VirFinder: R package for identifying viral sequences from metagenomic data using sequence signatures
Version: 1.1
Authors: Jie Ren, Nathan Ahlgren, Yang Lu, Jed Fuhrman, Fengzhu Sun
Maintainer: Jie Ren renj@usc.edu
The package provides functions to predict viral sequences in a fasta file, such as the assembled contigs from metagenomic data. The method has good prediction accuracy for short (~1kb) and noval viral sequences.
The prediction method is based on the sequence signatures (k-tuple word frequencies) that distinguish virus from host sequences. The model was trained using equal number of known viral and host sequences. For a query sequence, the number of occurrences of k-tuple words are first counted by a c++ program using a hash table. Then the sequence is predicted based on the k-tuple word frequencies using a logistic regression model trained with previously known sequences.
R packages "glmnet", "Rcpp" and "qvalue" are needed to be installed before Installation of VirFinder.
To install "glmnet" and "Rcpp", start R and enter,
install.packages("glmnet", dependencies=TRUE)
install.packages("Rcpp", dependencies=TRUE)
To install "qvalue", start R and enter,
## try http:// if https:// URLs are not supported; it also checks for out-of-date packages
source("https://bioconductor.org/biocLite.R")
biocLite("qvalue")
To install the R package VirFinder, follow the instuctions on http://cran.r-project.org/doc/manuals/r-release/R-admin.html#Installing-packages.
To quick start, first download the package file VirFinder_1.1.tar.gz or VirFinder_1.1.zip according to your operating system.
For Mac/Linux users, if you have a Graphic User Interfaces (GUI) of R, you fire up a R graphic window and type,
install.packages("<path_to_the_file>/VirFinder_1.1.tar.gz", repos = NULL, type="source")
library(VirFinder)
If you are not using GUI of R, you can install the package from the command line. Simply type the following to the command line,
R CMD INSTALL <path_to_the_file>/VirFinder_1.1.tar.gz
For Windows users, if you have a Graphic User Interfaces (GUI) of R, you first fire up a R graphic window. You can click "Install packages(s) from local files...", and choose the file VirFinder_1.1.zip. Or you can type,
install.packages("<path_to_the_file>/VirFinder_1.1.zip", repos = NULL, type="source")
library(VirFinder)
If you are not using GUI of R, you can install the package from the command line. Simply type the following to the command line,
Rcmd INSTALL <path_to_the_file>\VirFinder_1.1.zip
Please refer to VirFinder-manual.pdf for usage instruction.
To quick start, one can predict the viral contigs using the command,
library(VirFinder)
predResult <- VF.pred(<path_to_the_fasta_file>)
As an example, the package provides a small testing data containing 30 metagenomically assembled contigs,
