This protocol is derived from Rao et al., 2021.
Method 1:
If git command is available on the machine you want to run the pipeline, it can simply be downlaod using the following command:
git clone https://github.com/satyanarayan-rao/star_protocol_enhancer_cooperativity.git
Method 2
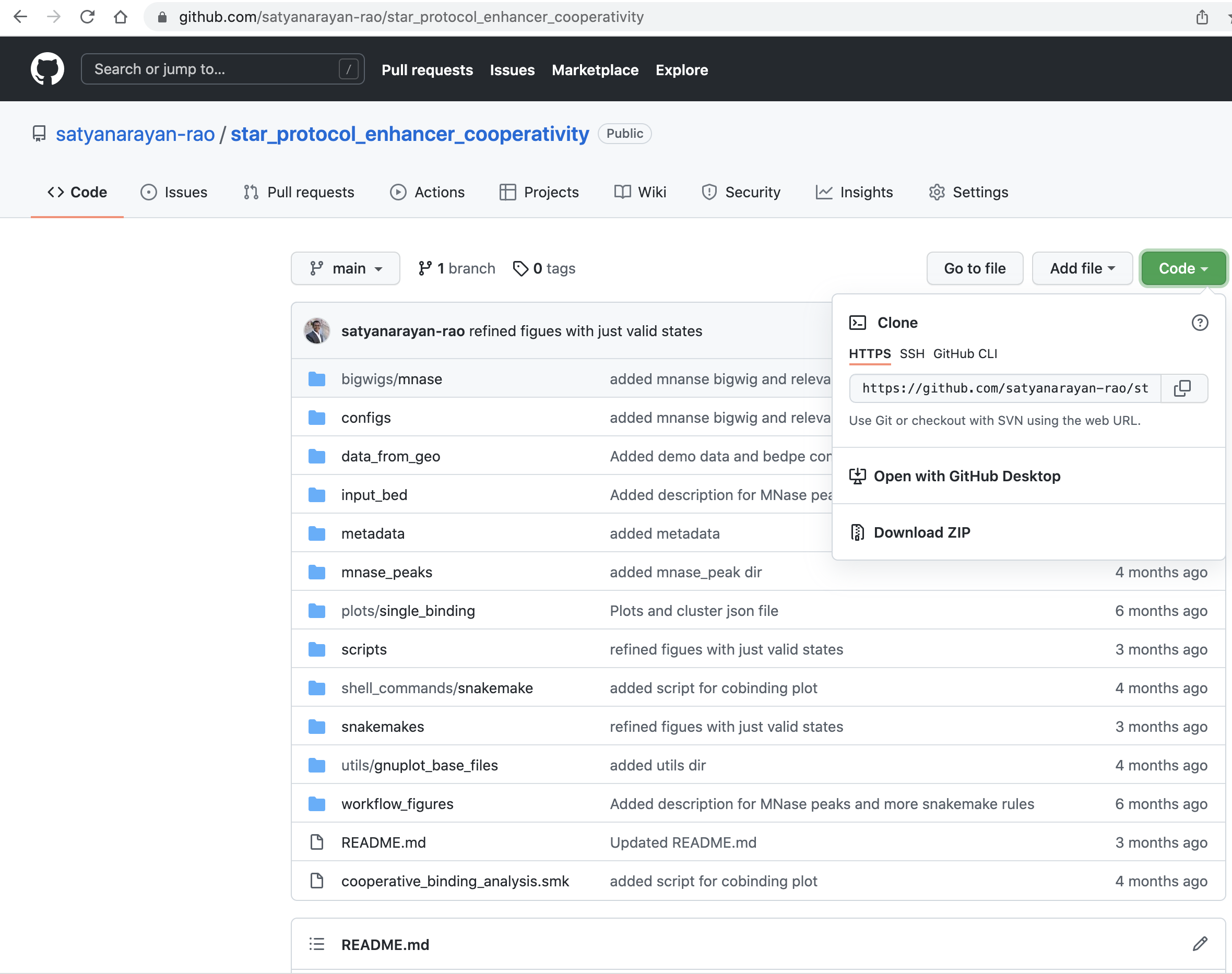
Please visit the github repository here. Please click on the code and choose "Download Zip" option as shown in the image below.
This pipeline is Linux/Unix-based system compatible.
Please install Anaconda Individual Edition first.
Please follow the steps below to build right environment to run the pipeline.
- Create an environment
dsmf_vizusing the command:conda create -n dsmf_viz python=3.6 - Activate this this environment using command
source activate dsmf_viz - Run
install_required_packages.shto install required packages mentioned below:- Bowtie2
- Bismark
- Trim Galore
- Snakemake
- Bedtools
- Samtools
- Bamtools
- pyBigWig
- Pandas
- Numpy
- Tbb
- Gnuplot
- Ghostscript
- Perl
CAUTION: Please run install_required_packages.sh only after activating the virtual environment (dsmf_viz) to avoid conflicts with existing package installations
Please run the following command to download dm3 reference genome.
