Project bulk/single-cell RNA-seq data to given LSI space.
The methods are implemented according to the article Granja, J.M., Klemm, S., McGinnis, L.M. et al. Single-cell multiomic analysis identifies regulatory programs in mixed-phenotype acute leukemia. Nat Biotechnol 37, 1458–1465 (2019) and the code of this paper available.
More features may be added in future. (etc. supporting for ATAC-seq)
devtools::install_github("sajuukLyu/projectLSI")Next, let's try to project a single-cell RNA-seq dataset pbmc4k and a bulk RNA-seq dataset bulk.data into the same single-cell RNA-seq dataset pbmc3k, keeping the original LSI & UMAP coordinate unchanged.
pbmc3k and pbmc4k datasets are from package TENxPBMCData, and bulk.data is part of GSE74246.
library(Seurat)
library(projectLSI)
library(patchwork)
data(pbmc3k)
data(pbmc4k)
data(bulk.data)
pbmc3k
## An object of class Seurat
## 32738 features across 2700 samples within 1 assay
## Active assay: RNA (32738 features)
pbmc4k
## An object of class Seurat
## 33694 features across 4340 samples within 1 assay
## Active assay: RNA (33694 features)
dim(bulk.data)
## [1] 25498 20The preprocess pipeline is same with standard Seurat pipeline.
# for pbmc3k
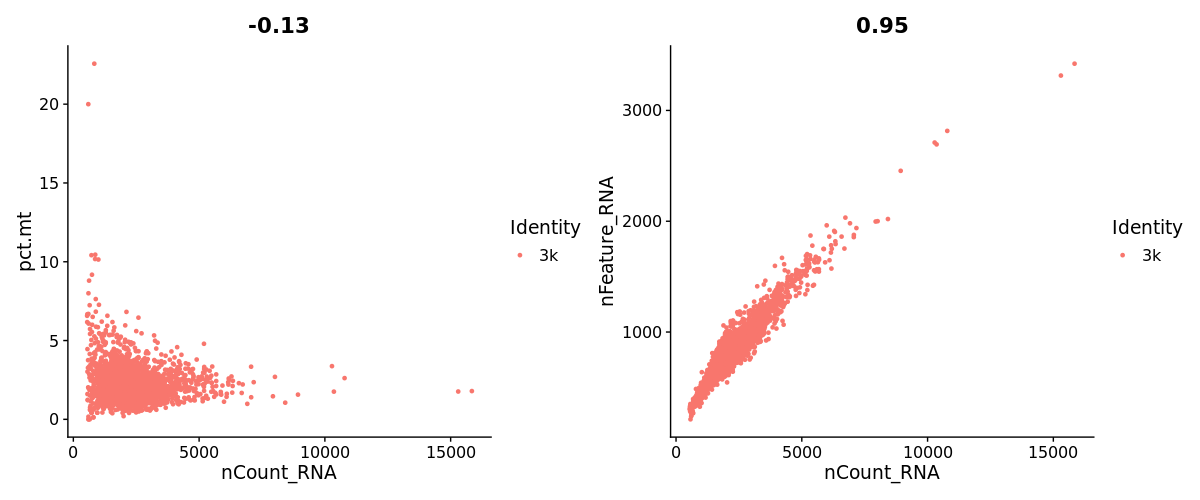
pbmc3k$pct.mt <- PercentageFeatureSet(pbmc3k, pattern = "^MT-")
FeatureScatter(pbmc3k, feature1 = "nCount_RNA", feature2 = "pct.mt") +
FeatureScatter(pbmc3k, feature1 = "nCount_RNA", feature2 = "nFeature_RNA")pbmc3k <- subset(pbmc3k, subset = nFeature_RNA > 200 & nFeature_RNA < 2500 & pct.mt < 5)
pbmc3k <- NormalizeData(pbmc3k)
## Performing log-normalization
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
pbmc3k <- FindVariableFeatures(pbmc3k, nfeatures = 2000)
## Calculating gene variances
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
## Calculating feature variances of standardized and clipped values
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
