This example shows how to simulate from a beta regression model. First, I do this using the rstanarm package for a fully Bayesian model. Second, I do this with a maximum likelihood model fit with the betareg package by modifying the function arm::sim().
Beta regression was just added into rstanarm, so you will need to install the package from GitHub. This will require a C++ compiler. Sometime in the near future I imagine you'll just be able to install the version from CRAN.
# install.packages("devtools")
# run this once to install rstanarm!!
# devtools::install_github("stan-dev/rstanarm", args = "--preclean", build_vignettes = FALSE)
library(rstanarm)
#> Loading required package: Rcpp
#> rstanarm (Version 2.14.1, packaged: )
#> - Do not expect the default priors to remain the same in future rstanarm versions.
#> Thus, R scripts should specify priors explicitly, even if they are just the defaults.
#> - For execution on a local, multicore CPU with excess RAM we recommend calling
#> options(mc.cores = parallel::detectCores())
options(mc.cores = parallel::detectCores())
library(betareg)First simulate some data to fit our models to. This is modified from the help file for rstanarm::stan_betareg.
set.seed(1)
N <- 150
x <- rnorm(N, 2, 1)
mu <- binomial(link = "logit")$linkinv(0.8 + 0.3*x)
phi <- exp(1.5)
y <- rbeta(N, mu * phi, (1 - mu) * phi)
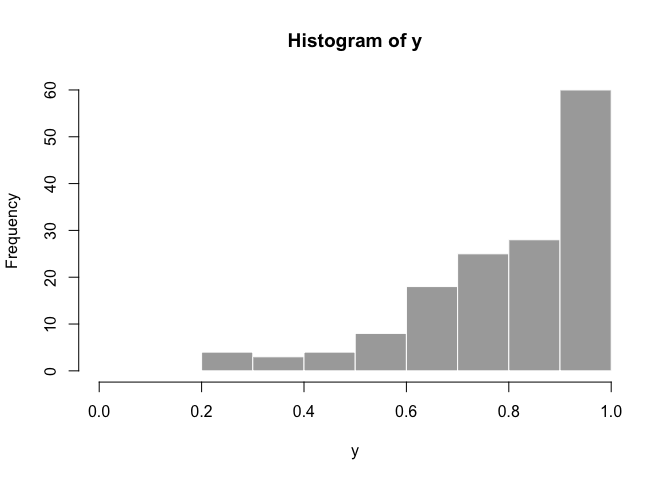
hist(y, col = "dark grey", border = FALSE, xlim = c(0,1))fake_dat <- data.frame(y, x)Now we can fit the model with rstanarm::stan_betareg:
fit <- stan_betareg(y ~ x, data = fake_dat, link = "logit")
print(fit)
#> stan_betareg(formula = y ~ x, data = fake_dat, link = "logit")
#>
#> Estimates:
#> Median MAD_SD
#> (Intercept) 0.6 0.2
#> x 0.4 0.1
#> (phi) 4.3 0.5
#>
#> Sample avg. posterior predictive
#> distribution of y (X = xbar):
#> Median MAD_SD
#> mean_PPD 0.8 0.0
fit$stanfit
#> Inference for Stan model: continuous.
#> 4 chains, each with iter=2000; warmup=1000; thin=1;
#> post-warmup draws per chain=1000, total post-warmup draws=4000.
#>
#> mean se_mean sd 2.5% 25% 50% 75% 97.5% n_eff
#> (Intercept) 0.55 0.00 0.19 0.18 0.43 0.56 0.68 0.93 3188
#> x 0.43 0.00 0.09 0.26 0.37 0.43 0.49 0.60 2948
#> (phi) 4.29 0.01 0.48 3.40 3.96 4.27 4.59 5.29 2594
#> mean_PPD 0.80 0.00 0.02 0.76 0.78 0.80 0.81 0.83 3134
#> log-posterior 101.04 0.03 1.21 97.84 100.46 101.36 101.93 102.41 1806
#> Rhat
#> (Intercept) 1
#> x 1
#> (phi) 1
#> mean_PPD 1
#> log-posterior 1
#>
#> Samples were drawn using NUTS(diag_e) at Mon Jan 9 17:45:33 2017.
#> For each parameter, n_eff is a crude measure of effective sample size,
#> and Rhat is the potential scale reduction factor on split chains (at
#> convergence, Rhat=1).
