The goal of strutilities is to perform obscure string manipulations.
You can install the development version of strutilities like so:
install.packages('pak')
pak::pak('mjfrigaard/strutilities')library(strutilities)process_text() is designed to standardize the columns names and text
contents in a dataset (sort of a low-budget combination of a
janitor::clean_names() and map(df, tolower)):
names(datasets::iris)
#> [1] "Sepal.Length" "Sepal.Width" "Petal.Length" "Petal.Width" "Species"
names(process_text(datasets::iris))
#> [1] "sepal_length" "sepal_width" "petal_length" "petal_width" "species"It has an optional fct argument that will convert factors to lowercase
characters, too.
str(datasets::InsectSprays)
#> 'data.frame': 72 obs. of 2 variables:
#> $ count: num 10 7 20 14 14 12 10 23 17 20 ...
#> $ spray: Factor w/ 6 levels "A","B","C","D",..: 1 1 1 1 1 1 1 1 1 1 ...
str(process_text(datasets::InsectSprays, fct = TRUE))
#> 'data.frame': 72 obs. of 2 variables:
#> $ count: num 10 7 20 14 14 12 10 23 17 20 ...
#> $ spray: chr "a" "a" "a" "a" ...Below you’ll find the structure of the tests/ folder:
#> tests
#> ├── testthat
#> │ ├── _snaps
#> │ ├── fixtures
#> │ │ ├── make-test_data.R
#> │ │ └── test_data.rds
#> │ ├── helper.R
#> │ ├── setup.R
#> │ ├── test-pivot_term_long.R
#> │ ├── test-process_text.R
#> │ └── test-sep_cols_mult.R
#> └── testthat.R
In the test below, the process_text() function uses the source .csv
version of palmerpenguins::penguins_raw as a test fixture (loaded in
from tests/testthat/fixtures/make-test_data.R and exported to
tests/testthat/fixtures/test_data.rds)