# for pbmc4k
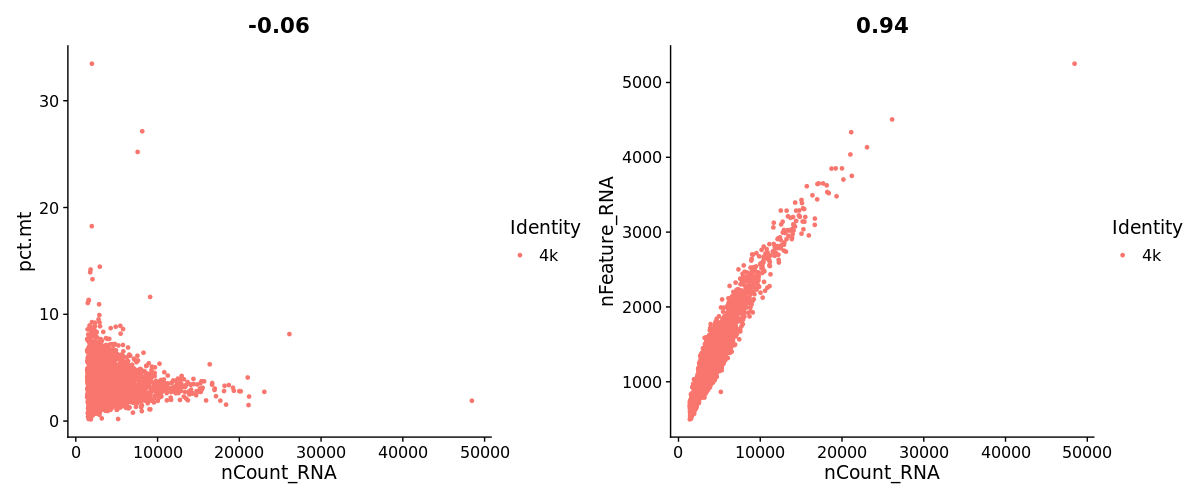
pbmc4k$pct.mt <- PercentageFeatureSet(pbmc4k, pattern = "^MT-")
FeatureScatter(pbmc4k, feature1 = "nCount_RNA", feature2 = "pct.mt") +
FeatureScatter(pbmc4k, feature1 = "nCount_RNA", feature2 = "nFeature_RNA")pbmc4k <- subset(pbmc4k, subset = nFeature_RNA > 200 & nFeature_RNA < 3500 & pct.mt < 8)
pbmc4k <- NormalizeData(pbmc4k)
## Performing log-normalization
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
pbmc4k <- FindVariableFeatures(pbmc4k, nfeatures = 2000)
## Calculating gene variances
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
## Calculating feature variances of standardized and clipped values
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|Then we can perform term frequency–inverse document frequency (TF-IDF) transformation and latent semantic indexing (LSI) on the normalized pbmc3k data. Notice that we should use variable genes as input.
pbmc3k.lsi <- calcLSI(pbmc3k[["RNA"]]@data[VariableFeatures(pbmc3k), ])
pbmc3k[["pca"]] <- CreateDimReducObject(
embeddings = pbmc3k.lsi$matSVD,
loadings = pbmc3k.lsi$fLoad,
assay = "RNA",
stdev = pbmc3k.lsi$sdev,
key = "PC_")
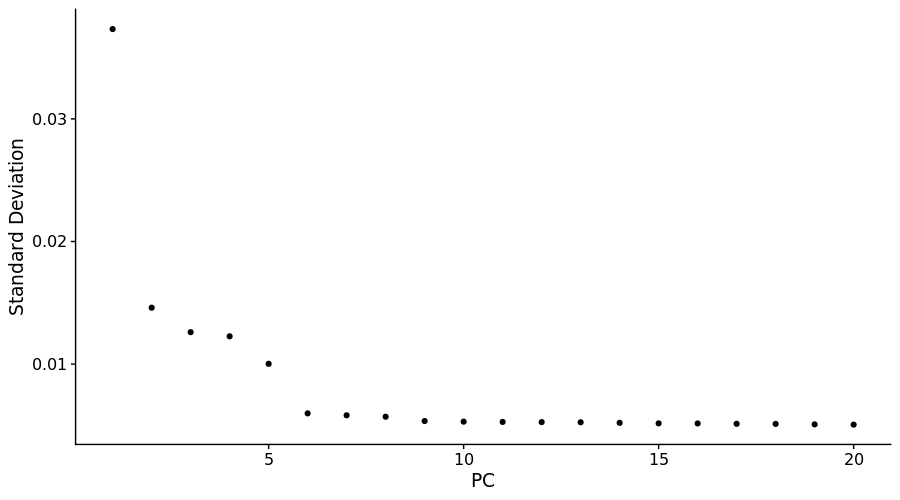
ElbowPlot(pbmc3k)Then we can use Seurat v3 provided functions to cluster the cells.
pbmc3k <- FindNeighbors(pbmc3k, dims = 1:10)
## Computing nearest neighbor graph
## Computing SNN
pbmc3k <- FindClusters(pbmc3k, resolution = 0.6)
## Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
##
## Number of nodes: 2638
## Number of edges: 97177
##
## Running Louvain algorithm...
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
## Maximum modularity in 10 random starts: 0.8542
## Number of communities: 9
## Elapsed time: 0 secondsNotice that the first 10 PCs have enough information of the whole dataset. So we can perform UMAP on the first 10 PCs using R package uwot. Notice that ret_model parameter should be TRUE for later projection.
