library(tidyverse)## ── Attaching packages ───────────────────────────────── tidyverse 1.3.0 ──
## ✓ ggplot2 3.3.2 ✓ purrr 0.3.4
## ✓ tibble 3.0.1 ✓ dplyr 1.0.0
## ✓ tidyr 1.1.0 ✓ stringr 1.4.0
## ✓ readr 1.3.1 ✓ forcats 0.5.0
## ── Conflicts ──────────────────────────────────── tidyverse_conflicts() ──
## x dplyr::filter() masks stats::filter()
## x dplyr::lag() masks stats::lag()
Let’s create 5000 data points from a multi-group comparison with no difference in group means
n_data <- 5000
n_per_group <- 10
n_groups <- 3
overall_mean <- 0
overal_sd <- 1set.seed(1)
sim_df <- bind_rows(lapply(seq_len(5000), function(x){
tibble(
id = x,
value = rnorm(n = n_per_group * n_groups,
mean = overall_mean,
sd = overal_sd),
group = rep(letters[seq_len(n_groups)], each = 10)
)
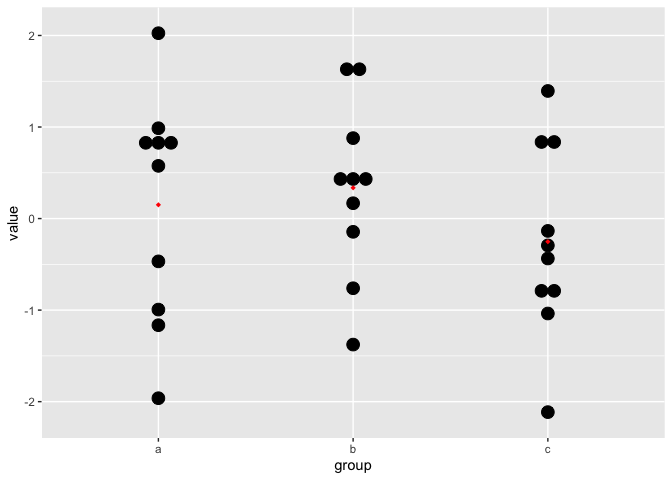
}))As an example we will inspect one of the instances:
filter(sim_df, id == 20) %>%
ggplot(aes(x = group, y = value)) +
geom_dotplot(binaxis='y', stackdir='center') +
stat_summary(fun=mean, geom="point",
shape=18, color="red")## `stat_bindot()` using `bins = 30`. Pick better value with `binwidth`.
Now we will compute, both the group mean and the overall mean for each instance (unique id):
sim_df <- sim_df %>%
group_by(id) %>%
mutate(obs_overall_mean = mean(value)) %>%
group_by(id, group) %>%
mutate(obs_group_mean = mean(value)) %>%
ungroup()
sim_df## # A tibble: 150,000 x 5
## id value group obs_overall_mean obs_group_mean
## <int> <dbl> <chr> <dbl> <dbl>
## 1 1 -0.626 a 0.0825 0.132
## 2 1 0.184 a 0.0825 0.132
## 3 1 -0.836 a 0.0825 0.132
## 4 1 1.60 a 0.0825 0.132
## 5 1 0.330 a 0.0825 0.132
## 6 1 -0.820 a 0.0825 0.132
## 7 1 0.487 a 0.0825 0.132
## 8 1 0.738 a 0.0825 0.132
## 9 1 0.576 a 0.0825 0.132
## 10 1 -0.305 a 0.0825 0.132
## # … with 149,990 more rows
And now we’ll compute the residuals within groups and globally:
sim_df <- sim_df %>%
mutate(overall_residuals = value - obs_overall_mean,
