A Python front-end of [cpptraj program] (https://github.com/Amber-MD/cpptraj) (a data analysis package for biomolecular simulation): http://amber-md.github.io/pytraj
- Features
- Install
- How to get started?
- Contributors and Acknowledgement
- Citation
- Question/Suggestion?
- nglview with pytraj in Jupyter notebook
- Demo: Interactive data exploration with Jupyter notebook
- License
- support more than 80 types of data analyses (rmsd, radgyr, autoimage, pca, clustering,...)
- read/write various file formats (.nc, .dcd, .trr, .pdb, .mol2, ...)
- fast (core codes were written in C++ and Cython)
- support parallel calculation with trivial installation (openmp, multiprocessing, mpi, ...)
- interactive analysis with large trajectory data that does not fit to memory
- [>> many more with comprehensive tutorials] (http://amber-md.github.io/pytraj)
-
from source:
git clone https://github.com/amber-md/pytraj cd pytraj # linux python ./setup.py install # osx: pytraj will use clang to compile cython extension. # if you want to change compiler, please check setup.py file python setup.py install --disable-openmp # Note: pytraj will install the most updated cpptraj. We recommend to have netcdf library.
-
from conda (develop version for linux):
conda install -c https://conda.anaconda.org/ambermd pytraj-dev -
from AMBER suite distribution http://ambermd.org/. AMBER16 will be available soon (around April 2016).
-
getting trouble? : check our webpage
-
examples:
import pytraj as pt traj = pt.iterload("data.nc", "top.parm7") pt.rmsd(traj, ref=0, mask='@CA') pt.dssp(traj, ':2-16')
-