set.seed(42)
umap.pbmc3k <- uwot::umap(pbmc3k.lsi$matSVD[, 1:10],
n_neighbors = 30,
min_dist = 0.5,
metric = "euclidean",
ret_model = T,
verbose = T)
## 00:58:06 UMAP embedding parameters a = 0.583 b = 1.334
## 00:58:06 Read 2638 rows and found 10 numeric columns
## 00:58:06 Using Annoy for neighbor search, n_neighbors = 30
## 00:58:06 Building Annoy index with metric = euclidean, n_trees = 50
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
## 00:58:06 Writing NN index file to temp file /tmp/RtmpbWcqgH/file17b95ca52051
## 00:58:06 Searching Annoy index using 8 threads, search_k = 3000
## 00:58:06 Annoy recall = 100%
## 00:58:07 Commencing smooth kNN distance calibration using 8 threads
## 00:58:07 Initializing from normalized Laplacian + noise
## Spectral initialization failed to converge, using random initialization instead
## 00:58:07 Commencing optimization for 500 epochs, with 110674 positive edges
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
## 00:58:12 Optimization finished
umap.pbmc3k.emb <- umap.pbmc3k$embedding
rownames(umap.pbmc3k.emb) <- colnames(pbmc3k)
colnames(umap.pbmc3k.emb) <- paste0("UMAP_", seq_len(ncol(umap.pbmc3k.emb)))
pbmc3k[["umap"]] <- CreateDimReducObject(
embeddings = umap.pbmc3k.emb,
assay = "RNA",
key = "UMAP_")
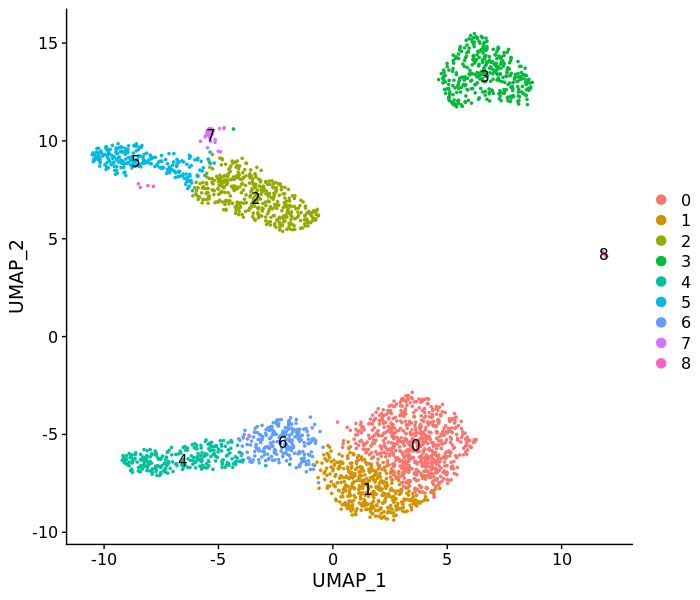
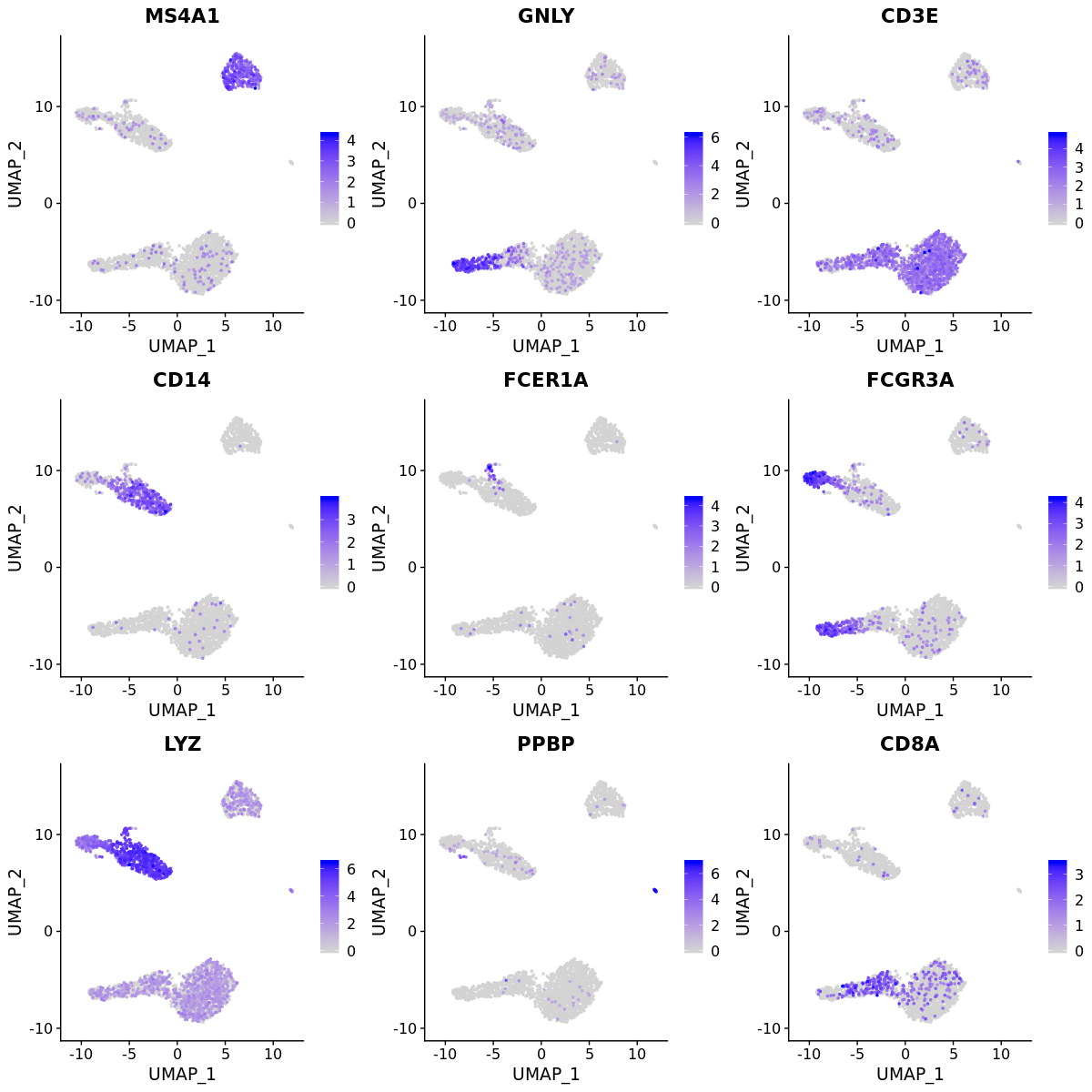
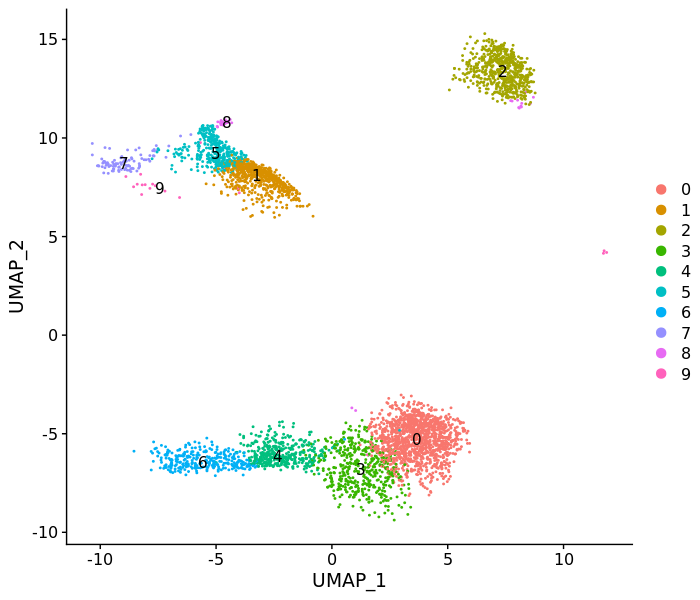
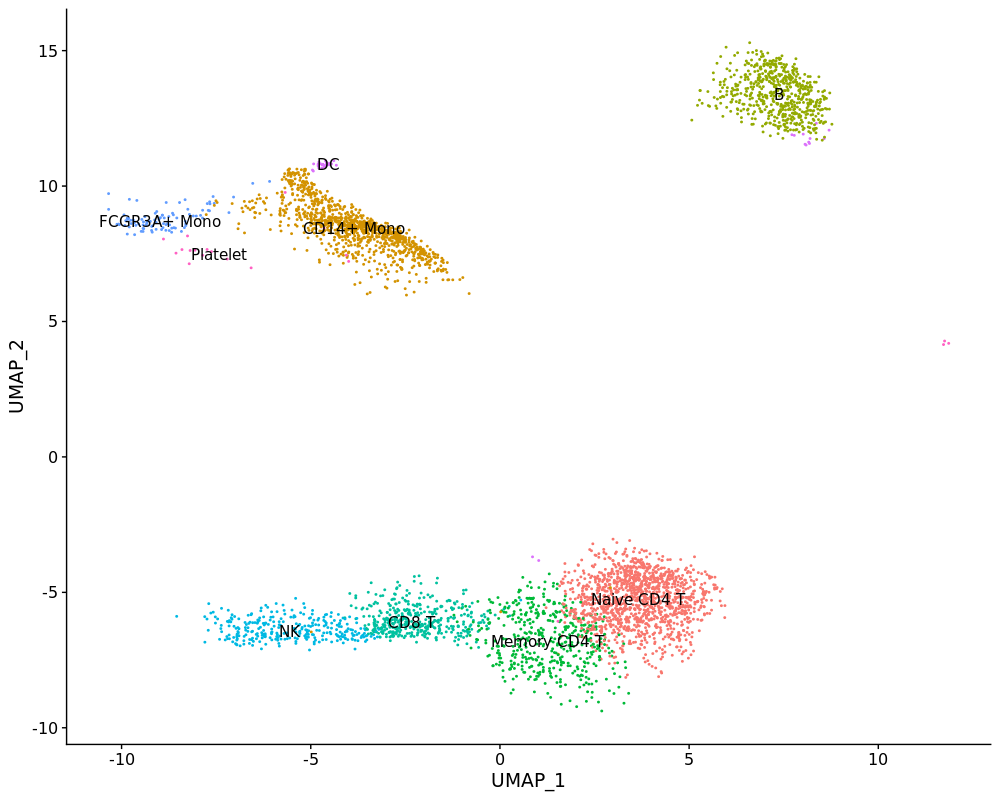
DimPlot(pbmc3k, label = T)FeaturePlot(pbmc3k, c("MS4A1", "GNLY", "CD3E",
"CD14", "FCER1A", "FCGR3A",
"LYZ", "PPBP", "CD8A"), order = T)We can use canonical markers to easily match the unbiased clustering to known cell types.
