Open https://neo4j.het.io/browser/ and practise following cypher queries:
- Collect Symptoms for each of the disease
MATCH (d:Disease)--(s:Symptom) RETURN d.name,collect(s.name)
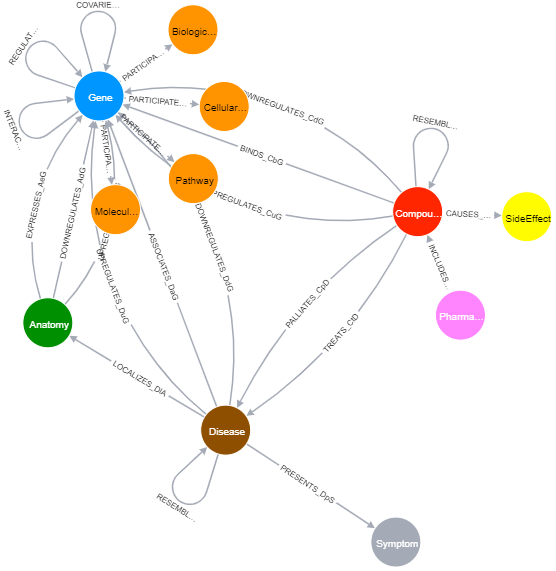
- Know schema of the graph data
CALL db.schema()
- Collect genes associated with each of the disease
MATCH (d:Disease)--(g:Gene) RETURN d.name,collect(g.name)
- Count gene and pathways for each diseases
MATCH (d:Disease)--(g:Gene)--(p:Pathway) RETURN d.name,count(g.name),count(p.name)
- Count and collect associated compound with disease
MATCH (d:Disease)--(c:Compound)--(s:SideEffect)
RETURN d.name,count(DISTINCT c.name),collect(DISTINCT c.name)
- Visualize a subgraph of associated compound with disease and side effects (100 nodes)
MATCH p=(d:Disease)--(c:Compound)--(s:SideEffect) RETURN p LIMIT 100
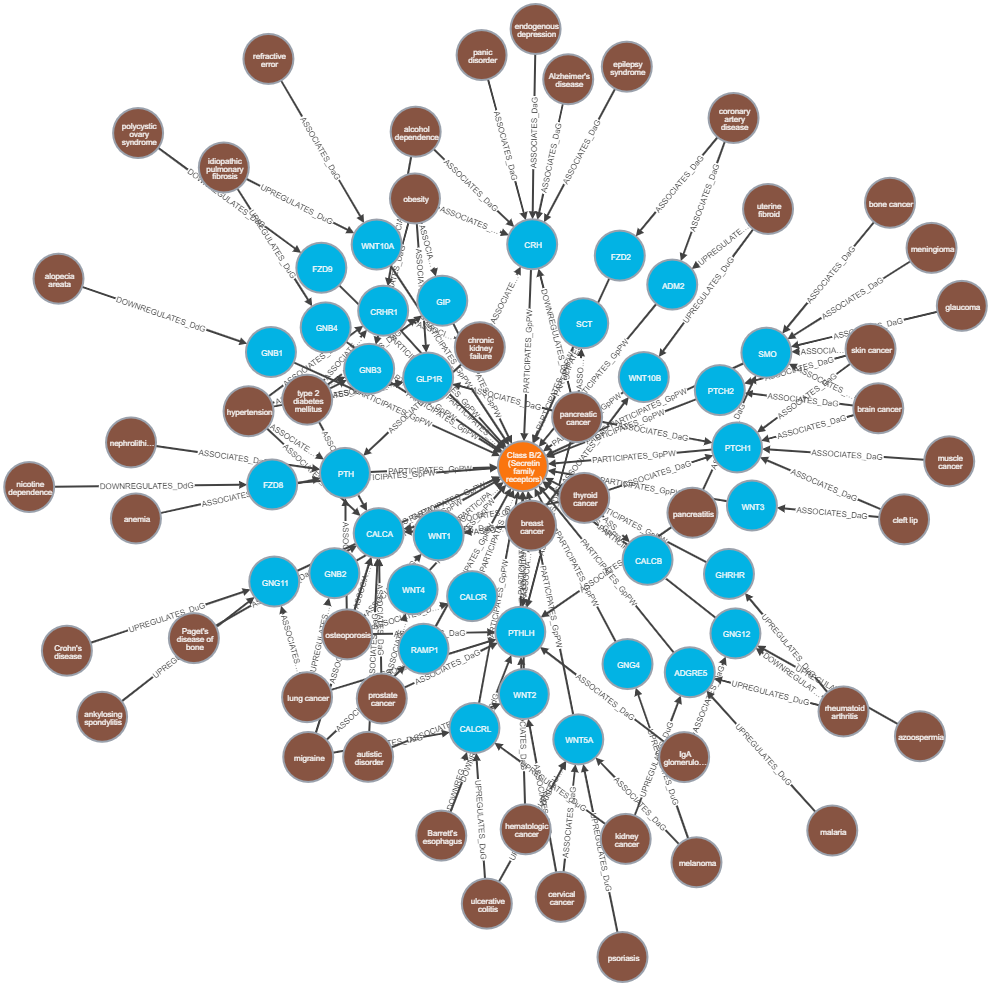
- Visualize the subgraph of Disease, Gene and Pathways (100 nodes)
MATCH path=(d:Disease)--(g:Gene)--(p:Pathway) RETURN path LIMIT 100