Exam Format:
- "Cheat Sheet": double-sided, can be typed
- Multiple-choice questions (for breadth)
- Smaller in-depth questions (for depth): may require bit of code (like PySpark) or explaining how to solve a problem w/ Pseudocode
- Content is material from homework, lectures, readings (emphasis on first two)
- Acquisition, access: needs to be accessible
- Wrangling: data may be in wrong form
- Integration, representation: data relationships may not be captured
- Cleaning, filtering: data may have variable quality
- Hypothesizing, querying, analyzing, modeling: from data to information
- Understanding, iterating, exploring: build knowledge from information
- Ethical obligations
- Need to protect data, follow good statistical practices, present results in a non misleading way
Goal is Raw Data to Structured Data to Information to Knowledge
- Myth:
- Learn "bottom up" using fancy statistics and machine learning
- "Turn up crank" and pop out insights
- Reality:
- Rely on human expertise to impose models over the data
- Deep learning for feature selection
- * No clear consensus
- Too complex for humans to understand directly
- Doesn't fit single uniform memory space (variables in python)
- Need more than brute algorithms to analyze
- May require multiple computers
- Possible high dimensionality (feature selection and dimensionality reduction)
- Netflix Prize
- High-Throughput Gene Sequencing
- Epileptic Seizure Prediction
- Highly dimensional, hard to understand, requires understanding computation & IO costs
- Relies on extracting data and selecting features and adding semantic structure to data
- Logical representation of data that isn't dependent on
- Where things are located
- Specific in-memory data structures (array vs. linkedlist, tree vs. map)
- What pointers, indices are available
- "implicit" properties like order of access
- "Physical data independence":
- Led to relational database
- Needs abstraction more general than in-memory data structure
- Split across a cluster?
- Should allow for efficient bulk computation (filter, merge, extract)
Essentially a kind of table "think as a relation across values or a list of maps w/ the same keys/values"
- Camps of Big Data People:
- Programming languages and compiler developers
- Software engineers
- R and Matlab users
- Database developers and users
- MapReduce / Spark
- Basic Abstraction: Dataframe
- Default: table w/ named columns and integer-identified rows
pandas.DataFrame.from_dict()pandas.read_csv()pandas.read_json()
- Accessing Elements of DataFrame:
- Rows as Tuples:
for iter in my_frame.iterrows():row = iter[1] // (v1, v2, v3...)
- Rows by Keys & Series
for col in my_frame:my_frame[col] // k-v
- Element by Index
my_frame.ix[0] // first rowmy_frame.ix[:,'first'] // first columnmy_frame.ix[0,'first'] // first cell
- Rows as Tuples:
- Setting Value:
my_frame.set+value(0, 'first', 1.0)(sets[0, 'first']to1.0)
- Iteration in python is slow
- Order doesn't matter as long as final output combined the right way
- Way to do multiple operations at the same time?
- The basis of parallel computing, GPU computing, vector instructions, databases, MapReduce
- Extracting subsets of a DataFrame
- Selection / filtering: rows of certain data values
- Projection
- Want certain columns
- Composition of DataFrames:
- Concatenation: append a table to another
- Joining: merge rows
- Can select rows by passing in Boolean vector of the same length
my_frame[[True, False, True]]
- DataFrames that use
NaNto represent a "null" value dropna: remove rowsfillna: substitute value
- Map each element by applying a function w
applymap - ex:
my_frame.applymap(my_fn)
pd.DataFrame(my_frame['first']): frames of columnfirstpd.DataFrame(my_frame['first'] > 1.0): boolean column representing if which rows havefirst> 1.0pd.DataFrame(my_frame(my_frame['first'] > 1.0)): all rows where the condition"first" > 1.0is true
my_frame.append(my_frame2): concatenates rows of two framesmy_frame.append(my_frame2).drop_duplicates(): concatenates rows of two frames without repeat rows
- pseudo-code ex: matches all 'to' in left w 'from' in right
dict_x = {'from': 1, 'to': 2}
left, right = pd.DataFrame.from_dict(dict_x)
left.merge(right, left_on=['to'], right_on='from')- pseudo-code ex: non-matched are included with NaN on the un-matched column
left.merge(right, left_on='to', right_on='from', how='outer', suffixes=('_l', '_r'))- Renaming column:
my_frame.rename(columns={'first':'spam'}) - Changing index columns:
my_frame.reset_index()ormy_frame.set_index('second')- reset index keeps previous index as a new column
- Change column type:
pd.to_numeric(my_frame['third'], errors='coerce')(forces NaN if non-numeric),my_frame['second'].astype(str)