## (1) set the input fasta file name.
library(VirFinder)
inFaFile <- system.file("data", "contigs.fa", package="VirFinder")
## (2) prediction
predResult <- VF.pred(inFaFile)
predResult
#### (2.1) sort sequences by p-value in ascending order
predResult[order(predResult$pvalue),]
#### (2.2) estimate q-values (false discovery rates) based on p-values
predResult$qvalue <- VF.qvalue(predResult$pvalue)
#### (2.3) sort sequences by q-value in ascending order
predResult[order(predResult$qvalue),]
The package also has the reference sequence of crAssphage for users to test,
inFaFile <- system.file("data", "crAssphage.fasta", package="VirFinder")
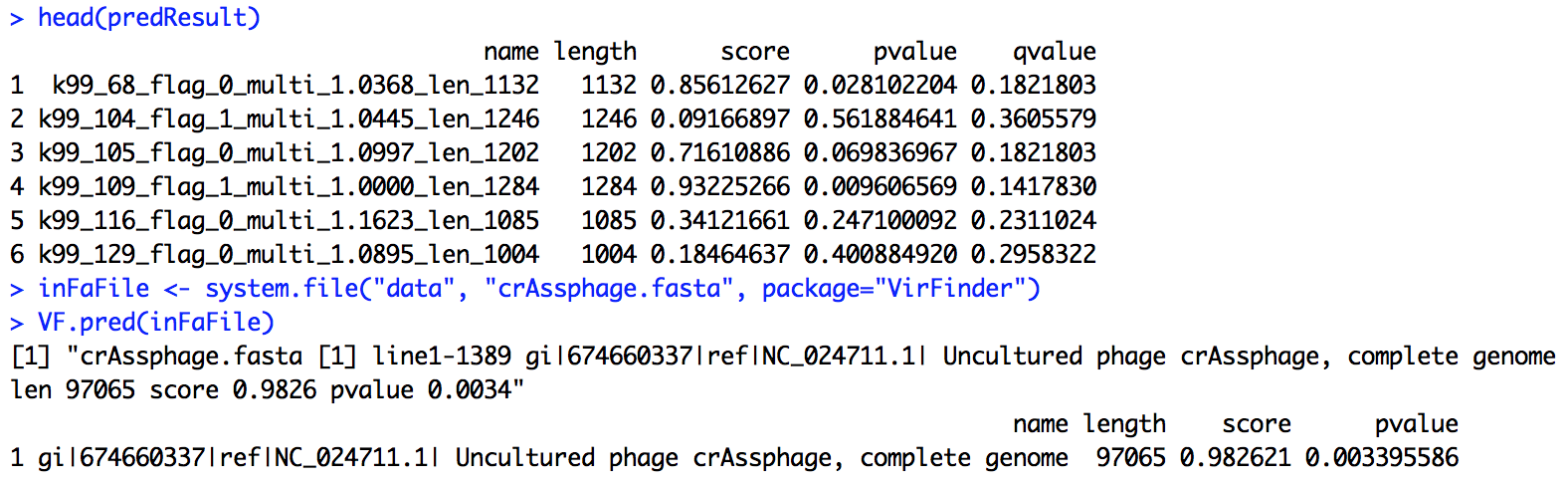
VF.pred(inFaFile)
The result will be something like the following. Each row represents a contig/sequences, with name, length, score, p-value and q-value. The higher score or lower p-value indicate higher likelihood of being a viral sequence. The q-value measures the proportion of false positives incurring when predicting viral sequences using the corresponding p-value as a threshold.
A new function has been added to the package to allow users to train the prediction model using their own database for viral sequences and host sequences.
To start with, two fasta formated files, containing viral sequences and host sequences respectively, need to be specified. The directory where the file of the trained model will be saved and the name of the model need to be set as well.
## (1) specifiy the fasta files of the training contigs
#### (1.1) one for virus and one for prokaryotic hosts
trainFaFileHost <- system.file("data", "tara_host.fa", package="VirFinder")
trainFaFileVirus <- system.file("data", "tara_virus.fa", package="VirFinder")
#### (1.2) specify the directory where the trained model will be saved, and the name of the model
userModDir <- file.path(find.package("VirFinder"))
userModName <- "modTara"
The input sequences are then split into non-overlapping fragments of fixed lengths of 0.5 kb, 1kb and 3kb. The k-tuple frequencies are counted for each fragments. The length of the k-tuple need to be specified. The longer k-tuple can describe better the difference between virus and host sequences, but if the data is not enough, it can lead to an overfitting problem. Given the database, users are suggested to test different lengths of k-tuple in order to get the best model.
Three different models are trained based on the k-tuple frequencies of viral and host fragments of three different lengths. User can specify if equal numbers of virus and host fragments are used for training by setting equalSize=TRUE. The default model is FALSE. The models are used for prediction of sequences of different lengths. For query sequences of length < 1 kb, the model trained using 0.5 kb fragments is used for predicting. For sequences of length ranging from 1 kb to 3 kb, the model trained using 1 kb fragments is used, and for sequences > 3 kb, the model trained using 3 kb fragments is used for prediction.
