Pipelines and scripts associated with the article
Garner RE, Gregory-Eaves I, Walsh DA. 2020. Sediment Metagenomes as Time Capsules of Lake Microbiomes. mSphere 5:e00512-20. https://doi.org/10.1128/mSphere.00512-20.
- 01_pipeline.md Bioinformatics pipeline that begins with raw metagenome reads, (1) trims the raw reads, (2) assembles metagenomes from the trimmed reads, and (3) maps trimmed reads to the metagenomes.
- 02_rnagenes.R Identify scaffolds containing conserved genes (ribosomal RNA or transfer rRNA genes) and count reads mapped (after removing scaffolds containing conserved genes).
- 03_idhist.R Read mapping vs. sequence identity in captured metagenomes.
- 04_genecontent.R Calculate gene counts on scaffolds in free and captured metagenomes.
- 05_covlength.R Scaffold coverage vs. length.
- 06_taxonomy.R Taxonomic analyses of free and captured metagenomes.
- 07_genera.R Gene counts for selected taxa in free metagenomes.
- 08_virusphylogeny.R Identify genes encoding Caudovirales terminase large subunit.
- 09_mapgenomes.R Map metagenome reads to reference reference genomes accessed from IMG.
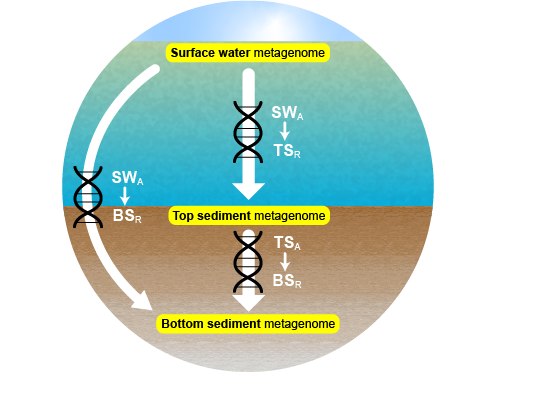
Note on metagenome file names: Numeric codes sometimes stand in for lake names: Lac Paula 06-126, Eightmile Lake 17-054, Grand lac Touradi 17-067. Surface water metagenomes are named watercolumn, top sediment metagenomes top, and bottom sediment metagenomes bottom.