check our website: [http://amber-md.github.io/pytraj] (http://amber-md.github.io/pytraj)
If you would like to acknowledge our work, please cite both cpptraj and pytraj papers
-
[PTRAJ and CPPTRAJ] (http://pubs.acs.org/doi/abs/10.1021/ct400341p): Software for Processing and Analysis of Molecular Dynamics Trajectory Data Daniel R. Roe and Thomas E. Cheatham, III Journal of Chemical Theory and Computation 2013 9 (7), 3084-3095
-
PYTRAJ: Interactive data analysis for molecular dynamics simulations. Hai Nguyen, Daniel R. Roe, Jason Swails, David A. Case. (2016)
- code issue and stuff relating directly to
pytraj, create Issue - ask question about data analysis in general, send email to [AMBER Mailing List] (http://lists.ambermd.org/mailman/listinfo/amber)
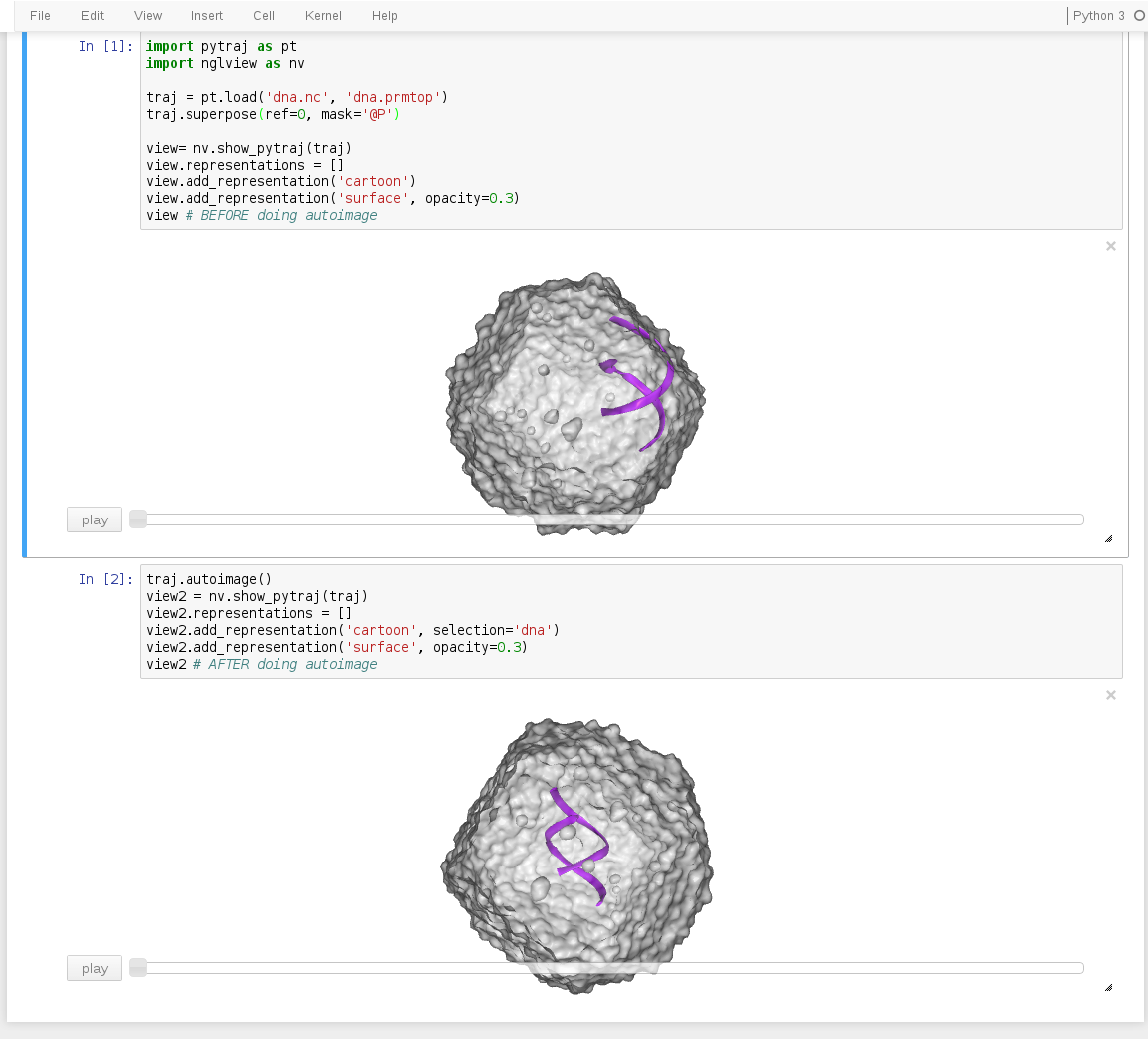
nglview with pytraj in Jupyter notebook
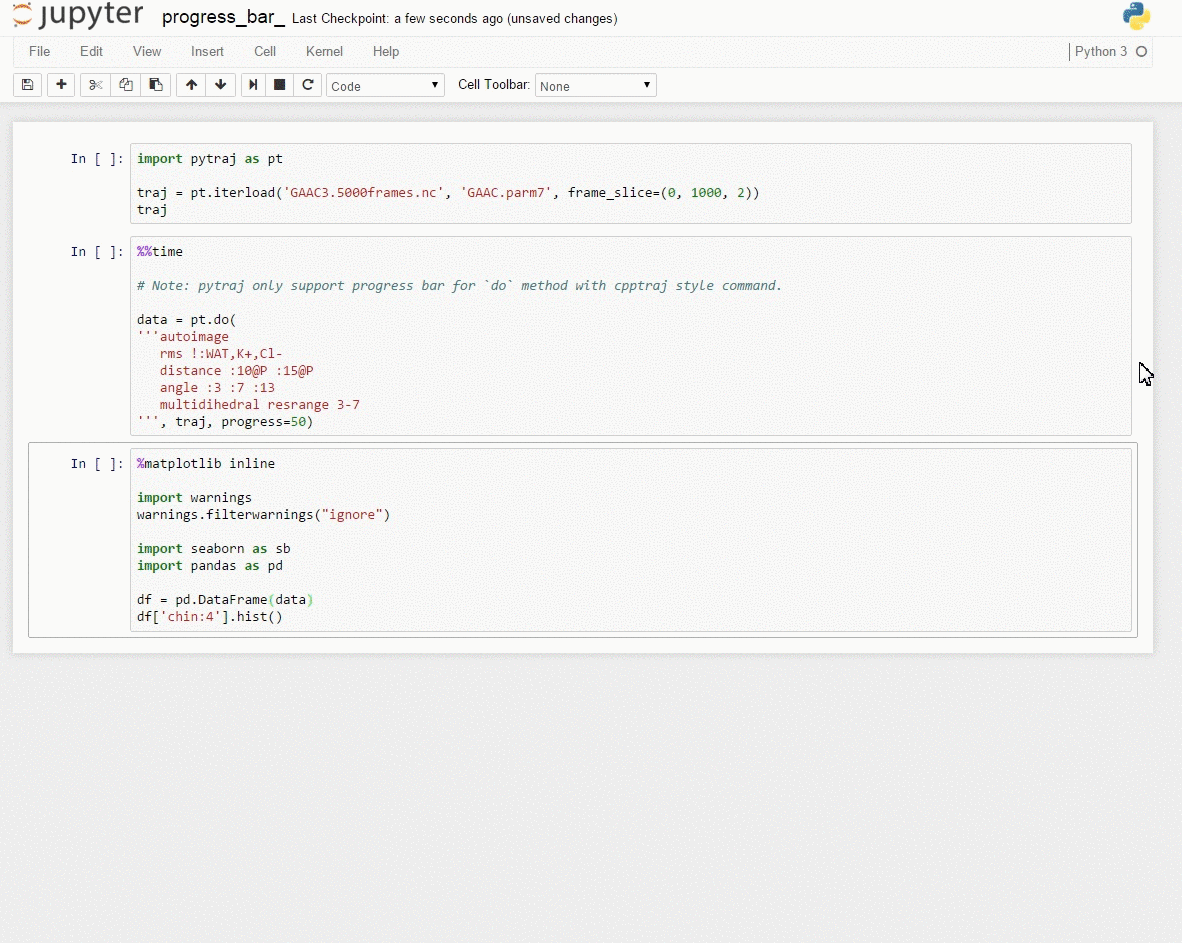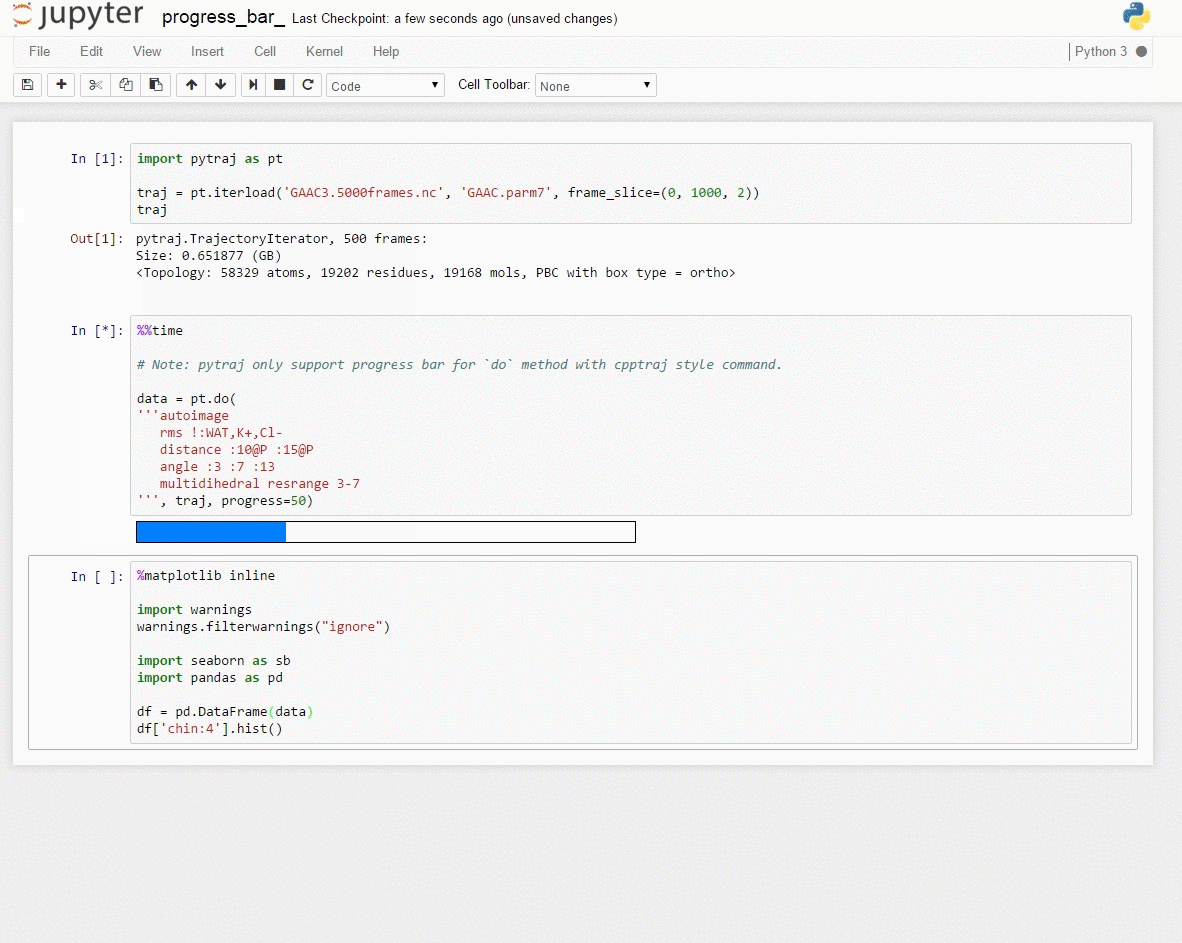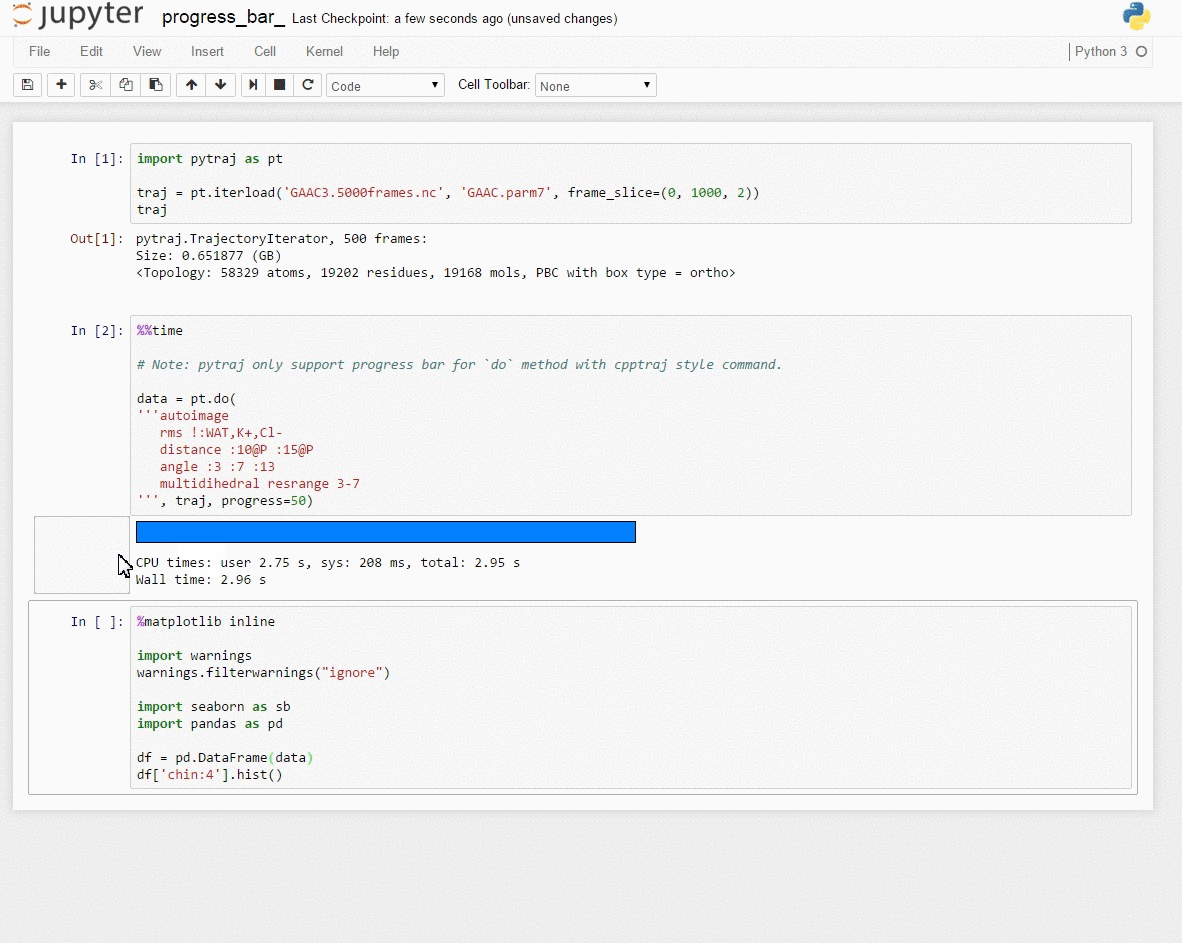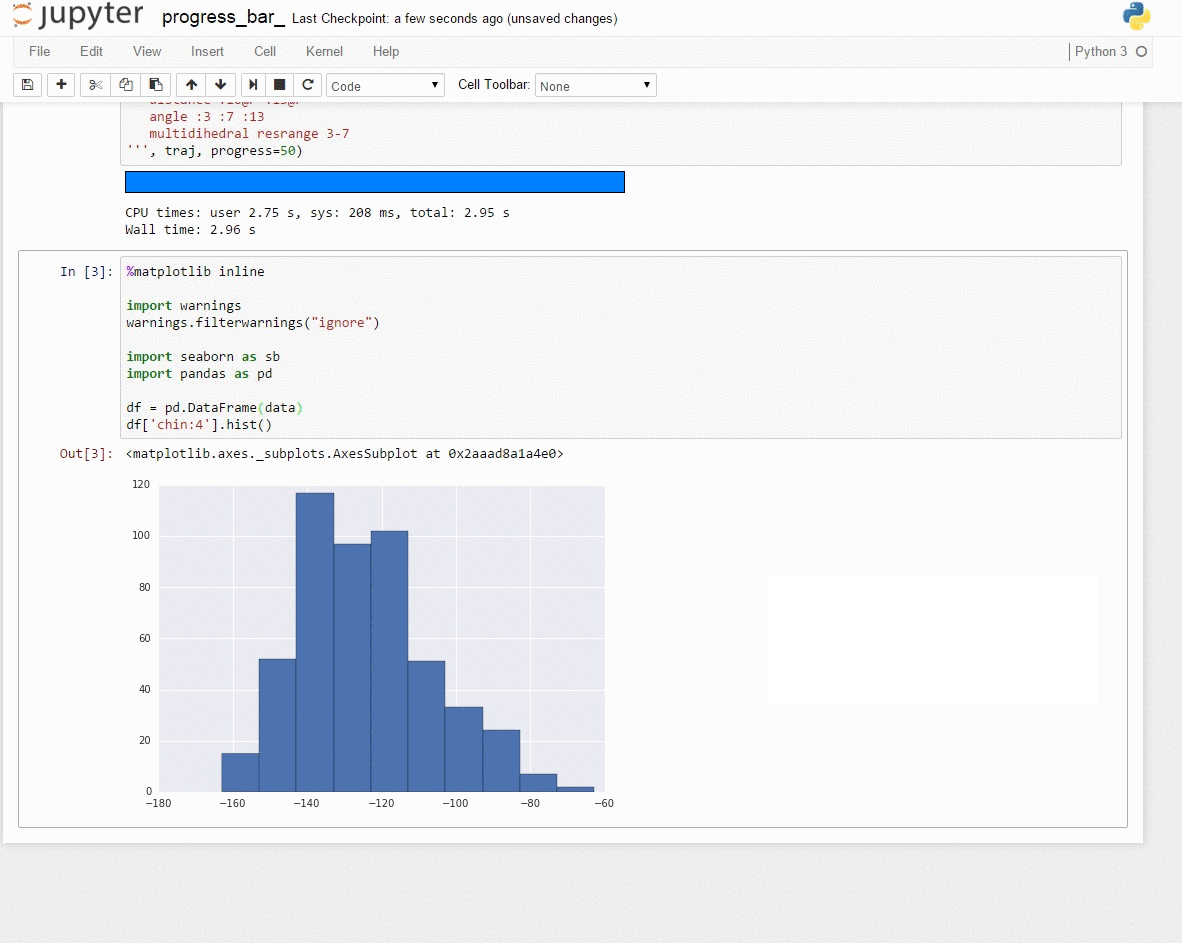
Demo: Interactive data exploration with [Jupyter notebook] (http://jupyter.org/)
BSD 2-Clause