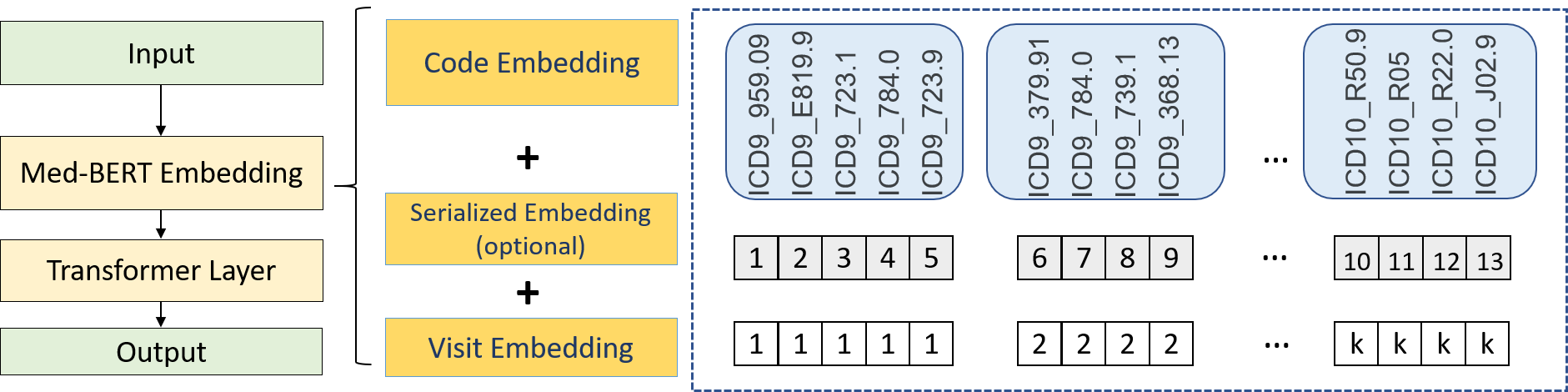
This repository provides the code for pre-training and fine-tuning Med-BERT, a contextualized embedding model that delivers a meaningful performance boost for real-world disease-prediction problems as compared to state-of-the-art models.
Med-Bert adapts bidirectional encoder representations from transformers (BERT) framework and pre-trains contextualized embeddings for diagnosis codes mainly in ICD-9 and ICD-10 format using structured data from an EHR dataset containing 28,490,650 patients.
 Please refer to our paper Med-BERT: pre-trained contextualized embeddings on large-scale structured electronic health records for disease prediction for more details.
Please refer to our paper Med-BERT: pre-trained contextualized embeddings on large-scale structured electronic health records for disease prediction for more details.
To reproduce the steps necessary for pre-training Med-BERT
python preprocess_pretrain_data.py <data_File> <vocab/NA> <output_Prefix> <subset_size/0forAll>
python create_BERTpretrain_EHRfeatures.py --input_file=<output_Prefix.bencs.train> --output_file='output_file' --vocab_file=<output_Prefix.types>--max_predictions_per_seq=1 --max_seq_length=64
python run_EHRpretraining.py --input_file='output_file' --output_dir=<path_to_outputfolder> --do_train=True --do_eval=True --bert_config_file=config.json --train_batch_size=32 --max_seq_length=512 --max_predictions_per_seq=1 --num_train_steps=4500000 --num_warmup_steps=10000 --learning_rate=5e-5
You can find an example for the construction of the data_file under Example data as well as images showing the construction of preprocessed data and the BERT features. Additional details are available under Pretraining Tutorial
Note: We run our code using mainly GPU, while CPU and TPU options migt be available in the code they were not tested.
To see an example of how to use Med-BERT for a specific disease prediction task, you can follow the Med-BERT DHF prediction notebook
Kindly note that you need to use the following code for preparing the fine-tunning data using (create_ehr_pretrain_FTdata.py) in a similar way of preparing the pretraining data.
