RNA and Protein expression analysis from Human Protein Atlas
A simple script to compare expression of a set of genes using data from HPA. It uses bioconductor and associated hparand biomaRt library.
The script will plot:
- Expression (pTPM - protein transcripts per million) in a host of cell lines - grouped by origin.
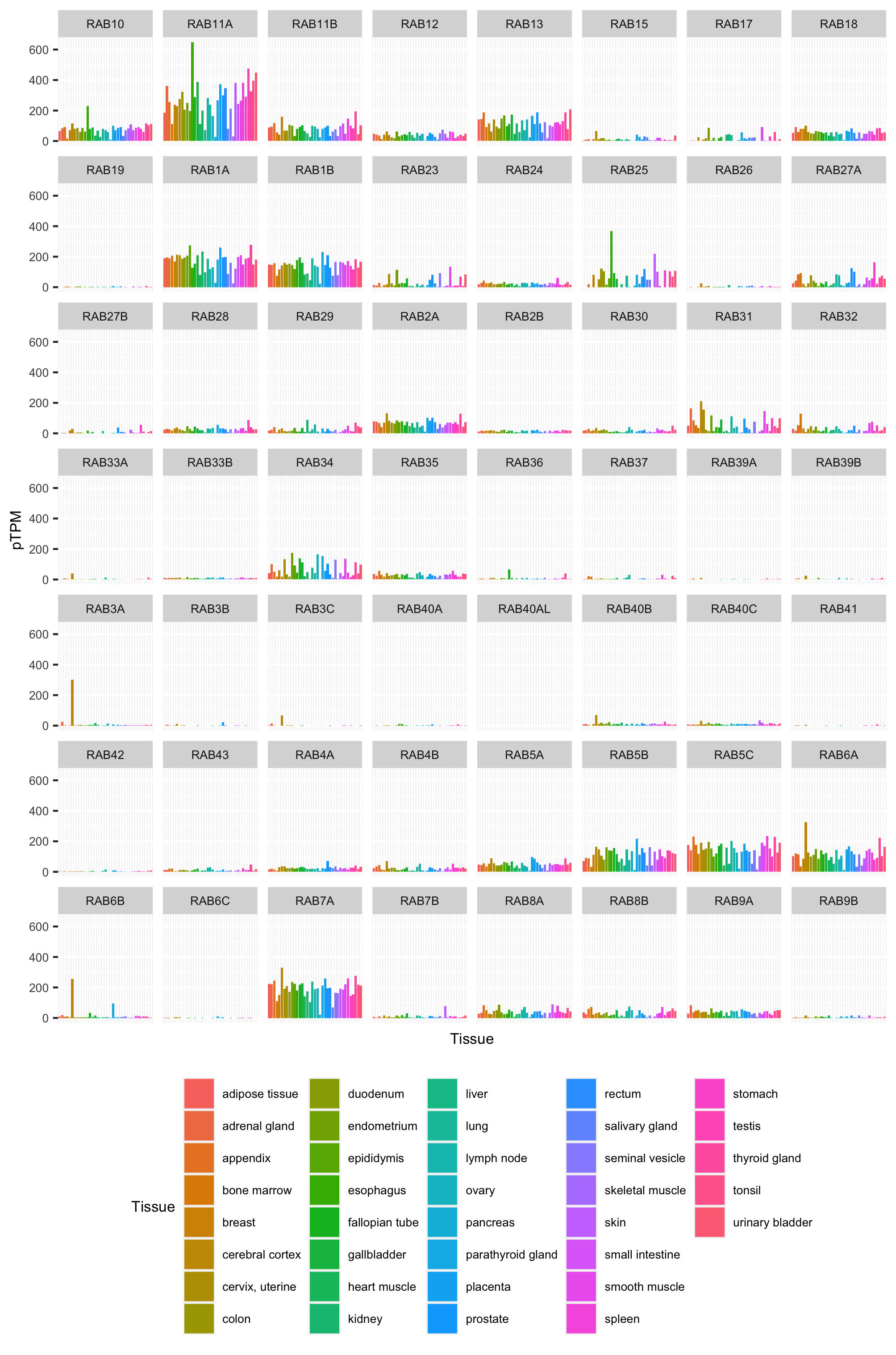
- Expression (pTPM) in different (normal) tissues.
- Protein expression in different tissues. Annoying this is not terribly useful as it is limited to antibody availability and also cannot be compared between proteins.
- Mean and stdev plots of pTPM expression to compare across the queried gene set.
Input is via a character vector of Ensembl IDs. In the script there are three ways illustrated to do the input:
- Manually input of a list of ids.
- Get a list of ids in some other format and convert them using
biomaRt. - Retrieve a list of Ensembl ids directly using a GO Term.
Some example graphs are shown in Output/Plots/ they show a comparison of all human Rab GTPase genes.
Directory setup for RStudio is as described here.