Benchpress [1] is a Snakemake workflow where structure learning algorithms, implemented in possibly different languages, can be executed and compared. The computations scale seamlessly on multiple cores or "... to server, cluster, grid and cloud environments, without the need to modify the workflow definition" - Snakemake. The documentation is found at https://benchpressdocs.readthedocs.io. This section shows an overview of the supported evaluation methods.
- Snakemake ≥ 5.2 (installation instructions)
- Singularity ≥ 3.2 (installation instructions)
- Linux (Singularity currently only has a Beta release for OSX which is not enough)
As benchpress is a snakemake workflow, once the requirements are installed it requires no further installation but cloning the repository as
$ git clone https://github.com/felixleopoldo/benchpress.git
$ cd benchpress
On some systems, you might also have to explicitly install squash-tools. This can be done using conda as
$ conda install -c conda-forge squash-tools
Benchpress supports five different data scenarios built from combining different sources of graph parameters and data.
| Graph | Parameters | Data | |
|---|---|---|---|
| I | - | - | Fixed |
| II | Fixed | - | Fixed |
| III | Fixed | Fixed | Generated |
| IV | Fixed | Generated | Generated |
| V | Generated | Generated | Generated |
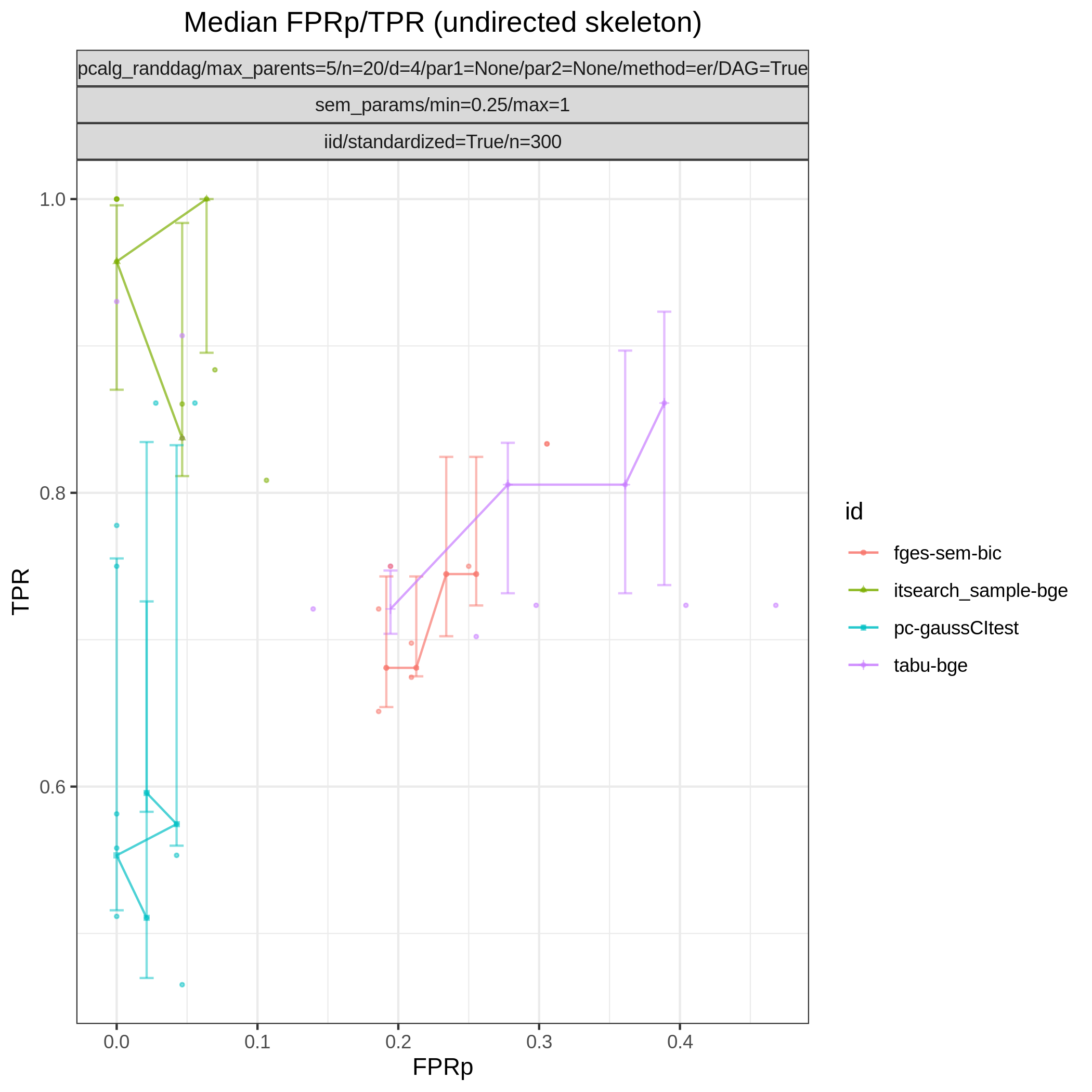
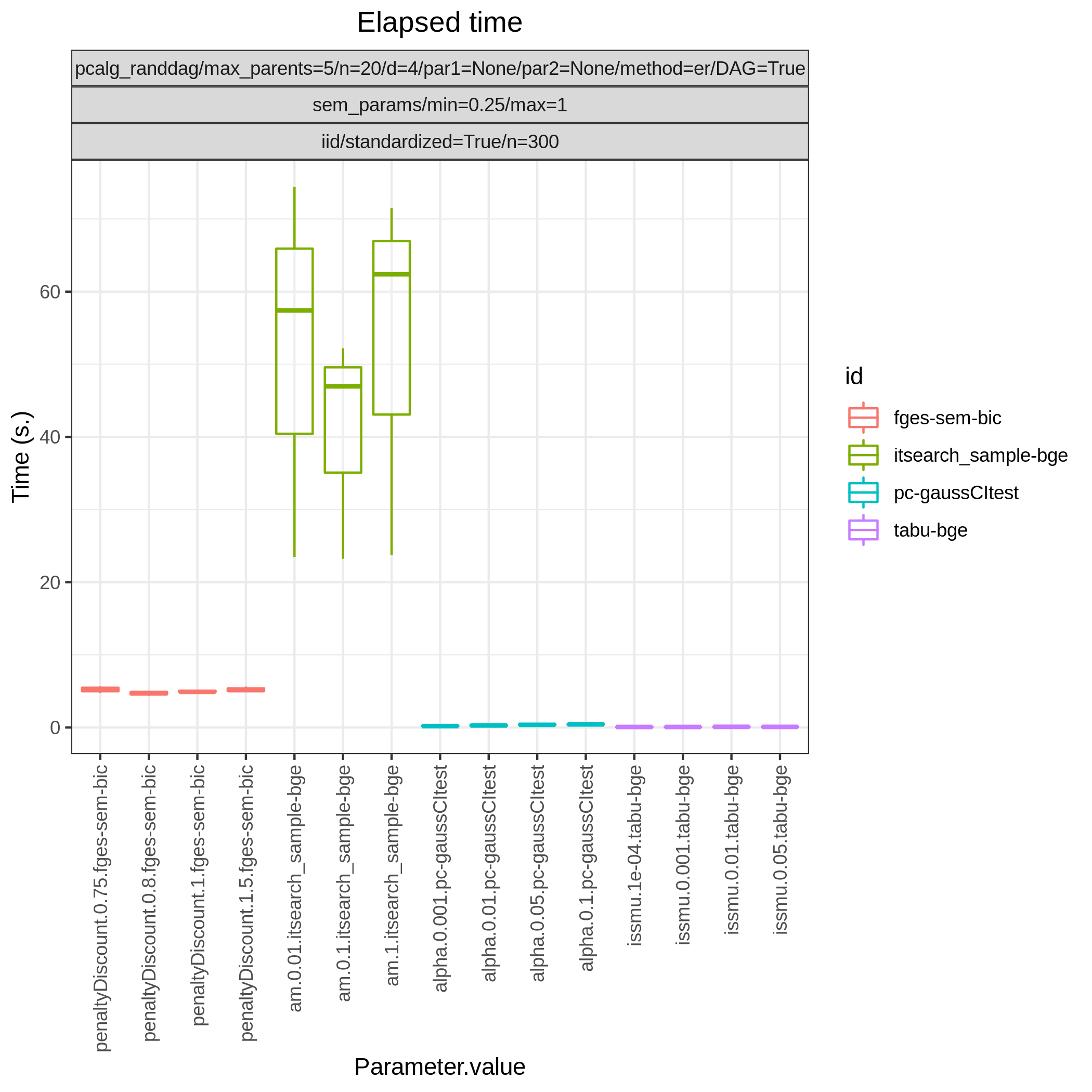
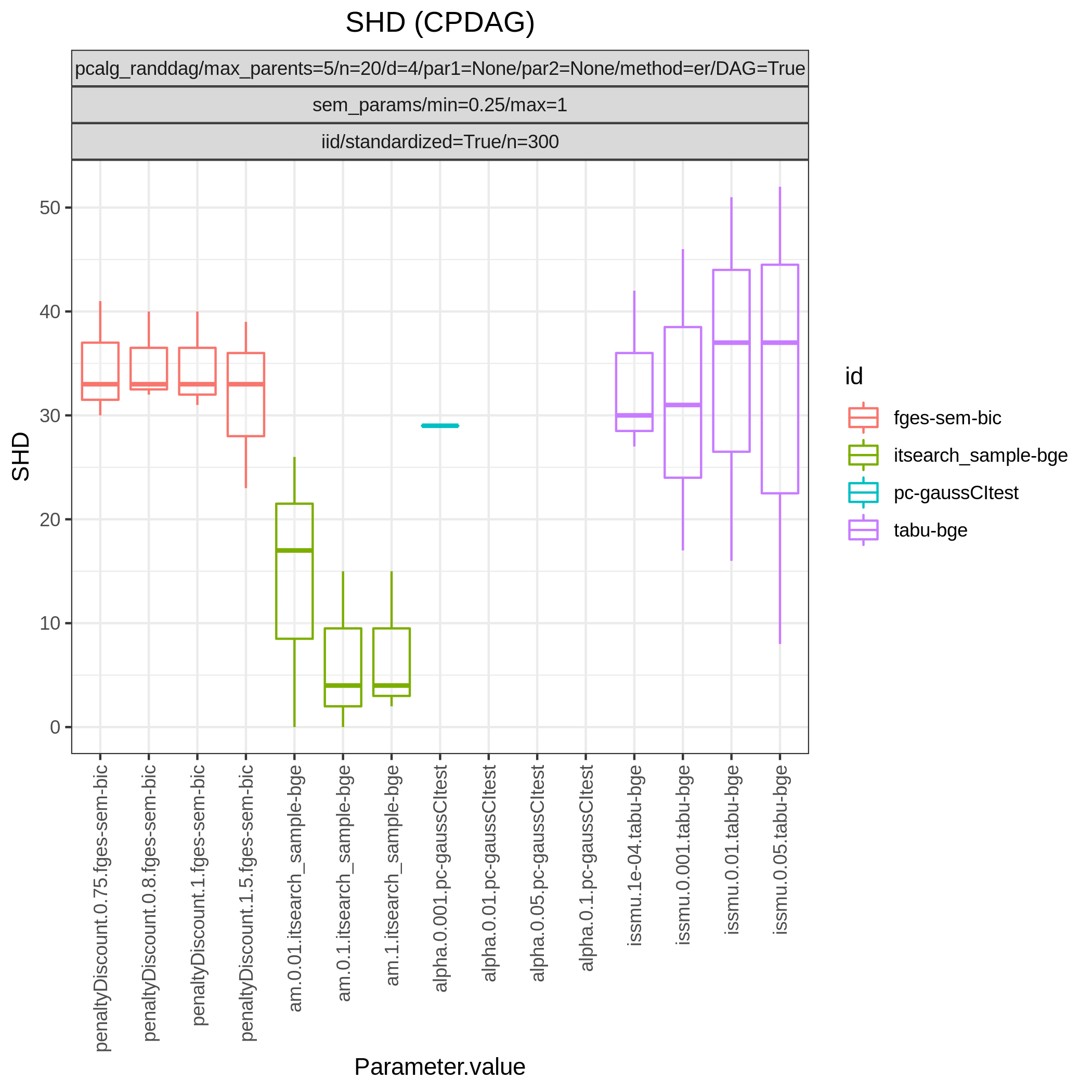
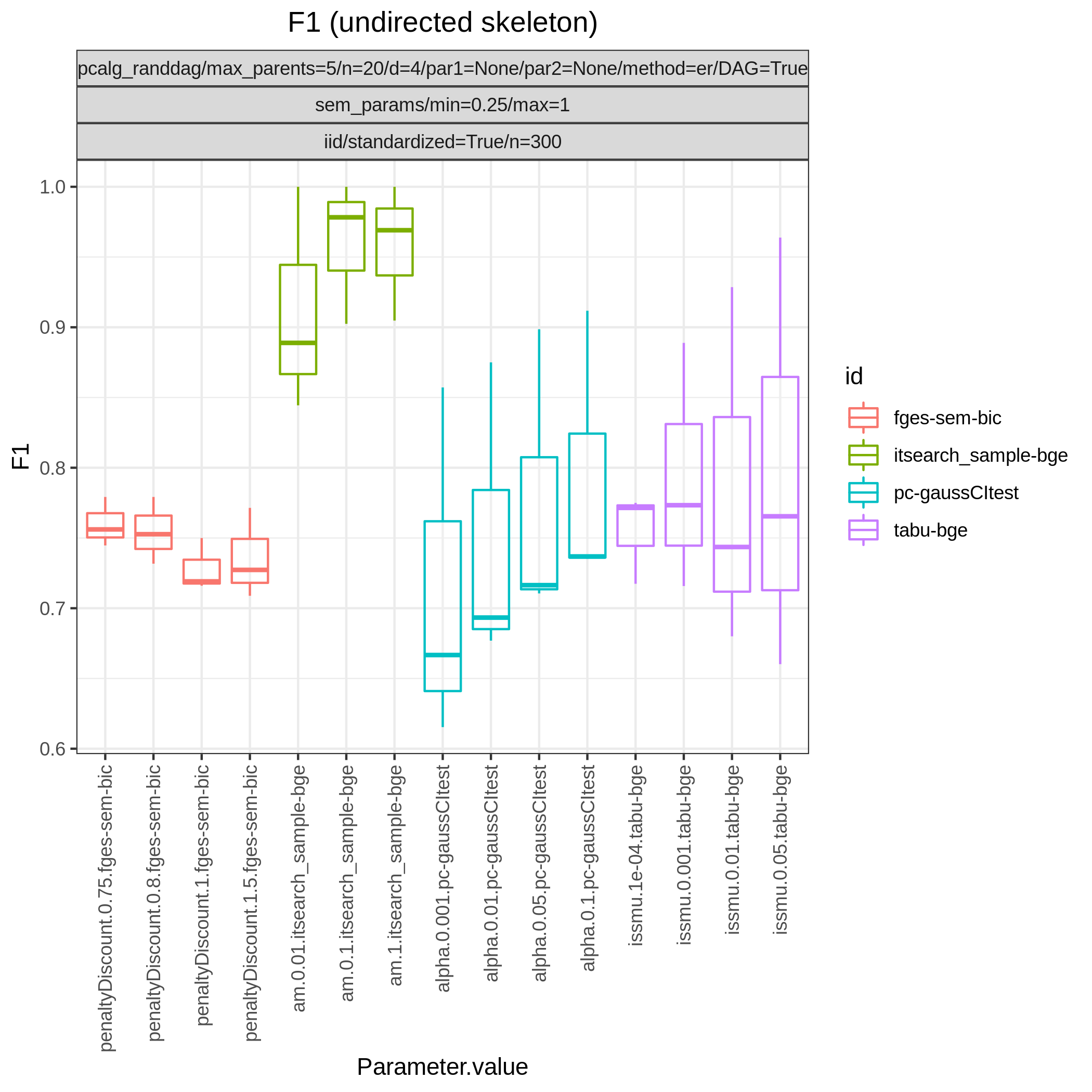
This study is an example of data scenario V based on three continuous datasets corresponing to three realisations of a random linear Gaussian structural equation model (SEM) with random DAG. The DAGs are sampled from a restricted Erdős–Rényi distribution using the pcalg_randdag module and the weight parameters are sampled uniformly using the sem_params module. For simplicity we use only a few structure learning modules here (bidag_itsearch, tetrad_fges, bnlearn_tabu, pcalg_pc) with different parameter settings. The full setup is found here config/ex.json.
To run this study (300 jobs ~ 10 minutes on a 2-cores laptop) type
$ snakemake --cores all --use-singularity --configfile config/ex.json
The following plots are generated by the roc module
| Method | Graph | Language | Library | Version | Module id |