| Cluster ID | Markers | Cell Type |
|---|---|---|
| 0 | IL7R, CCR7 | Naive CD4+ T |
| 1 | IL7R, S100A4 | Memory CD4+ |
| 2 | CD14, LYZ | CD14+ Mono |
| 3 | MS4A1 | B |
| 4 | GNLY, NKG7 | NK |
| 5 | FCGR3A, MS4A7 | FCGR3A+ Mono |
| 6 | CD8A | CD8+ T |
| 7 | FCER1A, CST3 | DC |
| 8 | PPBP | Platelet |
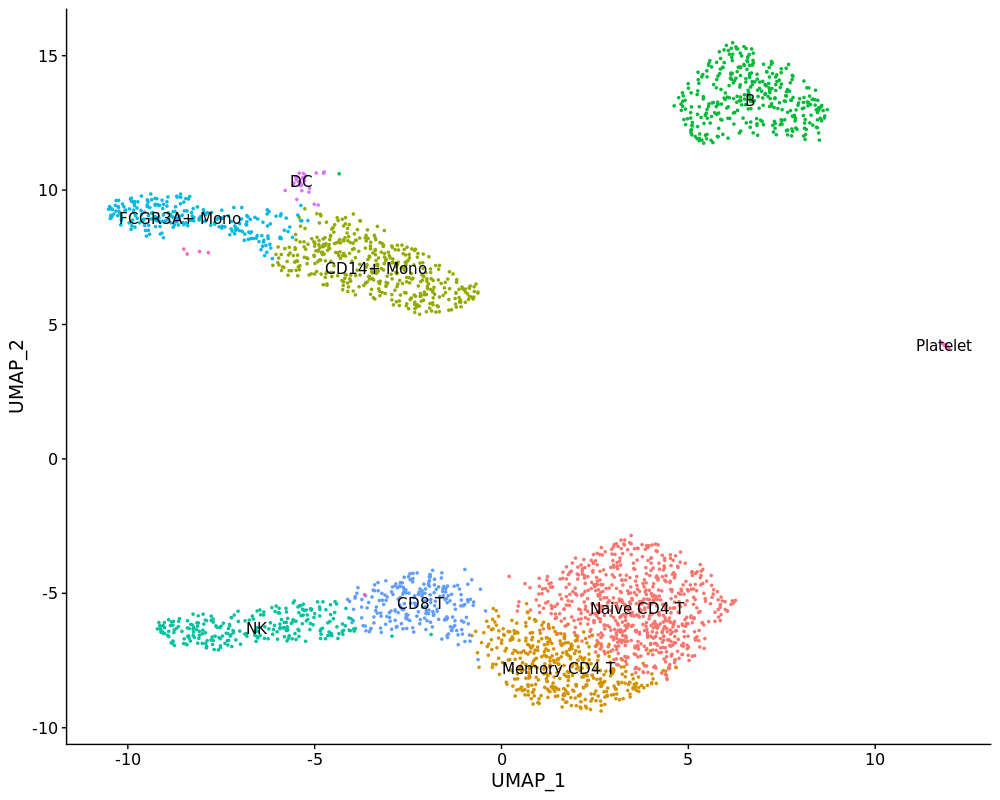
new.cluster.ids <- c("Naive CD4 T", "Memory CD4 T", "CD14+ Mono",
"B", "NK", "FCGR3A+ Mono",
"CD8 T", "DC", "Platelet")
names(new.cluster.ids) <- levels(pbmc3k)
pbmc3k <- RenameIdents(pbmc3k, new.cluster.ids)
DimPlot(pbmc3k, label = T) + NoLegend()Now we can project the normalized pbmc4k data into pre-calculated pbmc3k LSI space using projectLSI function.
matSVD.pbmc4k <- projectLSI(pbmc4k[["RNA"]]@data, pbmc3k.lsi)
pbmc4k[["pca"]] <- CreateDimReducObject(
embeddings = matSVD.pbmc4k,
loadings = pbmc3k.lsi$fLoad,
assay = "RNA",
key = "PC_")
# cluster cells using projected LSI
pbmc4k <- FindNeighbors(pbmc4k, dims = 1:10)
## Computing nearest neighbor graph
## Computing SNN
pbmc4k <- FindClusters(pbmc4k, resolution = 0.6)