$ sh download_reference_genome.sh
Data for demo is included in this github repository, but to visualize at your
sites of interest, please download the sequencing data, and keep them in
data_from_geo/. Here is the list of URLs for the sequencing data.
ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR313/006/SRR3133326/SRR3133326_1.fastq.gz
ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR313/006/SRR3133326/SRR3133326_2.fastq.gz
ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR313/007/SRR3133327/SRR3133327_1.fastq.gz
ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR313/007/SRR3133327/SRR3133327_2.fastq.gz
ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR313/008/SRR3133328/SRR3133328_1.fastq.gz
ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR313/008/SRR3133328/SRR3133328_2.fastq.gz
ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR313/009/SRR3133329/SRR3133329_1.fastq.gz
ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR313/009/SRR3133329/SRR3133329_2.fastq.gz
-
configs/: contains configuration file for the pipeline. Please see the exmapledemo_S2inconfigs/config.yamlto add your own sample information.configs/cluster.jsoncontains information for submitting jobs on cluster. Plese contact your cluster system administrator to configure this json file accordingly. -
input_bed/: Here user should keep regions of interest in a bed file. Please look atinput_bed/example.bedfor mapping binding at single sites, and seeinput_bed/example_cobinding.bedpefor mapping binding at pair of sites. -
data_from_geo/: This directory contains raw sequencing reads -
ref_genome/: This directory contains reference genome of your interest -
metadata/: This directory contains meta information, for example, genome size file,metadata/dm3.chrom.sizes. Please use appropriate genome size correspoding to the reference genome! -
plots/: Contains subdirectories with output pdf visualizing footprints and methylation maps -
utils/gnuplot_base_files/: Contains gnuplot commands in files that are used while plotting -
scripts/: Contains required scripts to run the pipeline -
snakemakes/: Contains modularized snakemake files. File names are self-explanatory -
workflow_figures/: Contains snakemake workflow image. Names of rules in the image can be traced in the snakemake files
Please run the following single command.
snakemake --snakefile cooperative_binding_analysis.smk plots/single_binding/suppressed_merged_demo_S2_to_example_spanning_lf_15_rf_15_extended_left_150_right_150_roi_peak_229.fp.pdf plots/single_binding/suppressed_merged_demo_S2_to_example_spanning_lf_15_rf_15_extended_left_150_right_150_roi_peak_229.methylation.pdf --configfile configs/config.yaml
snakemake --snakefile cooperative_binding_analysis.smk plots/cobinding_bedpe/suppressed_merged_demo_S2_to_example_cobinding_lf_15_rf_15_extended_left_300_right_300_roi_peak_110_4_and_peak_110_6.fp.pdf plots/cobinding_bedpe/suppressed_merged_demo_S2_to_example_cobinding_lf_15_rf_15_extended_left_300_right_300_roi_peak_110_4_and_peak_110_6.methylation.pdf --configfile configs/config.yaml
The advantage of Snakemake is that a user can incorporate parameters in file names. Related to this, below I expand on parameters placed in the output file names:
File name: plots/single_binding/suppressed_merged_demo_S2_to_example_spanning_lf_15_rf_15_extended_left_150_right_150_roi_peak_229.fp.pdf
-
demo_S2: points to the samples. Please take a look at samples starting withdemo_S2indata_from_geo/samples.tsvand also look atbam_merge_config->demo_S2inconfigs/config.yamlfile -
example: points toinput_bed/example.bed -
15: span 15bp from the ROI center;lfmeans span left, andrfmeans span right. This parameter is used in defining TF footprint. -
150: span 150 bp from ROI center. This is for visualization purpose. A dSMF molecule in principle could be as long as 300 bp, thus spanning 150 bp left and right respectively. -
peak_229: Name of the ROI. This name can be found as the fourth column ininput_bed/example.bed
File name: plots/cobinding_bedpe/suppressed_merged_demo_S2_to_example_cobinding_lf_15_rf_15_extended_left_300_right_300_roi_peak_110_4_and_peak_110_6.fp.pdf
-
demo_S2: Same as above -
example_cobinding: points toinput_bed/example_cobinding.bedpe; CRITICAL: the file name should have.bedpeextension and should followbedpeformat. -
15: same as above: this parameter will be used for defining TF footprints at both ROIs -
300: span 300bp from the left ROI (Chromosom location of ROIleft < ROIright) -
peak_110_4_and_peak_110_6:name_of_left_ROIandname_of_right_ROI; this name can be found ininput_bed/example_cobinding.bedpe