|---|---|---|---|---|---|
| randDAG | DAG,UG | R | pcalg | 2.7-3 | pcalg_randdag |
| graph.sim | DG,UG | R | BDgraph | 2.64 | bdgraph_graphsim |
| CTA | DG | Python | trilearn | 1.2.3 | trilearn_cta |
| AR | DG | Python | trilearn | 1.2.3 | bandmat |
| AR random lag | DG | Python | trilearn | 1.2.3 | rand_bandmat |
| Fixed adjacency matrix | * | .csv | - | - | - |
| Distribution | Method | Graph | Language | Library | Version | Module id |
|---|---|---|---|---|---|---|
| Graph Wishart | rgwish | DG, UG | R | BDgraph | 2.64 | bdgraph_rgwish |
| Hyper Dirichlet [2] | - | DG | Python | trilearn | 1.2.3 | trilearn_hyper-dir |
| Graph intra-class | - | DG | Python | trilearn | 1.2.3 | trilearn_intra-class |
| Random SEM parameters | - | DAG | R | - | - | sem_params |
| Random probability tables | - | DAG | R | - | - | bin_bn |
| Fixed bn.fit object | - | DAG | .rds | bnlearn | - | - |
| Fixed SEM parameter matrix | - | DAG | .csv | - | - | - |
| Algorithm | Graph | Language | Library | Version | Module id |
|---|---|---|---|---|---|
| GOBNILP | DAG | C | GOBNILP (bitbucket) | #e60ef14 | gobnilp |