Python: 3.7+
Pytorch 1.5.0
Tensorflow 1.13.1+
Pandas
Pickle
tqdm
pytorch-transformers
Google BERT
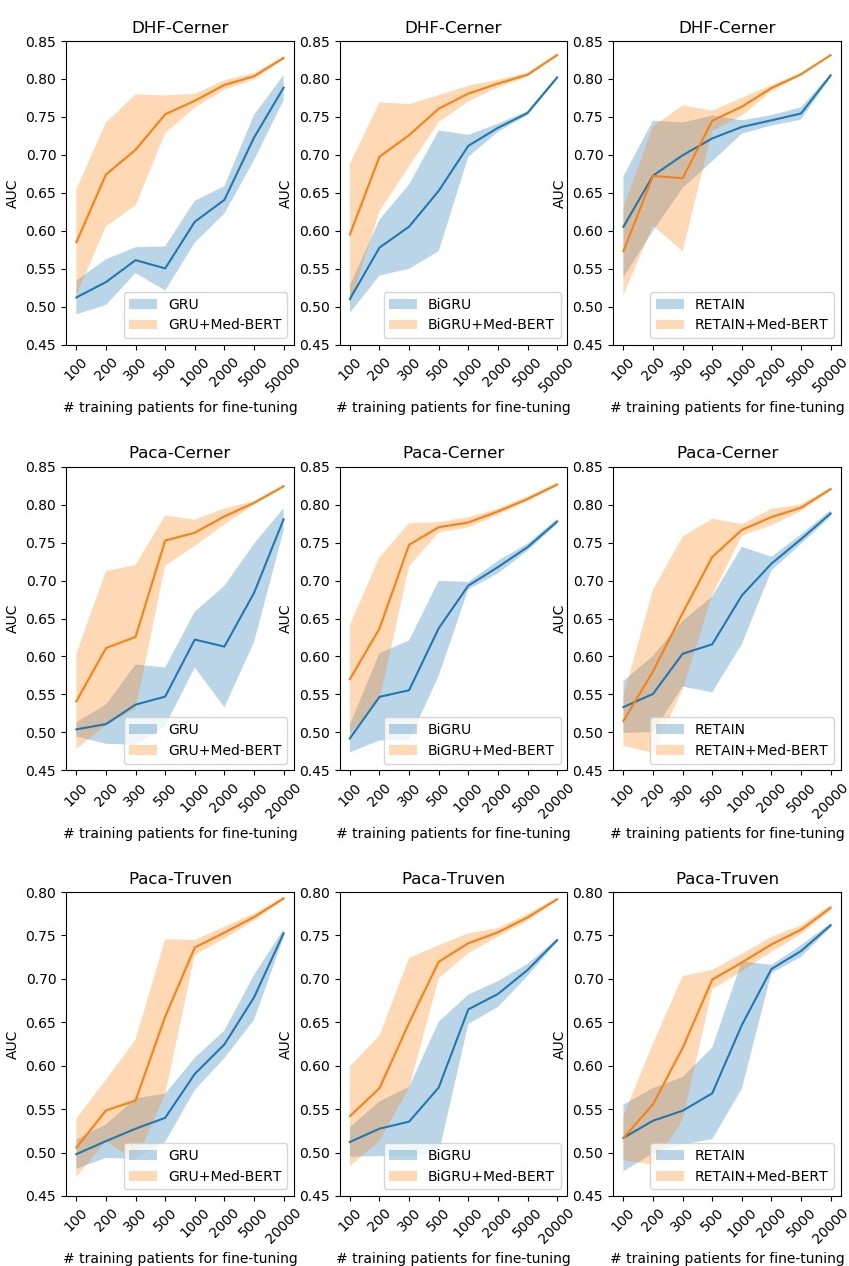
Prediction results for the evaluation sets by training on different sizes of data on DHF-Cerner (top), PaCa-Cerner (middle), and PaCa-Truven (bottom). The shadows indicate the standard deviations. Please refer to our paper for more details.
Details are still being worked out on how to share the pre-trained model in a responsible manner. Stay tuned.
Please post a Github issue if you have any questions.
Please acknowledge the following work in papers or derivative software:
Laila Rasmy, Yang Xiang, Ziqian Xie, Cui Tao, and Degui Zhi. "Med-BERT: pre-trained contextualized embeddings on large-scale structured electronic health records for disease prediction." arXiv preprint arXiv:2005.12833 (2020).