Python package for geometrical learning on protein structures.
To install this library, as is under development, we need to use testpipy index, it can be install with a simple command:
pip install -i https://test.pypi.org/simple/ protgeom -U
There is a helper to obtain a protein by name and parse the atomic structure
mol_7q0b = fetchPDB('7Q0B')
mol_7q0b.num_atoms21148
Atom positons are read as 3D cordines into a cartesian plance with Å as unit.
mol_7q0b.cordsarray([[193.53 , 206.683, 173.089],
[193.285, 208.002, 173.678],
[194.431, 208.461, 174.578],
...,
[210.995, 129.905, 137.104],
[210.455, 128.536, 137.509],
[209.865, 130.913, 136.954]])
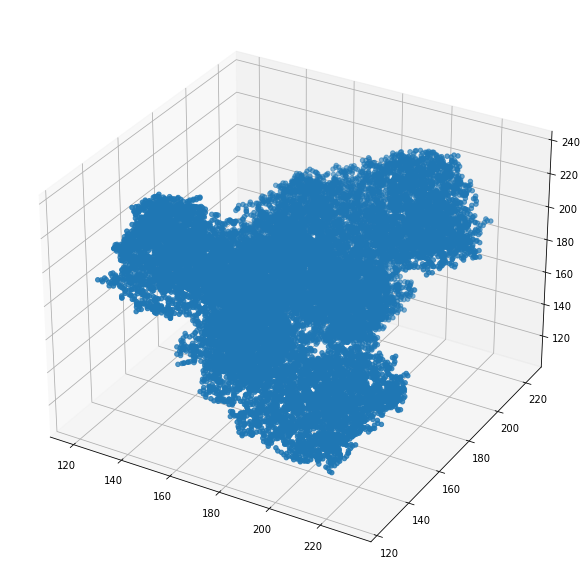
Once the libary loaded the 3D structure we can easily work with the molecule vectors:
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3Dfig = plt.figure()
fig.set_size_inches(18.5, 10.5)
ax = plt.axes(projection='3d')
ax.scatter3D(mol_7q0b.cords[:,0], mol_7q0b.cords[:,1], mol_7q0b.cords[:,2])