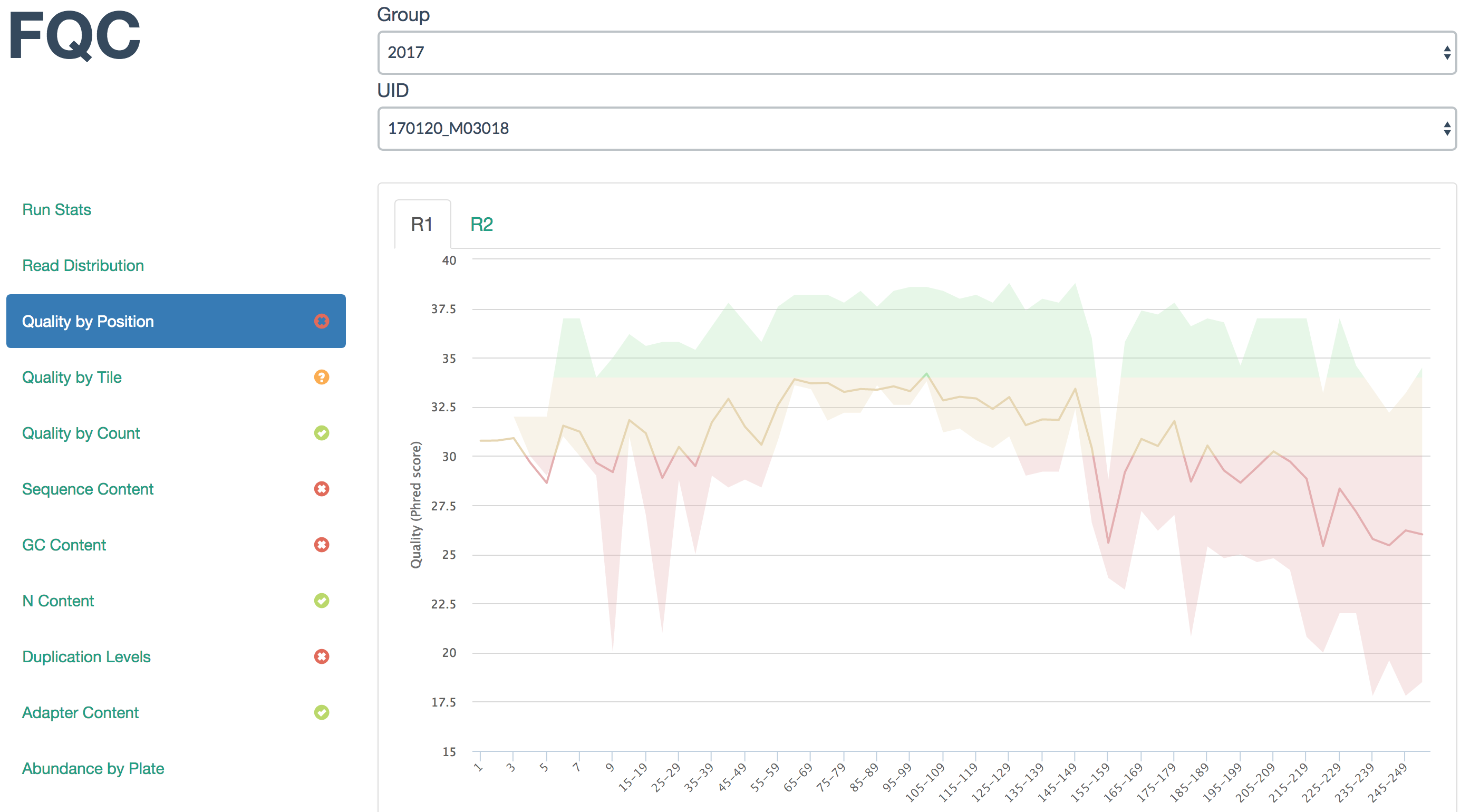
FQC is designed to better group FastQC result data across groups where each group is comprised of FASTQs related to an experiment or sequencing batch. Individual samples are grouped into paired-end sets when available and the dashboard's extensibility allows a user to add plots or tables as desired.
To see the full documentation and get started, see:
https://pnnl-fqc.readthedocs.io/
Live example:
Parsing the table and running FastQC is performed with code written for
Python 3. We recommend using Anaconda to
install the FastQC dependency.
conda config --add channels conda-forge
conda config --add channels defaults
conda config --add channels bioconda
conda install fastqc
The dashboard reads local files, so install where you will eventually be serving the site.
git clone https://github.com/pnnl/fqc.git
cd fqc
python setup.py install
This installs fqc command-line tool to process FASTQs and create the
dashboard.
Then to deploy a local copy from within the fqc directory, you can run:
python -m http.server --bind localhost 8000
And navigate to localhost:8000 in your browser.
By default, this will show the test data QC as determined by the data
directory in js/fqc.js:
var filePath = "/example/plot_data/"
Edit fqc.js to your local path within the fqc directory tree.
The first time this is run, it will build the entire backend of the site. Additional FASTQs being written to the same output directory are added to the backend according to Group ID and UID.
$ fqc qc group sample sample_r1.fastq.gz
[2016-07-26 13:24 INFO] Writing data to: plot_data/group/sample
[2016-07-26 13:24 INFO] Running FastQC
[2016-07-26 13:27 INFO] Extracting data from FastQC archives
[2016-07-26 13:27 INFO] Processing of sample1 complete
fqc qc --r2 sample_r2.fastq.gz 2016 sample sample_r1.fastq.gz
Joseph Brown, Meg Pirrung, Lee Ann McCue; FQC Dashboard: integrates FastQC results into a web-based, interactive, and extensible FASTQ quality control tool. Bioinformatics 2017 doi: 10.1093/bioinformatics/btx373