Quantum Computing Exploration for Drug Discovery on AWS (abbrev. QCEDD), an open-sourced solution customers can launch to design and run computational studies in the area of drug discovery, e.g. molecular docking and protein folding. With this solution, customers can design and load new, or existing algorithms, into Jupyter notebooks, then orchestrate the classical and quantum compute resources using AWS Batch and AWS Step Functions. Due to its design, customers are free to scale their classical simulations by launching a performant HPC resource instead of a single EC2 instance. At the same time customer’s quantum experiments benefit from the growing number of quantum resources available through the Amazon Braket service.
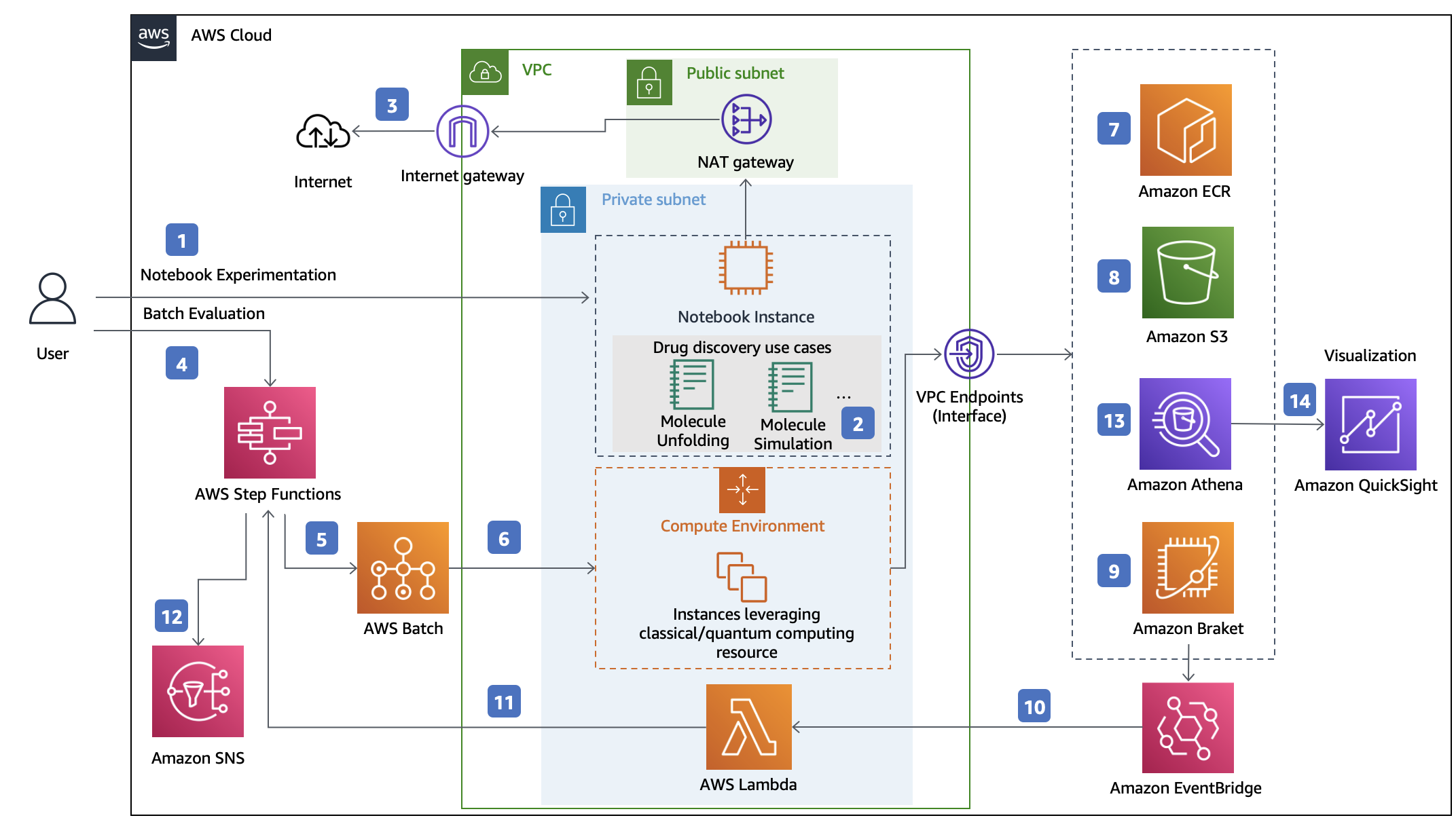
The overall architecture is shown as below:
This solution deploys the Amazon CloudFormation template in your AWS Cloud account and provides three URLs. One for Visualization. The others provide user with two approaches to study drug discovery problems: Notebook Experiment and Batch Evaluation:
- Notebook Experiment
The solution deploys the notebook for user to study different drug discovery problems. These problems will be studied using classical computing or quantum computing.
- Batch Evaluation
The solution provides user the way to evaluate a particular problem based on different computing resources , classical computing or quantum computing. For example, for the problem of molecular unfolding, the performance difference between quantum annealer and simulated annealer can be compared.
- Visualization
The solution provides user the way to visualize the comparing results of batch evaluation (e.g. performance, time)
For detailed description of architecture, please refer to the Architecture Page
| Problem Name | Methods | Function | Dataset | Reference | |
| Molecular Unfolding | QUBO | single solver | ✔ | 117 mol2 | Quantum Molecular Unfolding(2021) |
| qbsolv | |||||
| RNA Folding | QUBO | single solver | ✔ | bpRNA-1m | RNA folding using quantum computers(2022) |
| qbsolv | QHack 2022 winner | ||||
Upon successfully cloning the repository into your local development environment, you will see the following file structure in your editor:
├── CHANGELOG.md [Change log file]
├── CODE_OF_CONDUCT.md [Code of conduct file]
├── CONTRIBUTING.md [Contribution guide]
├── LICENSE [LICENSE for this solution]
├── NOTICE [Notice for 3rd-party libraries]
├── README.md [Read me file]
├── docs [Solution's document]
│ ├── en
│ ├── index.html
│ ├── mkdocs.base.yml
│ ├── mkdocs.en.yml
│ ├── mkdocs.zh.yml
│ └── zh
└── source [Solution's source and test code]
├── README.md
├── cdk.json
├── package-lock.json
├── package.json
├── src [Solution's source code]
│ ├── main.ts
│ ├── molecular-unfolding
│ │ ├── batch-images
│ │ ├── cdk
│ │ ├── lambda
│ │ ├── molecular_unfolding.ipynb
│ │ ├── molecule-data
│ │ ├── requirements.txt
│ │ └── utility
│ └── stack.ts
├── test [Solution's unit test code]
├── tsconfig.jest.json
├── tsconfig.json
└── version.json
Refer to prerequisites and deployment
The /source/run-all-tests.sh script is the centralized script for running all unit, integration, and snapshot tests for both the CDK project as well as any associated Lambda functions or other source code packages.
cd ./source
chmod +x ./run-all-tests.sh
./run-all-tests.sh