| ASOBS | DAG | R/Java | r.blip | 1.1 | rblip_asobs |
| FGES | CPDAG | Java | TETRAD (causal-cmd) | 1.1.3 | tetrad_fges |
| FCI | DAG | Java | TETRAD (causal-cmd) | 1.1.3 | tetrad_fci |
| RFCI | CPDAG | Java | TETRAD (causal-cmd) | 1.1.3 | tetrad_rfci |
| GFCI | DAG | Java | TETRAD (causal-cmd) | 1.1.3 | tetrad_gfci |
| PC | CPDAG | R | pcalg | 2.7-3 | pcalg_pc |
| Dual PC | CPDAG | R | dualPC (github) | 4a5175d | giudice_dualpc |
| No tears | DAG | Python | jmoss20 (github) | #0c032a0 | notears |
| HC | DAG | R | bnlearn | 4.7 | bnlearn_hc |
| MMHC | DAG | R | bnlearn | 4.7 | bnlearn_mmhc |
| Inter-IAMB | CPDAG | R | bnlearn | 4.7 | bnlearn_interiamb |
| GS | DAG | R | bnlearn | 4.7 | bnlearn_gs |
| Tabu | DAG | R | bnlearn | 4.7 | bnlearn_tabu |
| PC stable | CPDAG | R | bnlearn | 4.7 | bnlearn_pcstable |
| IAMB | DAG | R | bnlearn | 4.7 | bnlearn_iamb |
| Fast IAMB | DAG | R | bnlearn | 4.7 | bnlearn_fastiamb |
| IAMB FDR | DAG | R | bnlearn | 4.7 | bnlearn_iambfdr |
| MMPC | DAG | R | bnlearn | 4.7 | bnlearn_mmpc |
| SI HITON-PC | DAG | R | bnlearn | 4.7 | bnlearn_sihitonpc |
| Hybrid PC | DAG | R | bnlearn | 4.7 | bnlearn_hpc |