## Modularity Optimizer version 1.3.0 by Ludo Waltman and Nees Jan van Eck
##
## Number of nodes: 4284
## Number of edges: 154662
##
## Running Louvain algorithm...
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
## Maximum modularity in 10 random starts: 0.8670
## Number of communities: 10
## Elapsed time: 0 seconds
# perform UMAP using first 10 PCs, just like pbmc3k
umap.pbmc4k.proj <- uwot::umap_transform(matSVD.pbmc4k[, 1:10], umap.pbmc3k, verbose = T)
## 01:37:48 Read 4284 rows and found 10 numeric columns
## 01:37:48 Processing block 1 of 1
## 01:37:48 Writing NN index file to temp file /tmp/RtmpbWcqgH/file17b933607747
## 01:37:48 Searching Annoy index using 8 threads, search_k = 3000
## 01:37:48 Commencing smooth kNN distance calibration using 8 threads
## 01:37:48 Initializing by weighted average of neighbor coordinates using 8 threads
## 01:37:48 Commencing optimization for 167 epochs, with 128520 positive edges
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
## 01:37:50 Finished
rownames(umap.pbmc4k.proj) <- colnames(pbmc4k)
colnames(umap.pbmc4k.proj) <- paste0("UMAP_", seq_len(ncol(umap.pbmc4k.proj)))
pbmc4k[["umap"]] <- CreateDimReducObject(
embeddings = umap.pbmc4k.proj,
assay = "RNA",