- Exist in other platforms too: SQL (relational algebra), MapReduce (specific patterns of map / reduce functions), Spark (combinations of patterns and built-in functions)
- Store in DataBase to pass DF between programs / store excess data
- Lingo:
import sqlite3
engine = sqlite3.connect('my_database')
my_frame.to_sql('my_table', engine, if_exists='replace')
pd.read_sql('select * from my_table', engine)- SQL vs. NoSQL:
- SQL offers transactional semantics / expects very regular structure
- NoSQL offers less clean support for updates but can offer faster queries as result
- MySQL vs. PostgreSQL vs. MongoDB
- Docker containers can install / use these quite easily
- Basic capabilities for item-at-a-time traversal (discouraged) + operators for filtering, merging, etc
- Grouping by some fields ("Show me the counts FOR EACH") into "bins"
- Aggregation: "compute the x for each bin"
df.sort_values(['count', 'source'], ascending=False)
df['count'].plot(kind='line', title='Ayy')
TODO
- Huge DFs requiring expensive processing
- Put data on disk
- Distribute data and computation: across CPUs / GPUs + across machines (compute cluster or cloud center)
- Followup Challenges: retrieving data efficiently (even if changing in flight)
- Examples:
- Logs in Web company: platforms like DataDog, Apache Flume
- Order Processing / Analytics (Amazon)
- Ad analytics (Facebook, Google)
- Twitter meme / bot detection
- Search engine
- Facial / entity recognition from images
- Gene sequence matching pipelines
- Social network friend recommendation
- Disk is slow (even SSD)
- Don't want to retrieve all data in order to do computation
- Need notation of indexing
- If queries + updates happen simultaneously, need to handle concurrency via isolation -* Requirements lead to database management system
- data is interrelated (notion of "schema")
- Ensures data is "clean" w constraints: notation of "key" (relationship w other tables through "foreign keys")
- Handles concurrent updates
- Improves performance by optimization
- Example:
Person(SSN, Name, DOB):- SSN is 9-digit Integer
- Name is alpha-character string
- DOB is a date
- Key: set of fields that uniquely identify a tuple (Ex: SSN)
- Relational databases have "designed" relationships b/w tables (table w/ courses, names and departments, table w/ departments and building have relationship of department in both)
- Ex: a department offers many courses
- Relational databases require atomic valued attributes {cells}
- No complex types (lists, dictionaries)
- "First normal form"
- Relationships are encoded using keys (the "One") and foreign keys (the "Many")
- Ex: Students take many courses, courses are taken by many students
- Not every relationship is hierarchical
- JSON-embedded relationships vs. JSON-referenced relationships:
- Embedded will repeat a value multiple times (a student with their name, SID, etc) instead of just pointing to the data w a reference
- Selecting Data:
- DF:
student_df[student_df["name"] == "Maya"] - SQL:
SELECT * FROM Student WHERE Name="Maya"
- DF:
- Projecting Columns:
- DF:
pd.DataFrame(student_df["Name"]) - SQL:
SELECT (DISTINCT) Name FROM Student(paren ~ optional)
- DF:
- Merging (Joining):
- DF:
pd.merge(student_df, takes_df, on="SID") - SQL:
SELECT Student.Name, Student.SID, Takes.Num FROM Student JOIN Takes ON Student.SID = Takes.SIDalsoSELECT * FROM Student NATURAL JOIN
- DF:
- Grouping: "for each course, count the number of students"
- DF:
pd.DataFrame(takes_df.groupby(['Num']).size(),columns={'count'}) - SQL:
SELECT Num, count(*) AS Count FROM Takes GROUP BY Num
- DF:
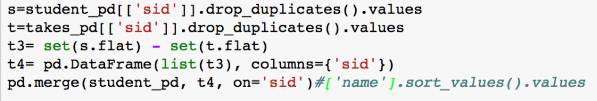
- Exists / Not-Exists: "Print all students who are not taking any course"
- SQL:
SELECT * FROM Student AS s WHERE NOT EXISTS (SELECT * FROM Takes AS t WHERE s.SID=t.SID) - DF:
- SQL:
- Data "in the wild" is in database
- Need to be able to store DFs in a DB and get from DB
- Paper by E. F. Codd [1970]: "relational model of data for large shared data banks"
- Separate physical implementation from logical
- Model the data independently from how it will be used (accessed, printed, etc)
- Describe the data minimally and mathematically
- use standard mathematical operations over the data (relational algebra, relational calculus)
- Optimization was key to relational systems
- [1969-70] Codd's original work
- [1976] Earliest relational database research
- [1979] Oracle 2.0
- Success was optimization: indexing, optimization based on rewriting queries, optimizing joins
- Indices reduce the number of rows in a table that must be examined
- Think index of a book
- Database indices work b/w disk and main memory
- Getting DF from DB comes with added index column
