Introduction
Multimodal models for Integrated Regulatory Analysis, or MIRA, is a python package for analyzing the dynamic processes of gene regulation using single-cell multiomics datasets.
MIRA works on top of Scanpy and Anndata to provide a rich, comprehensive framework integrating accessibility and expression data for more insights into your data. MIRA includes methods for:
- Multiomodal topic modeling
- Construction a joint representation of cells
- Regulator and functional enrichment
- Pseudotime trajectory inference
- Cis-regulatory modeling
- Finding divergences between local chromatin accessibility and gene expression
... And more! For mora, check out the MIRA preprint on bioarxiv.
Documentation
See MIRA's website for tutorials and API reference.
Data
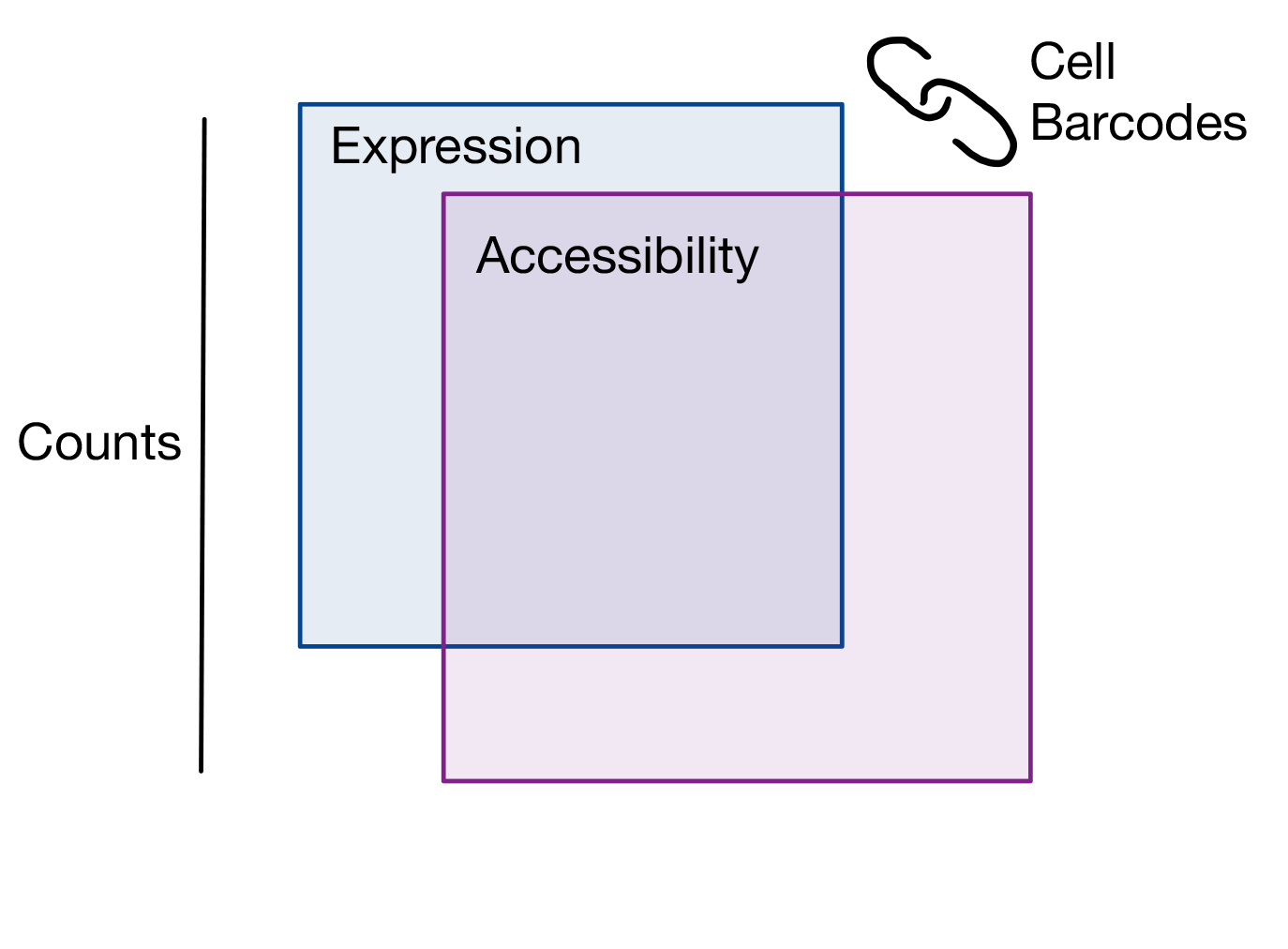
MIRA takes scRNA-seq and scATAC-seq count matrices from a single-cell multiomics experiment, where each cell is measured using both assays, and measurements are linked by a shared cell barcode. We demonstrated MIRA using SHARE-seq data and commercial 10X genomics multiome data, but MIRA's assumptions and models are extensible to other multiome protocols.
Installation
MIRA can be installed from either Conda or PyPI:
conda install -c conda-forge -c bioconda -c liulab-dfci mira-multiomeor
pip install mira-multiomeInstallation will take about a minute.
Please note, currently MIRA may only be installed on Python <3.8 due to some dependencies' requirements. We are working to make it accessible on newer Python versions. To set up an a new analysis, we recommend starting with a fresh environment:
conda create --name mira-env -c conda-forge -c pytorch -c bioconda mira-multiome scanpy jupyter leidenalg
conda activate mira-env
python -m ipykernel install --user --name mira-envTo use the environment in a jupyter notebook, start the notebook server, then go to Kernel > Change kernel > mira-env.
Installing with GPU support
Training on a GPU reduces the training time of MIRA topic models. To install MIRA with PyTorch compiled with GPU support, first install MIRA, as above. Then, follow instructions at pytorch.org to find the version of PyTorch that suits your system.
Learning Curve
If you have experience with Scanpy, we structured MIRA to follow similar conventions so that it would feel familiar and intuitive. In fact, most MIRA analyses seamlessly weave between MIRA and Scanpy functionalities for cleaning, slicing, and plotting the data. In general, the first positional argument of a MIRA function is an AnnData object, and the following keyword arguments change how the function transforms that object.
Dependencies
- pytorch
- pyro-ppl
- tqdm
- moods
- pyfaidx
- matplotlib
- lisa2
- requests
- networkx
- numpy
- scipy
- optuna
- anndata