## (2) train the model using user's database
w <- 4 # the length of the k-tuple word
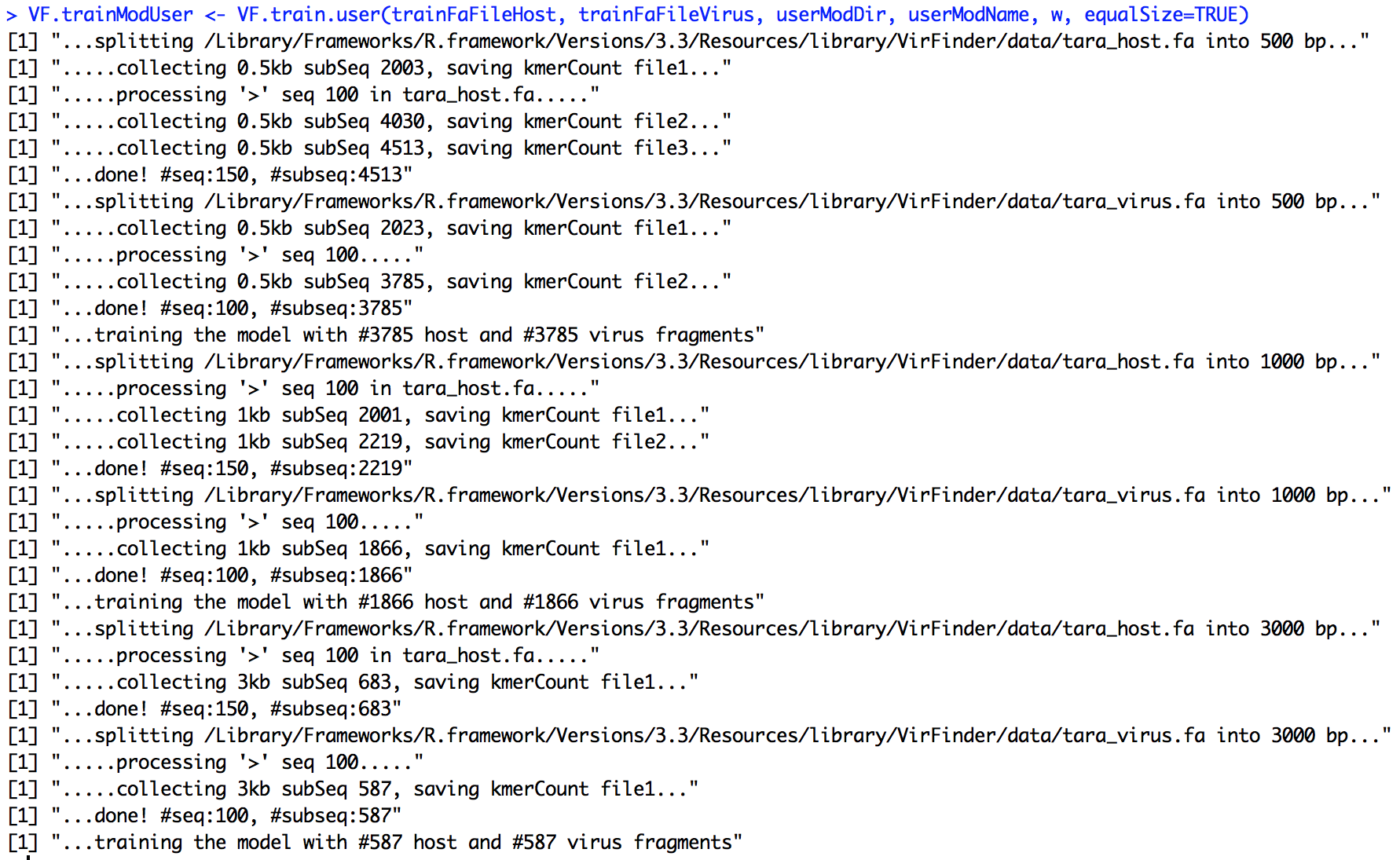
VF.trainModUser <- VF.train.user(trainFaFileHost, trainFaFileVirus, userModDir, userModName, w, equalSize=TRUE)
Collecting fragments and training the model may take some time. #Roughly, it takes 1 min to collect 100 contigs and count 8-mer frequencies. Decreasing kmer length can exponentially reduce the computing time. The screen output for the training process will be something like the following,
Once the trained model is returned, it can be used to predict viral sequences. Here we use the same example, the small testing data containing 30 contigs, for illustrate the usage.