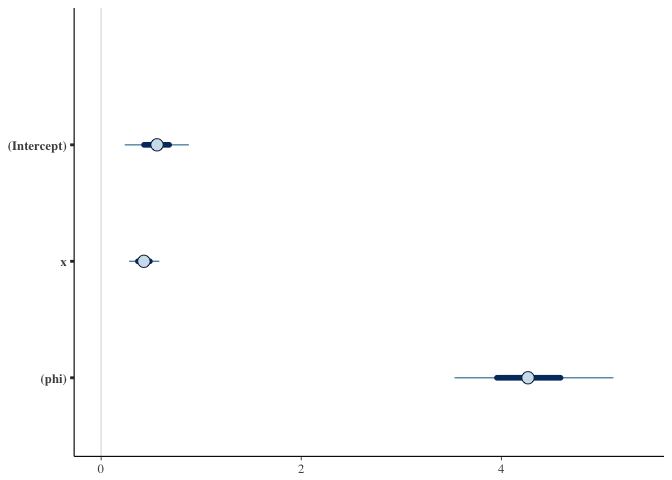
plot(fit)# pp_check(fit)
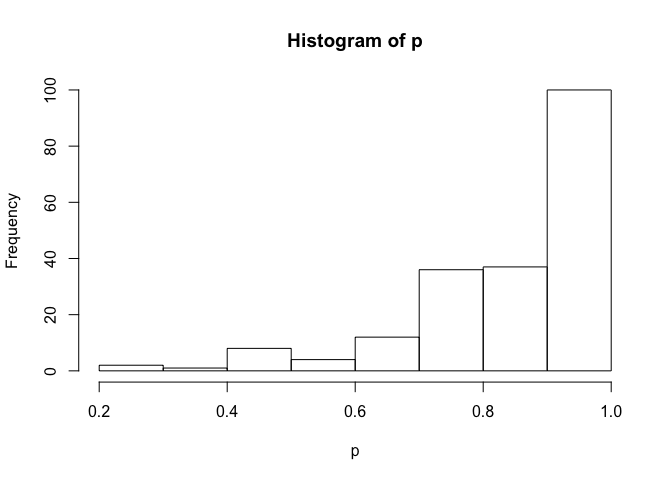
# prior_summary(fit)We can simulate new observations for a given set of predictor values. This includes both uncertainty in the mean and the beta observation distribution. Below, I picked 2 arbitrary values for x to simulate with.
newdata <- data.frame(x = 3)
p <- posterior_predict(fit, draws = 200, newdata = newdata)
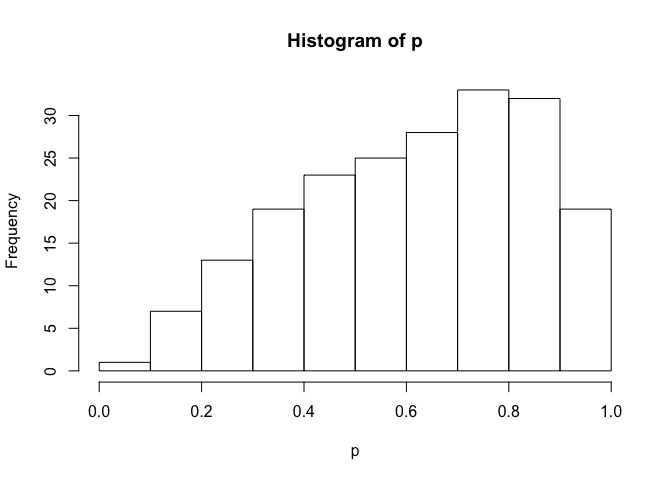
hist(p)newdata <- data.frame(x = -0.5)
p <- posterior_predict(fit, draws = 200, newdata = newdata)
hist(p)We can also take draws from the posteriors:
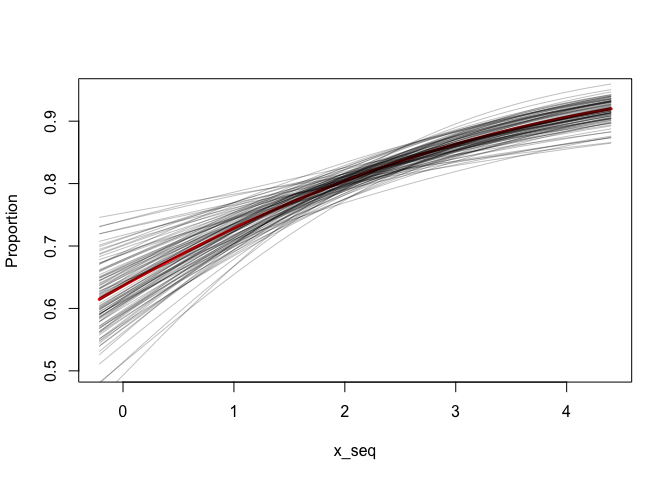
intercept <- as.matrix(fit)[,"(Intercept)"]
beta_x <- as.matrix(fit)[,"x"]
x_seq <- seq(min(x), max(x), length.out = 100)
plot(x_seq, plogis(coef(fit)[[1]] + coef(fit)[[2]] * x_seq), type = "l",
ylim = c(0.5, 0.95), col = "red", lwd = 3, ylab = "Proportion")
for (i in seq_len(100)) {
lines(x_seq, plogis(intercept[i] + beta_x[i] * x_seq), col = "#00000040")
}Now let's do something similar with the betareg package.
