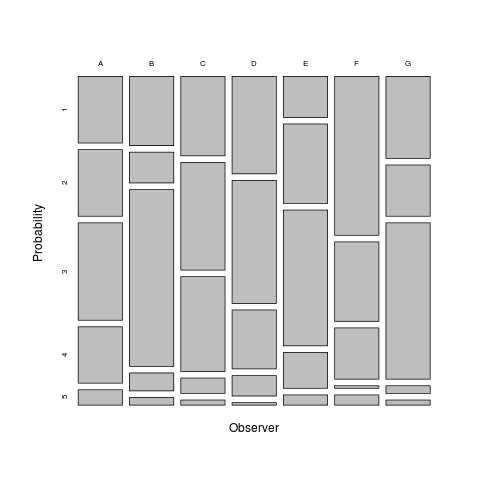
There are currently two plot types: the default type = “p1” gives a tabular plot for the marginal probabilities of each category by observed:
plot(magree(landis))#+RESULTS[ac7ce3b55597642e6c75f253cad009bda11d8338]:
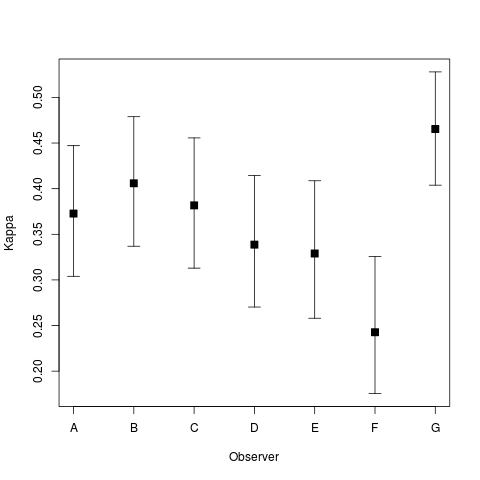
The second plot is type = “kappa by observer”, which shows the estimated average kappa by observer with intervals based on single standard errors:
plot(magree(landis),type="kappa by observer")#+RESULTS[8172ce7d6a34ee663adfe7d6e788b083b17c67f2]:
- Better describe the difference between the estimators that assume homogeneity and those that assume heterogeneity.
- For the average agreement by subject, calculate p-values compared with the average agreement across subjects (which equals the average agreement across observers). This will require an estimate of the covariance between the average agreement across subjects and the average agreement by subject. As the code is so fast, it may be possible to calculate these values using the jackknife or similar approach.
- Extend the O’Connell-Dobson estimators to include an arbitrary weight matrix.