The test helper function (test_logger()) is stored in
tests/testthat/helper.R:
describe(
"Feature: Process text from dataset
As a ...
I want to ...
So that I ...", code = {
it(
"Scenario: scenario
Given ...
When ...
Then ...", code = {
# helper
test_logger(start = "process_text()", msg = "names penguins_raw.csv")
# fixture
test_data <- readRDS(test_path("fixtures", "test_data.rds"))
# observerd data
processed_data <- process_text(raw_data = test_data, fct = TRUE)
# expected names
nms <- c("studyname",
"sample_number",
"species",
"region",
"island",
"stage",
"individual_id",
"clutch_completion",
"date_egg",
"culmen_length_mm",
"culmen_depth_mm",
"flipper_length_mm",
"body_mass_g",
"sex",
"delta_15_n_o_oo",
"delta_13_c_o_oo",
"comments")
expect_equal(object = names(processed_data), expected = nms)
test_logger(end = "process_text()", msg = "names penguins_raw.csv")
})
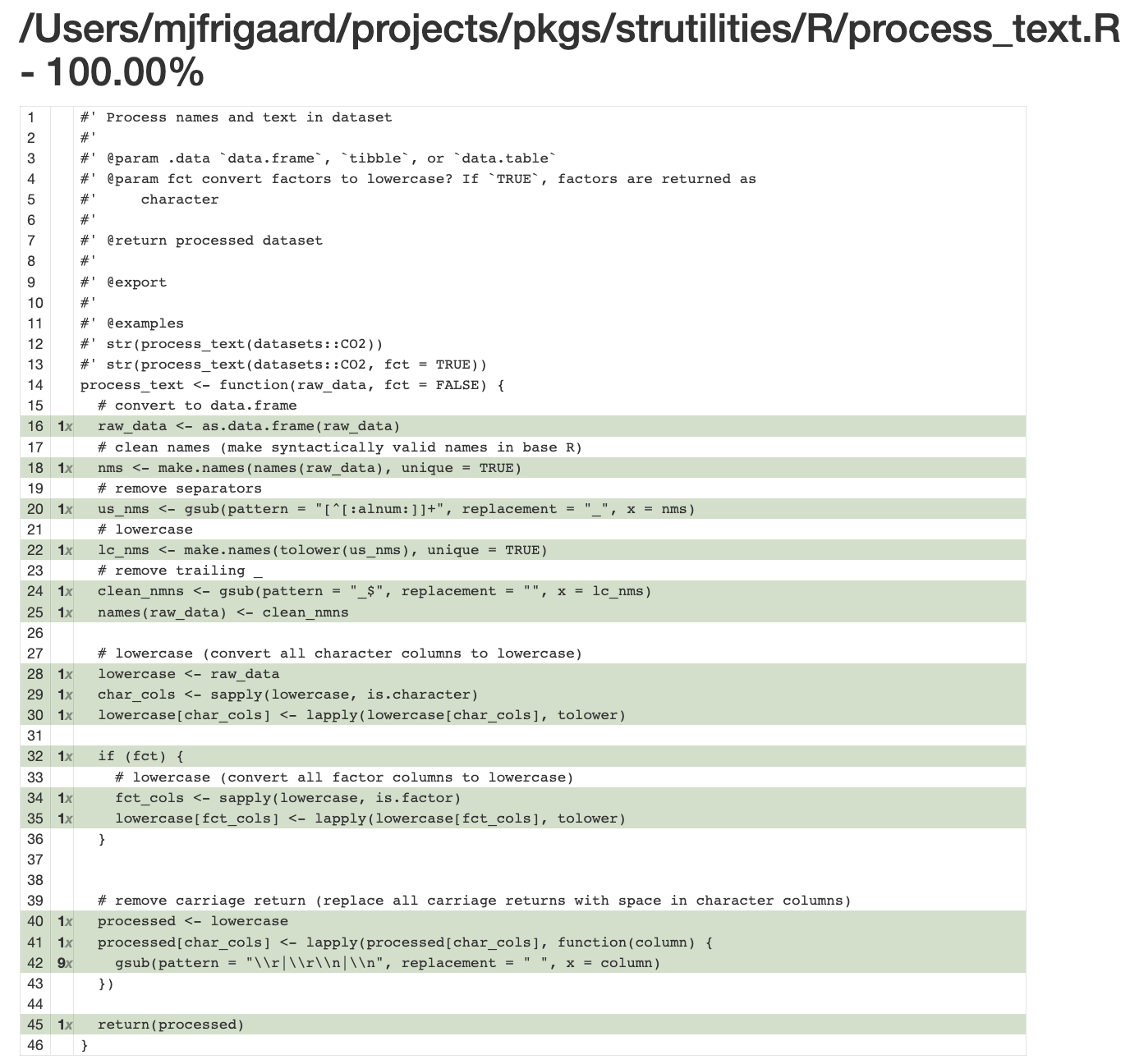
})As we can see below, the test runs fine (with the helper).
[ FAIL 0 | WARN 0 | SKIP 0 | PASS 0 ]
INFO [2023-11-09 14:59:57] [ START process_text() = names penguins_raw.csv]
[ FAIL 0 | WARN 0 | SKIP 0 | PASS 1 ]
INFO [2023-11-09 14:59:57] [ END process_text() = names penguins_raw.csv]
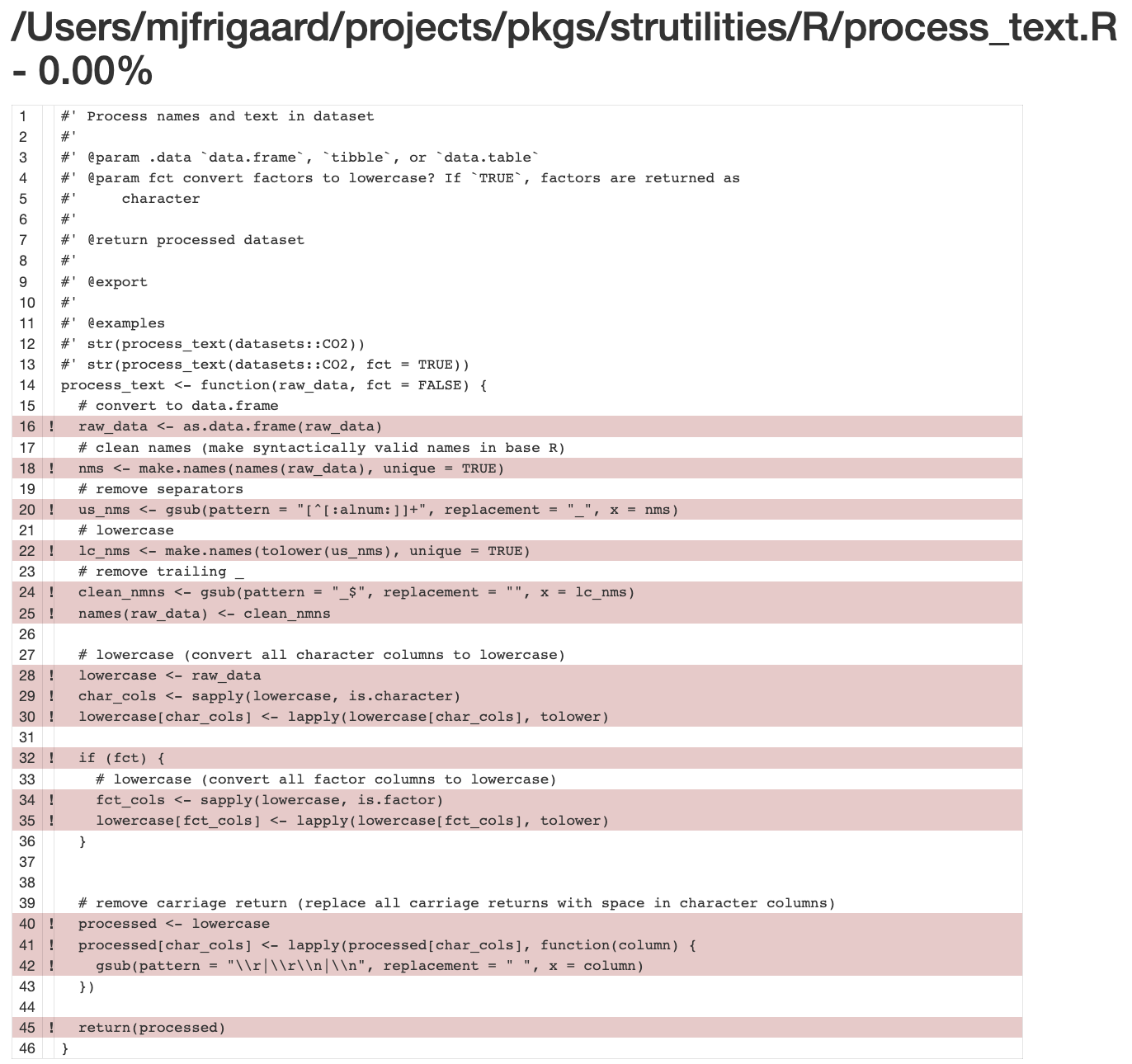
However, when I attempt to get the coverage for the test file, it shows 0.00% :(
I thought it might be it(), so I swapped it for test_that(), but
‘same same’ :(
To make sure it wasn’t the process_text() function or the helper, I
also tested loading the penguins_raw data directly from the
palmerpenguins package (i.e., not using the fixture):
describe(
"Feature: Process text from dataset
As a ...