First I extracted the function arm::sim() from https://github.com/cran/arm/blob/master/R/sim.R
Then I modified it as needed to match the output from the betareg package:
sim_betareg <- function(object, n.sims = 100) {
object.class <- class(object)[[1]]
summ <- summary (object, correlation=TRUE, dispersion = object$dispersion)
coef <- summ$coefficients$mean[,1:2,drop=FALSE]
dimnames(coef)[[2]] <- c("coef.est","coef.sd")
beta.hat <- coef[,1,drop=FALSE]
sd.beta <- coef[,2,drop=FALSE]
k <- length(object$coefficients$mean)
corr.beta <- cov2cor(vcov(object))[1:k, 1:k]
V.beta <- corr.beta * array(sd.beta,c(k,k)) * t(array(sd.beta,c(k,k)))
beta <- array (NA, c(n.sims,k))
dimnames(beta) <- list (NULL, dimnames(beta.hat)[[1]])
for (s in 1:n.sims){
beta[s,] <- MASS::mvrnorm (1, beta.hat, V.beta)
}
beta2 <- array (0, c(n.sims,length(coefficients(object))))
dimnames(beta2) <- list (NULL, names(coefficients(object)))
beta2[,dimnames(beta2)[[2]]%in%dimnames(beta)[[2]]] <- beta
beta2 <- beta2[, 1:k]
beta2
}Now we can fit the model and take draws from the coefficients.
fit2 <- betareg(y ~ x, data = fake_dat, link = "logit")
summary(fit2)
#>
#> Call:
#> betareg(formula = y ~ x, data = fake_dat, link = "logit")
#>
#> Standardized weighted residuals 2:
#> Min 1Q Median 3Q Max
#> -2.1103 -0.7360 -0.0685 0.5154 3.5817
#>
#> Coefficients (mean model with logit link):
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 0.5518 0.1842 2.996 0.00274 **
#> x 0.4297 0.0864 4.973 6.58e-07 ***
#>
#> Phi coefficients (precision model with identity link):
#> Estimate Std. Error z value Pr(>|z|)
#> (phi) 4.3140 0.4965 8.69 <2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Type of estimator: ML (maximum likelihood)
#> Log-likelihood: 108.2 on 3 Df
#> Pseudo R-squared: 0.1404
#> Number of iterations: 12 (BFGS) + 1 (Fisher scoring)
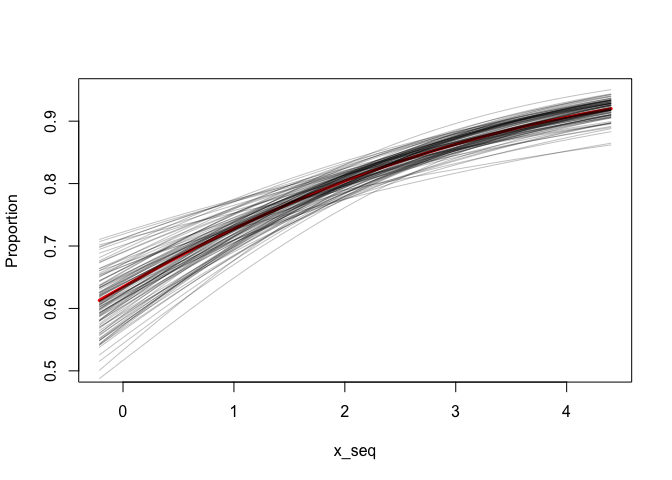
s <- sim_betareg(fit2, n.sims = 100)
x_seq <- seq(min(x), max(x), length.out = 100)
plot(x_seq, plogis(coef(fit2)[[1]] + coef(fit2)[[2]] * x_seq), type = "l",
ylim = c(0.5, 0.95), col = "red", lwd = 3, ylab = "Proportion")
for (i in seq_len(nrow(s))) {
lines(x_seq, plogis(s[i, 1] + s[i, 2] * x_seq), col = "#00000040")
}I think it's a little bit trickier if we also want to simulate new observations with the beta distribution because there is some correlation between the mean and the dispersion parameter.