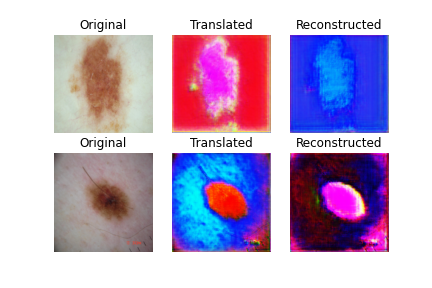
Result is after completing the epoches into one another (Kaggle Notebook)
- 1st result after 1 epoch
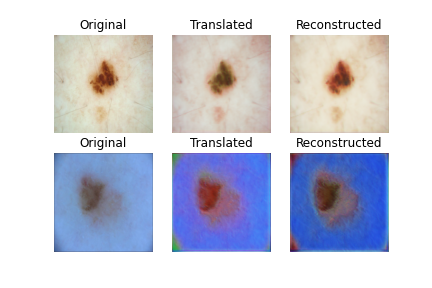
- 2nd result is after completing 15 epoch
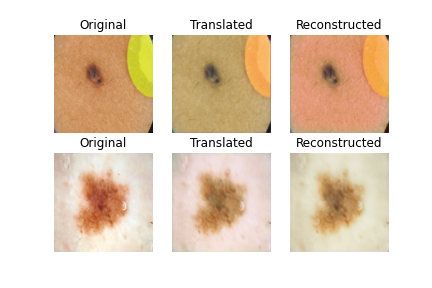
- 3rd result is completing 25 epochs
This code requires
- Python: 3.6
- Tensorflow: 2.0.0
- Keras: 2.3.1
- Download dataset from 2016, Task 3 (https://challenge.isic-archive.com/data)
- Clone this repo (obviously!)
- In this directory, make a folder in
datasetnamedisic2016and keep all files there - To build training set and test set
python data_process_isic2016.py
- To partition the dataset for training CycleGAN (two folders malignant and benign)
python data_process_gan.py
You will see that this script creates two folders trainA and trainB. Due to my utter laziness, I created testA and testB folders manually which are required for visualizing the training process of the CycleGAN. For my experiments, testA consisted of an image from trainA and vice versa.
Now that the data is partitioned according to its class label (trainA -> benign and trainB -> malignant), train CycleGAN on this data.
- Run
train_cyclegan.ipynb
This will result in two models: b2m.h5 and m2b.h5 which translate from benign -> malignant and malignant -> benign respectively. For generating the minority class (malignant) using the benign samples using the translation model:
- Run
upsampler.ipynbto oversample and balance the dataset. (Make sure you useb2m.h5if you train your model. The notebook uses the pretrained weight.)
Train the classification model using the oversampled and balanced dataset
- Train classifier using
train_ISIC_2016.ipynb
The notebook train_ISIC_2016.ipynb consists the code to evalute on the ISIC 2016 test set.