I want to ...
So that I ...", code = {
it(
"Scenario: scenario
Given ...
When ...
Then ...", code = {
# helper
test_logger(start = "process_text()", msg = "names palmerpenguins::penguins_raw")
# data frame package
test_data <- palmerpenguins::penguins_raw
# test
processed_data <- process_text(raw_data = test_data, fct = TRUE)
nms <- c("studyname",
"sample_number",
"species",
"region",
"island",
"stage",
"individual_id",
"clutch_completion",
"date_egg",
"culmen_length_mm",
"culmen_depth_mm",
"flipper_length_mm",
"body_mass_g",
"sex",
"delta_15_n_o_oo",
"delta_13_c_o_oo",
"comments")
expect_equal(object = names(processed_data), expected = nms)
test_logger(end = "process_text()", msg = "names palmerpenguins::penguins_raw")
})
})[ FAIL 0 | WARN 0 | SKIP 0 | PASS 0 ]
INFO [2023-11-09 15:03:36] [ START process_text() = names palmerpenguins::penguins_raw]
[ FAIL 0 | WARN 0 | SKIP 0 | PASS 1 ]
INFO [2023-11-09 15:03:36] [ END process_text() = names palmerpenguins::penguins_raw]
strutilities has two other weird functions (sep_cols_mult() and
pivot_term_long()) for manipulating strings/character columns (all
written in base R to keep dependencies at a minimum).
This is an odd version of pivot_wider() that’s been adapted for a
vectors:
pivot_term_long("A large size in stockings is hard to sell.")
#> unique_items term
#> 1 A A large size in stockings is hard to sell.
#> 2 large <NA>
#> 3 size <NA>
#> 4 in <NA>
#> 5 stockings <NA>
#> 6 is <NA>
#> 7 hard <NA>
#> 8 to <NA>
#> 9 sell <NA>You can pass multiple ‘terms’ and it returns a data.frame with each unique term:
terms <- c("A large size in stockings is hard to sell.", "The first part of the plan needs changing.")
pivot_term_long(terms)
#> unique_items term
#> 1 A A large size in stockings is hard to sell.
#> 2 large <NA>
#> 3 size <NA>
#> 4 in <NA>
#> 5 stockings <NA>
#> 6 is <NA>
#> 7 hard <NA>
#> 8 to <NA>
#> 9 sell <NA>
#> 10 The The first part of the plan needs changing.
#> 11 first <NA>
#> 12 part <NA>
#> 13 of <NA>
#> 14 the <NA>
#> 15 plan <NA>
#> 16 needs <NA>
#> 17 changing <NA>The is somewhat similar to tidyr::separate(), but always uses
"[^[:alnum:]]+" as the sep and keeps all the items resulting from
the regex.
d <- data.frame(value = c(29L, 91L, 39L, 28L, 12L),
full_name = c("John", "John, Jacob",
"John, Jacob, Jingleheimer",
"Jingleheimer, Schmidt",
"JJJ, Schmidt"))
d
#> value full_name
#> 1 29 John
#> 2 91 John, Jacob
#> 3 39 John, Jacob, Jingleheimer
#> 4 28 Jingleheimer, Schmidt
#> 5 12 JJJ, Schmidt
sep_cols_mult(data = d, col = "full_name", col_prefix = "name")
#> value full_name name_1 name_2 name_3
#> 1 29 John John <NA> <NA>
#> 2 91 John, Jacob John Jacob <NA>
#> 3 39 John, Jacob, Jingleheimer John Jacob Jingleheimer
#> 4 28 Jingleheimer, Schmidt Jingleheimer Schmidt <NA>
#> 5 12 JJJ, Schmidt JJJ Schmidt <NA>