key = "UMAP_")
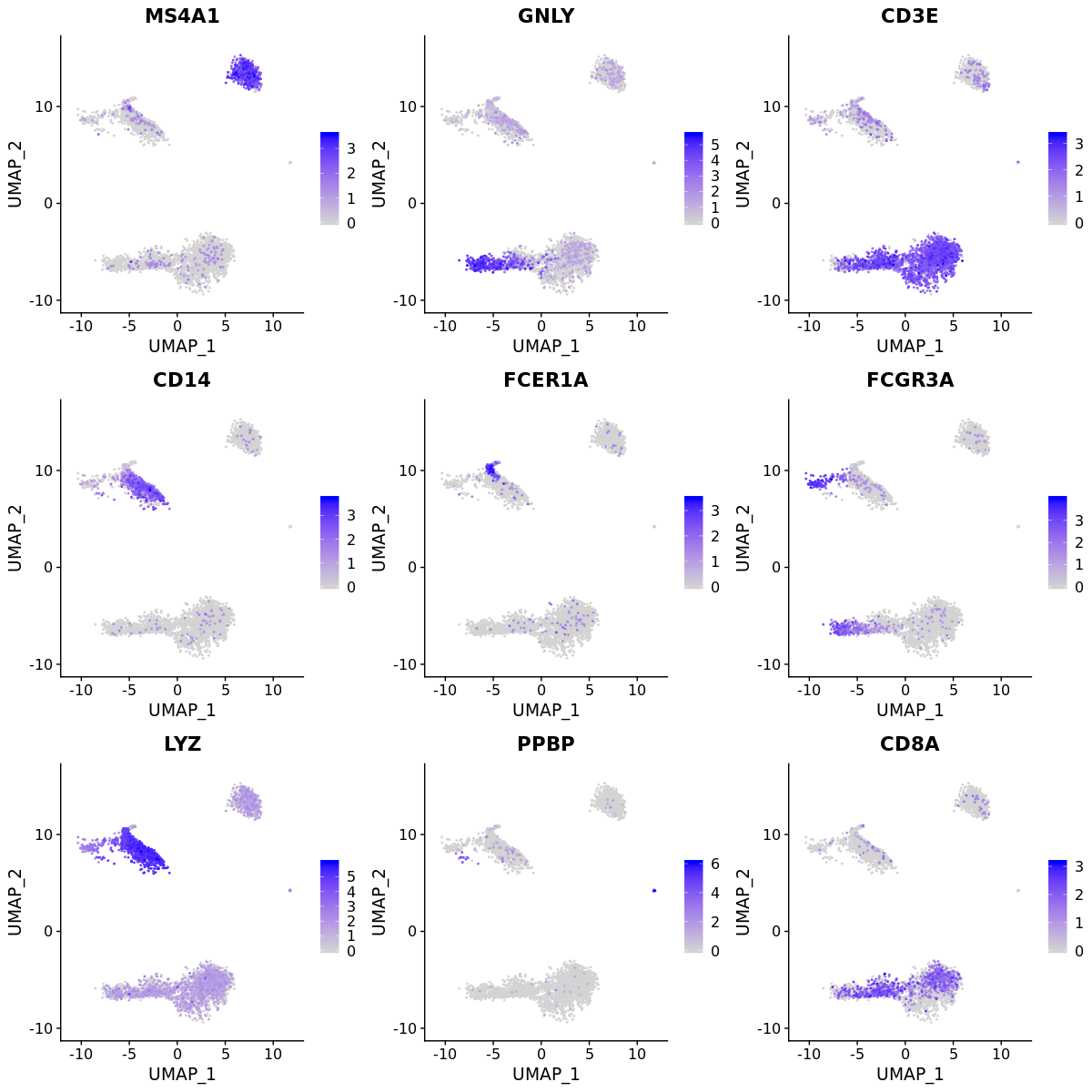
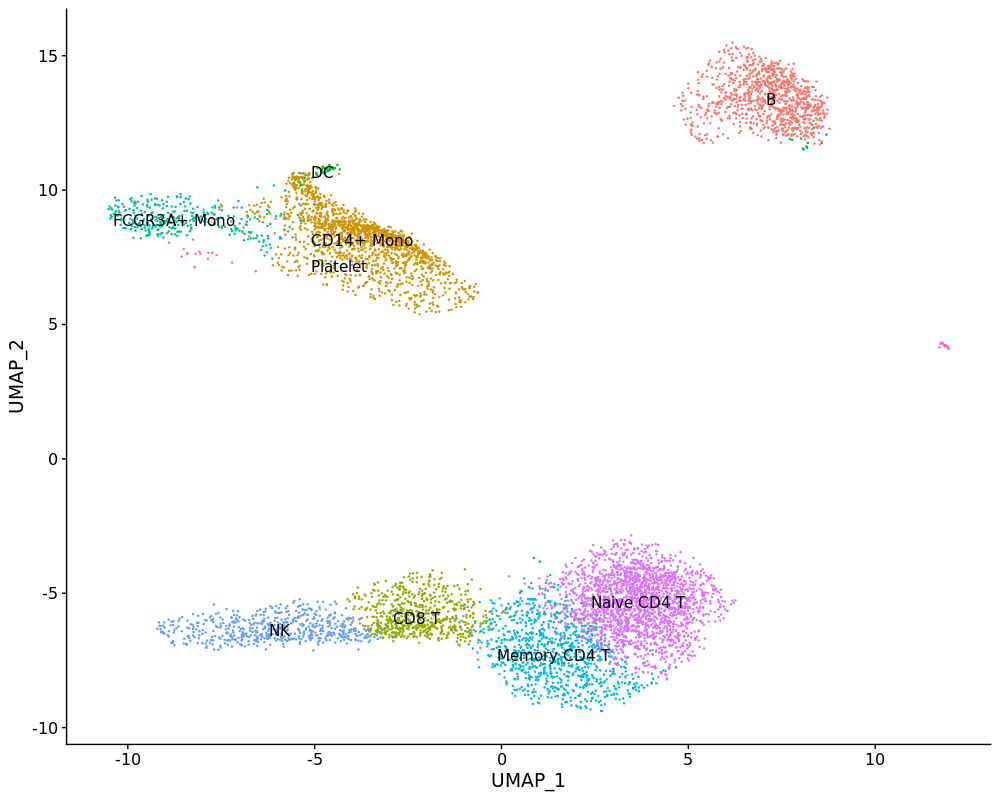
DimPlot(pbmc4k, label = T)FeaturePlot(pbmc4k, c("MS4A1", "GNLY", "CD3E",
"CD14", "FCER1A", "FCGR3A",
"LYZ", "PPBP", "CD8A"), order = T)new.cluster.ids <- c("Naive CD4 T", "CD14+ Mono", "B", "Memory CD4 T",
"CD8 T", "CD14+ Mono", "NK", "FCGR3A+ Mono",
"DC", "Platelet")
names(new.cluster.ids) <- levels(pbmc4k)
pbmc4k <- RenameIdents(pbmc4k, new.cluster.ids)
DimPlot(pbmc4k, label = T) + NoLegend()We can merge pbmc3k and pbmc4k together simply.
pbmc7k <- merge(pbmc3k, pbmc4k)
pbmc7k[["umap"]] <- CreateDimReducObject(
embeddings = rbind(pbmc3k[["umap"]]@cell.embeddings,
pbmc4k[["umap"]]@cell.embeddings),
assay = "RNA", key = "UMAP_")
DimPlot(pbmc7k, label = T) + NoLegend()pbmc7k$celltype <- Idents(pbmc7k)
Idents(pbmc7k) <- pbmc7k$orig.ident
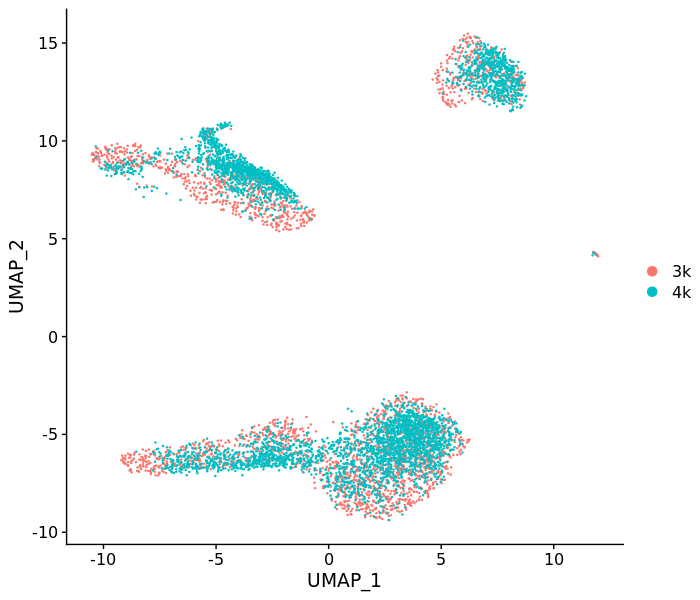
DimPlot(pbmc7k)We can see that the cells from pbmc4k locate on the UMAP space according to the pbmc3k cells of the same cell type.
First, we can down-sample the bulk RNA-seq data to get psudo-single-cell data.
psudo.all <- psudoSC(bulk.data, n = 100, depth = 3000)
## downsampling counts...
## merging all samples...
dim(psudo.all)