group_residuals = value - obs_group_mean)
sim_df## # A tibble: 150,000 x 7
## id value group obs_overall_mean obs_group_mean overall_residua…
## <int> <dbl> <chr> <dbl> <dbl> <dbl>
## 1 1 -0.626 a 0.0825 0.132 -0.709
## 2 1 0.184 a 0.0825 0.132 0.101
## 3 1 -0.836 a 0.0825 0.132 -0.918
## 4 1 1.60 a 0.0825 0.132 1.51
## 5 1 0.330 a 0.0825 0.132 0.247
## 6 1 -0.820 a 0.0825 0.132 -0.903
## 7 1 0.487 a 0.0825 0.132 0.405
## 8 1 0.738 a 0.0825 0.132 0.656
## 9 1 0.576 a 0.0825 0.132 0.493
## 10 1 -0.305 a 0.0825 0.132 -0.388
## # … with 149,990 more rows, and 1 more variable: group_residuals <dbl>
Next, we will compute residual sum of squares
rss_df <- sim_df %>%
group_by(id) %>%
summarise(RSS_0 = sum(overall_residuals^2),
RSS_1 = sum(group_residuals^2)) %>%
ungroup## `summarise()` ungrouping output (override with `.groups` argument)
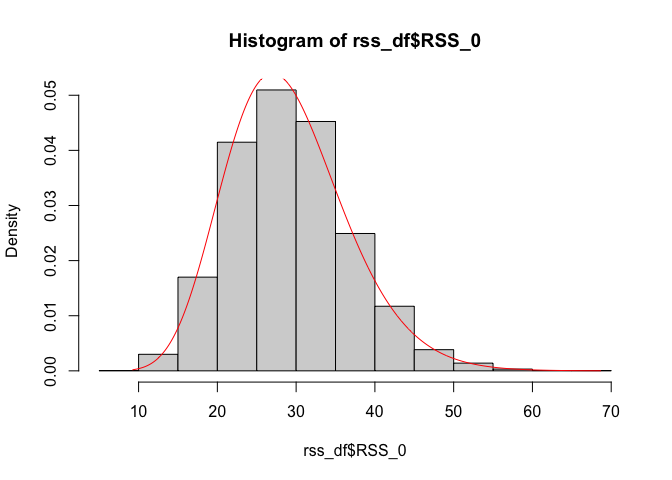
hist(rss_df$RSS_0, probability = TRUE)
lines(seq(min(rss_df$RSS_0), max(rss_df$RSS_0), 0.1),
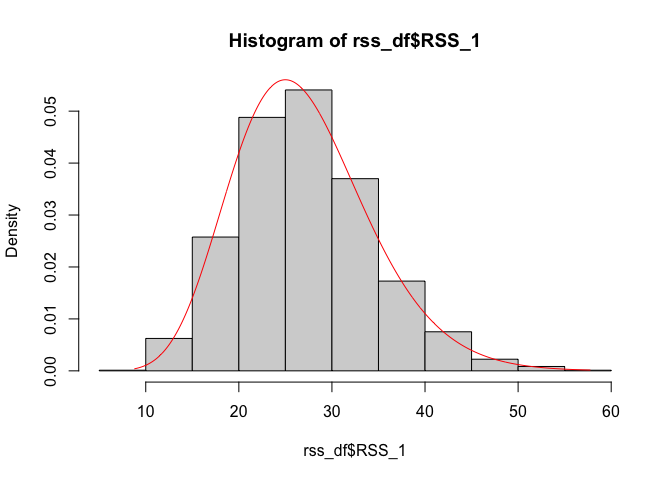
dchisq(seq(min(rss_df$RSS_0), max(rss_df$RSS_0), 0.1), df = n_groups * n_per_group - 1), col = "red")hist(rss_df$RSS_1, probability = TRUE)
lines(seq(min(rss_df$RSS_1), max(rss_df$RSS_1), 0.1),
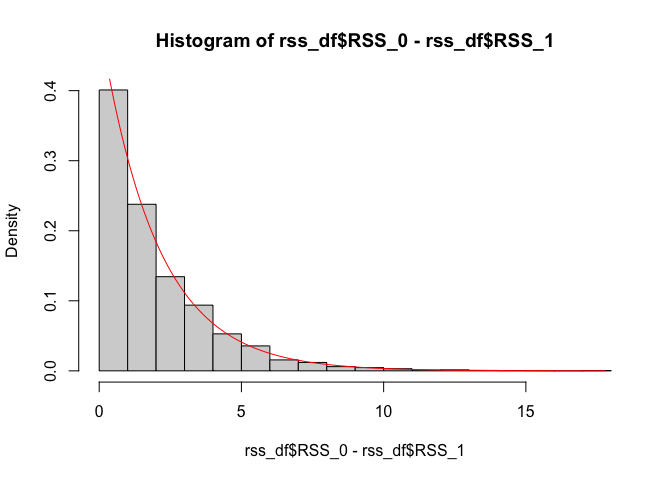
dchisq(seq(min(rss_df$RSS_1), max(rss_df$RSS_1), 0.1), df = n_groups * n_per_group - n_groups), col = "red")hist(rss_df$RSS_0 - rss_df$RSS_1, probability = TRUE)
lines(seq(min(rss_df$RSS_0 - rss_df$RSS_1),
max(rss_df$RSS_0 - rss_df$RSS_1), 0.1),
dchisq(seq(min(rss_df$RSS_0 - rss_df$RSS_1),
max(rss_df$RSS_0 - rss_df$RSS_1), 0.1), df = n_groups - 1), col = "red")d1 <- n_groups - 1
d2 <- n_groups * n_per_group - n_groups
Fstat <- (rss_df$RSS_0 - rss_df$RSS_1)/rss_df$RSS_1 * d2/d1
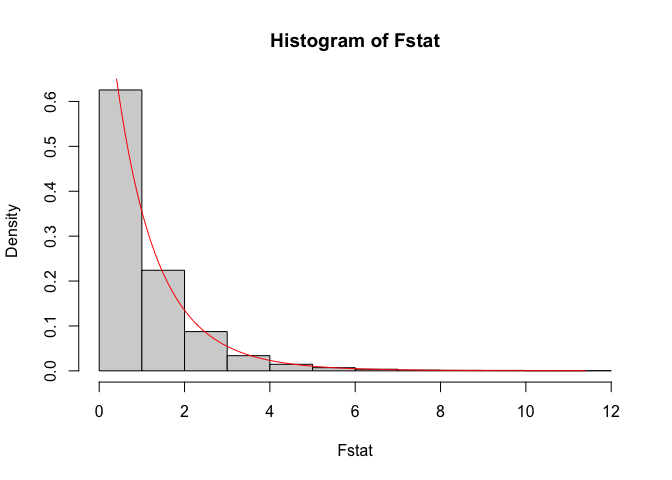
hist(Fstat, probability = TRUE)
lines(seq(min(Fstat), max(Fstat), 0.1),
df(seq(min(Fstat), max(Fstat), 0.1),
df1 = d1, df2 = d2), col = "red")