| H2PC | DAG | R | bnlearn | 4.7 | bnlearn_h2pc |
| RSMAX2 | DAG | R | bnlearn | 4.7 | bnlearn_rsmax2 |
| Iterative MCMC | DAG | R | BiDAG | 2.0.3 | bidag_itsearch |
| Order MCMC | DAG | R | BiDAG | 2.0.3 | bidag_order_mcmc |
| Partition MCMC | DAG | R | BiDAG | 2.0.3 | bidag_partition_mcmc |
| PGibbs | DG | Python | Trilearn | 1.2.3 | trilearn_pgibbs |
| GG99 single pair | DG | Java | A. Thomas | - | gg99_singlepair |
| GT13 multi pair | DG | Java | A. Thomas | - | gt13_multipair |
| GLasso | UG | Python | scikit-learn | 0.22.1 | sklearn_glasso |
| Method | Language | Module id |
|---|---|---|
| I.I.D. data samples | - | iid |
| Fixed data file | .csv | - |
| Function | Language | Library | Module id |
|---|---|---|---|
| Plot true graphs | - | graphviz | graph_true_plots |
| Plot true graphs properties | - | ggplot2 | graph_true_stats |
| Plot estimated graphs | - | graphviz | graph_plots |
| Timing and ROC curves for TPR,FPR,FNR,... | R | ggplot2 | roc |
| MCMC mean graph | Python | seaborn | mcmc_heatmaps |
| MCMC auto-correlation | Python | pandas | mcmc_autocorr_plots |
| MCMC trajectory | Python | pandas | mcmc_traj_plots |
Acronyms are used for Directed Acyclic Graphs (DAGs), Undirected Graphs (UGs), Decomposable Graphs (DGs), and Completed Partially DAGs (CPDAGs).
@misc{rios2021benchpress,
title={Benchpress: a scalable and platform-independent workflow for benchmarking structure learning algorithms for graphical models},
author={Felix L. Rios and Giusi Moffa and Jack Kuipers},
year={2021},
eprint={2107.03863},
archivePrefix={arXiv},
primaryClass={stat.ML}
}
- [1] Felix L. Rios and Giusi Moffa and Jack Kuipers Benchpress: a scalable and platform-independent workflow for benchmarking structure learning algorithms for graphical models. ArXiv eprints., 2107.03863, 2021.
- [2] Dawid AP, Lauritzen SL (1993). “Hyper Markov laws in the statistical analysis of decomposable graphical models.” The Annals of Statistics, 21(3), 1272–1317.
Contrubutions are very welcomed
- Fork it!
- Create your feature branch:
git checkout -b my-new-feature - Commit your changes:
git commit -am 'Add some feature' - Push to the branch:
git push origin my-new-feature - Open a pull request
This project is licensed under the GPL-2.0 License - see the LICENSE file for details