## [1] 25498 2000Then we can project the psudo-single-cell data into pbmc3k LSI just like pbmc4k.
psudo.so <- CreateSeuratObject(psudo.all, project = "bulk")
psudo.so <- NormalizeData(psudo.so)
## Performing log-normalization
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
Idents(psudo.so) <- rep(c("CD4T.bulk", "CD8T.bulk", "NK.bulk", "B.bulk", "Mono.bulk"), rep(400, 5))
bulk.matSVD <- projectLSI(psudo.so[["RNA"]]@data, pbmc3k.lsi)
umap.bulk.proj <- uwot::umap_transform(bulk.matSVD[, 1:10], umap.pbmc3k, verbose = T)
## 13:16:17 Read 2000 rows and found 10 numeric columns
## 13:16:17 Processing block 1 of 1
## 13:16:17 Writing NN index file to temp file /tmp/RtmplGf1gl/file1a6a91b3753
## 13:16:17 Searching Annoy index using 8 threads, search_k = 3000
## 13:16:17 Commencing smooth kNN distance calibration using 8 threads
## 13:16:17 Initializing by weighted average of neighbor coordinates using 8 threads
## 13:16:17 Commencing optimization for 167 epochs, with 60000 positive edges
## 0% 10 20 30 40 50 60 70 80 90 100%
## [----|----|----|----|----|----|----|----|----|----|
## **************************************************|
## 13:16:18 Finished
rownames(umap.bulk.proj) <- colnames(psudo.so)
colnames(umap.bulk.proj) <- paste0("UMAP_", seq_len(ncol(umap.bulk.proj)))
psudo.so[["umap"]] <- CreateDimReducObject(
embeddings = umap.bulk.proj,
assay = "RNA",
key = "UMAP_")
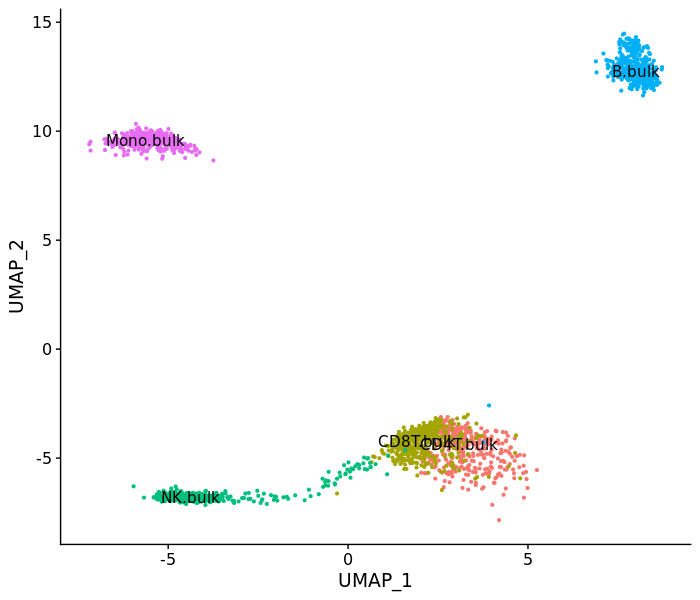
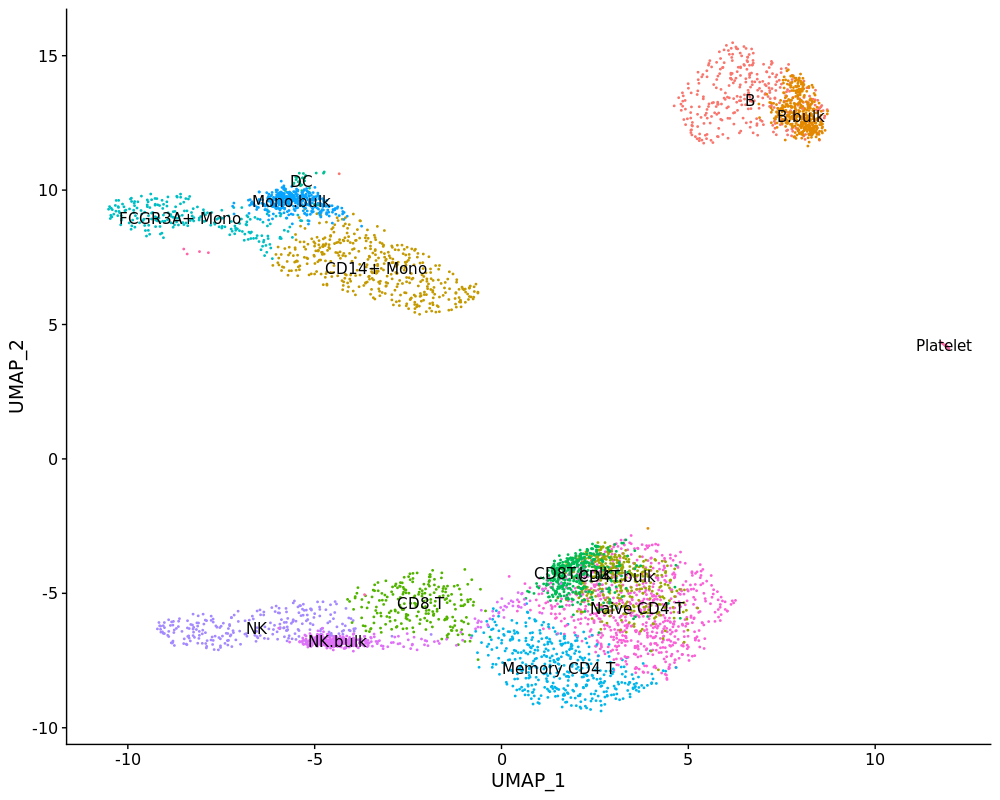
DimPlot(psudo.so, label = T)pbmc.mix <- merge(pbmc3k, psudo.so)
pbmc.mix[["umap"]] <- CreateDimReducObject(
embeddings = rbind(pbmc3k[["umap"]]@cell.embeddings,
psudo.so[["umap"]]@cell.embeddings),
assay = "RNA", key = "UMAP_")
DimPlot(pbmc.mix, label = T) + NoLegend()However, the CD8T cell bulk sample doesn't match single-cell data while others look good.