## (3) predict the contigs using the customized model
#### (3.1) specify the fasta file containing contigs for prediction
inFaFile <- system.file("data", "contigs.fa", package="VirFinder")
#### (3.2) prediction
predResultUser <- VF.pred.user(inFaFile, VF.trainModUser)
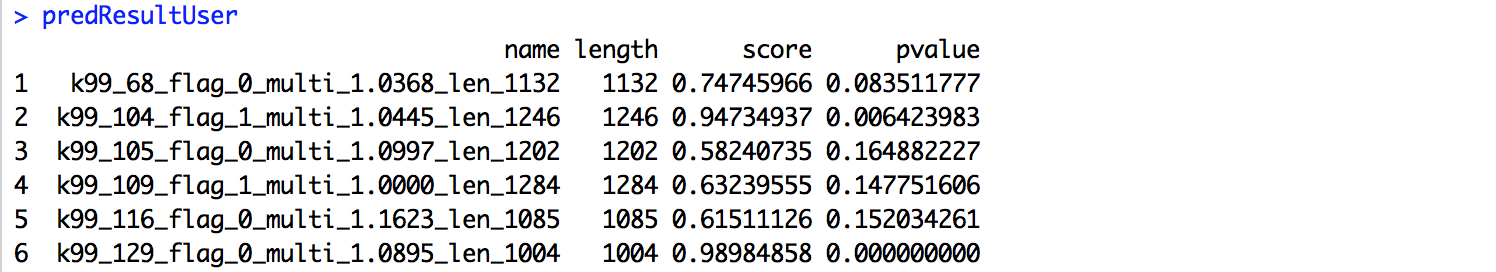
predResultUser
#### (3.3) sort sequences by p-value in ascending order
predResultUser[order(predResultUser$pvalue),]
The predict scores will be something like the following,
-
Users applying VirFinder to eukaryotic host associated microbiomes should take caution in filtering out eukaryotic sequences, as VirFinder may potentially mis-identify those sequences as viral, since eukaryotic sequences were not included in VirFinder’s training datasets.
-
VF.qvalue uses the existing function "qvalue" in the R package "qvalue" (Bass JDSwcfAJ, Dabney A and Robinson D (2015)) to estimate q-values given p-values. For some p-value distributions, VF.qvalue may prompt the error as "The estimated pi0 <= 0. Check that you have valid p-values or use a different range of lambda.". The error may also come up when the p-values are not fully covering the whole interval [0,1] or when there is a lack of p-values close or equal to 1. One possible case that leads to the above scenario is there are not enough bacteria contigs in the data such that the proportion of bacteria contigs cannot be estimated.
If you use VirFinder, please cite the following paper:
Copyright (C) 2017 University of Southern California
Authors: Jie Ren, Nathan Ahlgren, Yang Lu, Jed Fuhrman, Fengzhu Sun
This program is freely available as an R package at https://github.com/jessieren/VirFinder under the terms of the GNU General Public (version 3) as published by the Free Software Foundation (http://www.gnu.org/licenses/#GPL) for academic use.
Commercial users should contact Dr. Sun at fsun@usc.edu, copyright at the University of Southern California.
This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details.