- Create / get indices from table:
c = engine.cursor()
c.execute("create index my_index on my_table(second)")
pd.read_sql('pragma index_list("my_table")', engine)When is an index used:
- If the initial columns of the index (a, b, etc) appear in the WHERE clause terms as:
- column = expression
- column IS expression
- column in (subquery)
- column is NULL
- Right-most column of an index that is used can use inequality (restricting value for a column within a range)
- column > expression
- Example:
CREATE INDEX idx_R ON R(a,b,c)WHERE a=5 AND b=3 AND c=NULL: all columns usableWHERE a=5 AND b>10 AND b<50: first two columns usableWHERE a=5 AND b>12 AND c='hello': first two columns usableWHERE b=3 AND c='hello': index not usable
relational algebra can simplify an equation for easier queries
- Processing the Query: Query -> Optimizer -> Execution Engine (also inputs Storage System) -> Web Server
- Embody an "all or nothing" unit of work
- despite failures in system, concurrent activity, etc
- How should conflicts be handled: "serializable behavior"
- Say two transactions try to add different students but give both the same SID
- Old Model: CPUs and Memory Modules (RAM) fully connected
- Equal access to every processor and memory
- "crosswork network": slow, expensive
- If computer is not enough: buy many computers with small number of processors, connect on high-speed network into a cluster
- Non-uniform costs in parallel systems
- Recall lot of attention paid to disk algorithms:
- Internal CPU state (registers, cache)
- Memory: latency 1000s times slower
- SSD / disk: latency 1000s times slower
- Same is true with multiple processors / computers
- I/O latency and throughput / bandwidth
- Each processor should work "independently" and "locally" as much as poss.
- Recall lot of attention paid to disk algorithms:
-
Network switch: connects nodes w/ each other and other racks
-
Rack: holds nodes/blades (often identical)
-
* Data centers exists because cluster can become too big / hot
- Main issues with compute clusters: making computation parallel and minimzing communication and coordination
- Many of the same principals hold for multiple CPUs
- Analogy: 10,000 employees collate together on census form
- People take vacations, get sick, work at different rates, fill out form incorrectly, etc
- Abstract away into a Data Flow
-
Scheme 1: Server 1 does A-G, Server 2 does H-N, etc
- Critical Issue: Data skew
-
Scheme 2: Randomize placement of the data but make predictable (Use Spark)
- Hashing "spreads out" the values into predictable locations
-
Sharding: take index key, divide values into buckets, assign one bucket / machine, do work locally over the bucekets, exchange (shuffle) data, go to aggregate
-
Spark's Building Blocks: Simplest First
-
RDD {Resilient Distributed Dataset}: map from sharded index field to value
- Distributed data collection: set, dictionary, table
- It has an index or key to "shard" the data
- If the machine crashes, the computation will recover and reuse "resilient"
-
No program can directly iterate over all the elements of the RDD
- Instead define code to be applied on different machines / subsets of the data
-
TODO
- Simple Algorithm for a Centralized Graph:
- Input: Graph as adjacency list
- Output: dictionary (map) of degrees of all vertices
- Algorithm:
- For each vertex v
- Count pairs of (v, X) for all X in the table
- Store sum in D[v]
- Centrality: "most cited paper"
-
Random Walk: stateless way, randomly choose a neighbor and repeat when we return to our start node
- Given
$n$ nodes,$m$ edges: start at$u$ return from neighboring$v$ in$2m$ steps - E[Time to visit all nodes]
$\leq 2m(n-1)$
- Given
- Distance: number of edges on shortest path (use BFS)
-
Triadic Closure: one's friends become each other's friends
- Look to complete triangles to recommend friends
- Algorithm:
- Run BFS to find friends of friends
- For each such
$n$ , count how many friends are$n$ 's friends, rank each$n$ by how many friends in common
- Other Pair based Algorithms:
- Annotating edges with cost, distance, similarity
- MST: minimal cost tree connecting all vertices in graph
- Topological Sort, Steiner Tree (MST of certain nodes)
- Different notations of centrality have been defined based on connectivity
- "Betweenness centrality": measures how important a node is in bridging communities
- "Eigenvector centrality": Gives us a recursive measure of importance (ie do I connect to important nodes, do they connect to important nodes)
- Problem: Millions of pages contain query word
- Idea: hyper links encode human judgment (like citations)
- Intra-domain links: Created for navigation
- Inter-domain links: Confer measure of authority